Abstract
We report here on the characterization of a cDNA library from seeds
Introduction
The economic viability of using plant oils as renewable resources is critically dependent on the energetic balance of oil production in this system. Therefore, it is necessary to decipher the regulation of lipogenesis in maturing oilseeds.
Fatty acid synthesis occurs in the plastids. Many seeds accumulate large reservoirs of oil in the form of triacylglycerols (TAG). TAG assembly can be considered as proceeding by two distinct routes. 4 One of these, the Kennedy pathway, relies on the sequential acylation process of fatty acid biosynthesis of a glycerol-3-phosphate (Gly-3-P) backbone provided by GPDH. The second route relies on acyl-exchange between lipids and involves a phospholipid diacylglycerol acyltransferase (PDAT). Basically, TAGs are produced through the condensation of two fatty acid molecules with one molecule of phosphoric acid and one molecule of glycerol. The process of esterification produces various types of phosphatidic acids, which are subsequently dephosphorylated in the endoplasmic reticulum to form diacylglycerol (DAG). Finally, a fatty acid chain is added to the DAG by an acyltransferase, resulting in a TAG that is sequestered in lipid bodies for storage. 5 TAGs are used to support germination and early seedling growth. In this sense, fatty acids act as molecules of energy storage that are used to “fuel” the cell when other energy sources are not available.
During seed maturation, hexose phosphates from photo-assimilates are massively metabolized through the glycolytic pathway before being used for
Fatty acids chains longer than palmitate are formed by elongation reactions catalyzed by enzymes on the cytosolic side of the endoplasmic reticulum membrane. These reactions add two carbon units (Malonyl CoA) sequentially to the carboxyl ends of both saturated and unsaturated fatty acyl CoA substrates.
To be used as energy fuel, the fatty acids must be activated and transported into mitochondria for degradation ((3-oxidation). The fatty acids are then broken down in a step-by-step way into acetyl CoA, which is processed in the citric acid cycle. The conversion of fatty acid into energy (NADH, ATP) occurs in the citric acid cycle.
Acetyl CoA carboxylase plays an essential role in regulating fatty acid synthesis and degradation. Low levels of acetyl-CoA induce fatty acid (3-oxidation, which results in ADH2, NADH and acetyl-CoA. The excess of acetyl-CoA results in production of excess citrate (citric acid cycle), which is exported into the cytosol to give rise to cytosolic acetyl-CoA. A high level of citrate means that two-carbon units and ATP are available for fatty acid synthesis. Acetyl-CoA can be carboxylated into malonyl-CoA that is required for (i) synthesis of flavonoids and related polyketides, (ii) elongation of fatty acids to produce waxes, cuticule, and seed oils, and (iii) malonation of proteins and other phytochemicals such as terpenes, steroids and sterols. 7
The terpenoids constitute the largest class of natural products produced by plants. Terpenoids form groups of related structural types that allow the detection of chemosystematic relations among species. The plant families of
Despite the attractive features of its oil composition and productivity,
There is a critical need for scientific breeding of Jatropha guided by advanced DNA mapping technologies.15,16 Individuals of
The identification of quantitative traits that improve agricultural production allows breeders to introduce economically important traits into modern genetic backgrounds and to investigate the molecular mechanisms that regulate their effects. 27 For instance, map-based cloning of a diacylglycerol acyltransferase (DGAT) that catalyzes the final step in the glycerol biosynthetic pathway allowed the selection of a new protein variant influencing oil content and composition in maize seeds. 28
For this reason, this study aimed to (i) describe the complexity of the oil produced by this plant, (ii) describe the transcriptome complexity during seed maturation through the analysis of a cDNA library constructed from the mRNA of seeds at three different development stages, (iii) tag genes related to the route of fatty acids and toxic compound metabolism and (iv) measure the difference of expression level between leaves and fruits of four genes (palmitoyl-acyl carrier protein thioesterase, 3-ketoacyl-CoA thiolase B, lysophosphatidic acid acyltransferase and geranyl pyrophosphate synthase) involved in some economically important traits. This information will be useful for the investigation of genetic diversity and for the selective breeding for traits in relation to the optimization of biodiesel production and low-phorbol varieties of
Materials and Methods
Construction of the cDNA Library
Fruits from
Sequencing of cDNAs
The plates were inoculated with a 96 pins replicator into 96 deepwell plates containing 1.2 ml LB with ampicilin and incubated at 37
EST Processing
Expressed Sequence Tags (EST) were processed stepwise in order to (i) extract their regions corresponding to PHRED quality ^10, (ii) trim out vector sequences with CROSS-MATCH, 30 (iii) trim out polyA and polyT tail sequences with home-made Perl scripts, (iv) eliminate sequence redundancy by contig assembling with CAP3, 31 (v) annotate them for putative functions by comparison to GeneOntology (GO, http://www.geneontology.org) using Blast2GO 32 and selecting homologous pairs with E ≤ 0.0001, identity ≥ 40% and the homologous region ≥40 amino acids. These 931 ESTs were submitted to the dbEST section of GenBank with accession numbers GT228436-GT229366.
We also download the 13193 ESTs of
For the reasons outlined above, we compared the homology of the 5841 non-redundant ESTs from GenBank to the KEGG accessions including an EC number and selected the homologies that mapped (http://www.genome.jp/kegg/tool/color_pathway html) to the metabolism of fatty acids (map 61, 62, 71, 590, 591, 592, 1040), lipids (maps 561, 564, 565), terpenes (maps 900, 902, 904, 909), alkaloids (maps 901, 950, 1063, 1064, 1065, 1066), quinones (map 130), drugs (maps 982, 983) and hormones (maps 150, 905, 1070).
We compared (BLASTX) the 5841 non-redundant ESTs from GenBank (Rel. 174, Dec 14, 2009) with the KEGG sequences annotated with EC numbers (955689) used as the reference dataset.
Measure of Gene Expression by Real-Time PCR
The level of expression of palmitoyl-acyl carrier protein thioesterase, 3-ketoacyl-CoA thiolase B, lysophosphatidic acid acyltransferase and geranyl pyrophosphate synthase was compared to that of actin in a leaf sample and three seed samples corresponding to the three fruit stages described under “Construction of the cDNA library” (see above). The RNA was extracted from seeds and leaves of
The quality of amplicons was first checked with genomic DNA under conditions suitable to quantitative real-time PCR. The reference gene used as control to measure the constitutive expression was the actin gene from
The average and standard deviation were obtained from three replicates. The average of actin expression of seeds across the fruit stages was, first, normalized according to that of the leave sample. The linear correction performed according to the ΔΔCt method 35 was, then, applied to find the expression level of each gene in leaves and seeds. Finally, the average level of expression of each gene in seeds was divided by its corresponding value in leaves to obtain the multiplying factor associated to the over-expression of these genes in seeds at the three fruit stages.
Analysis of Fatty Acid Composition of Oil from J. Curcas
In addition to transcriptome characterization, we analyzed the fatty acid composition of
The derivatization of a sample of
Results
Assessing the Transcriptome and Fatty Acid Composition of J. Curcas Seeds
After analyzing the fatty acid composition
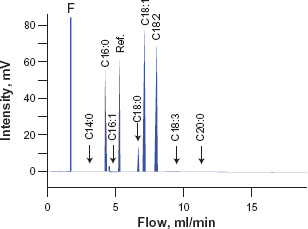
Gas chromatogram of
The double-stranded cDNA fragments obtained from the total RNA extract from seeds at three development stages (Fig. 2A) ranged from 300 to 2000 bp (Fig. 2B), as did the inserted fragments after cloning (Fig. 2C).
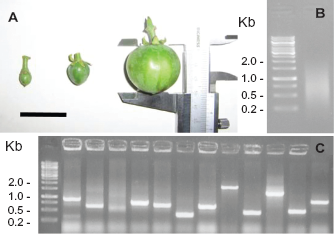
Steps toward library construction. Three different fruit stages were selected for seed RNA extraction. The black bar is three cm (
We sequenced 2200 cDNA clones, from which we recovered 1337 (60%) reads after quality control and trimming. Among them, 546 were clustered into 140 contigs while 791 remained singlets. Our final sample was therefore 931 non-redundant expressed sequence tags (EST) with PHRED quality higher than 10. The 140 contigs had an average size of 569 bp and the 791 singlets had an average size of 379 bp. The majority (64%) of contigs was made up of only two reads. The rest of the contigs were made up of three to eight reads, except four of them that were made up of a higher read number, ie, 9, 11, 12 and 28. This shows that our library demonstrated a low level of sequence redundancy.
Functional Characterization with BLAST2GO
We found 440 ESTs with homology to GO accessions, which allowed them to be grouped into three functional categories: those related to
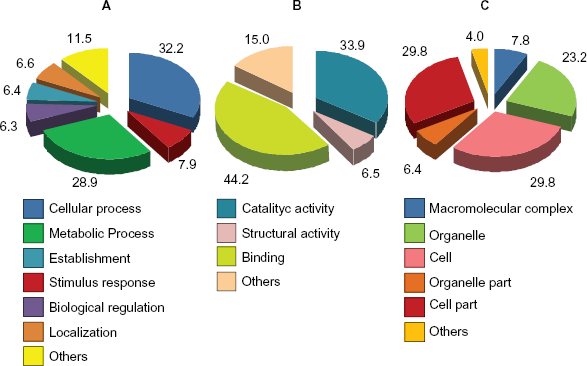
KEGG Pathway Annotations
Annotation using BLASTX and KEGG allowed the classification of ESTs in agreement with their function in the context of specific metabolic pathways.
We found homologous ESTs for most enzymes involved in the fatty acid biosynthesis (maps 61, 71, Fig. 4), ie, acetyl-CoA carboxylase, 3-oxoacyl-[acyl-carrier-protein] reductase, enoyl reductase, fatty acid synthase (Fig. 4, Table 1 of supplementary materials) and in particular for the enzymes involved in the last steps of oleic, stearic and palmitic acid synthesis (Table 2 of supplementary materials), ie, oleoyl-[acyl-carrier-protein] hydrolase, linoleic acyl-[acylcarrier-protein] (map 61) This was expected since oleic, linoleic and palmitic acids are the three major fatty acids from oil of
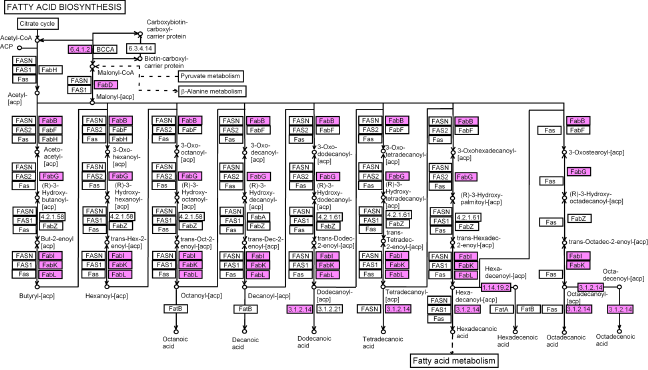
Fatty acids biosynthetic pathway (map 61). The pink boxes are for KEGG ECs that have homologies to
Features of genes and amplicons assessed by quantitative real-time PCR.
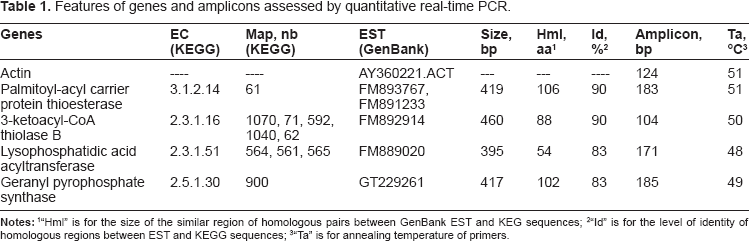
“Hml” is for the size of the similar region of homologous pairs between GenBank EST and KEG sequences
“Id” is for the level of identity of homologous regions between EST and KEGG sequences
“Ta” is for annealing temperature of primers.
In addition to fatty acid, lipid and glycerol pathways, the metabolism pathways of active compounds (terpenes, quinones and alkaloids) and hormones are interesting to consider here. On the one hand, understanding the regulation of active compounds is critical for the control of toxic secondary metabolites such as phorbol. On the other hand, understanding biochemical bases of regulatory mechanisms induced by hormones in the processes of organogenesis, defense and fruit maturation is obviously necessary to improve specific agronomical traits.
When considering the pathway of terpenoid backbone synthesis (map 900), we found representations related to enzymatic function downstream of isopentenyl pyrophosphate, suggesting that the activation of this pathway occurs through geranyl pyrophosphate. Some representations were also found in the biosynthesis of (i) monoterpenoids (map 902) and diterpenoids (map 904) (Table 4 of supplementary materials), (ii) quinones (map 130) and (iii) alkaloids (maps 901, 950) (Table 5 of supplementary materials). The drug metabolism was also found to be active (maps 982, 983) probably in relation to the metabolism of secondary metabolites (Table 6 of supplementary materials).
We found some ESTs putatively involved in auxine biosynthesis, cytokinin and brassinosteroid synthesis (map 905). Other enzymes involved in regulation were from the androgen and estrogen metabolism (map 150) that are known to regulate many processes of organogenesis and plant defenses (Table 7 of supplementary materials).
Assessing Gene Expression
Several ESTs from GenBank showed a high level of homology (>80%) over at least 150 bp with palmitoyl-acyl carrier protein thioesterase, 3-ketoacyl-CoA thiolase B, lysophosphatidic acid acyltransferase and geranyl pyrophosphate synthase genes (Table 1). This strongly suggests that these ESTs are involved in the same function as their homologous from KEGG. These genes are involved in the pathways of
In some cases, EST sequences tagging for one gene were showing nucleotide polymorphism, which is attractive for their use as DNA probe in breeding programs. The level of gene expression detected by quantitative real-time PCR was systematically higher in seeds at the three fruit stages than in leaves (Fig. 5A,B). The maximum of gene expression for geranyl pyrophosphate synthase and lysophosphatidic acid acyltransferase was found in seeds of fruits at stage 1. This was particularly strong for geranyl pyrophosphate synthase whose level o gene expression in seeds of fruits at stage 1 was ~25 times that found in leaves. The level of over-expression of this gene decreased to ~11 in seeds of fruits at stage 2 and to ~6 in seeds of fruits at stage 3. Even if the level of over-expression of lysophosphatidic acid acyltransferase was ~3 in seeds of fruits at stage 1, it was the only gene whose level of expression was lower than that of actin in fruits as well as in leaves. Its level of expression in seeds of fruits at stage 2 and 3 decreased to the same value as that found in leaves. The 3-ketoacyl-CoA thiolase B was over-expressed by a factor ~2 at stages 1 to 2 and ~5 at stage 3. Finally, palmitoyl was over-expressed in fruits by a factor ~5, but the profile of gene expression was flat across the 3 stages.
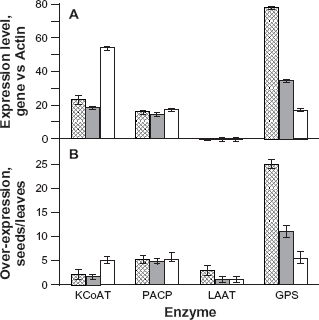
Assessment of the expression profile of four genes by real-time PCR in seeds of
Discussion
As previously mentioned,
The low genetic variability that has been found until now in
SSR could be an alternative source of genetic variability. Using SSRIT, we found that ~23% of our EST sample contains short repeats distributed in tandem di-, tri-, tetra-, penta- and hexa-nucleotides, with the largest amount represented by the di-nucleotides TC(n) and AG(n). The polymorphism associated with these repeats is under investigation (data not shown).
Despite its small size, the metabolic coverage of our EST sample was large at least in the first steps of fruit maturation and characterized by a low level of sequence redundancy. The 5841 non-redundant ESTs of GenBank account for ~20% of the total sample of coding sequences; even if it is still a small proportion, it is enough to start looking for correlations with quantitative trait loci (QTL).
The link between protein function and ESTs that has been established in this study could be questioned since false positives are common to this type of methodology. However, the filter used (Expected ≤0.0001, identity ≥40% and homology size ≥40 aa) is standard when describing the proportions among metabolic activities of a transcriptome at a given stage. Actually, a homologous region larger than 40 amino acids is generally significant when associated to both expected rate lower than 0.0001 and identity larger than 40%. As can be seen from data of supplementary materials, homologies are often larger than 80% similarity. Of course, paralogous genes may always appear as false positives, but would ultimately be eliminated if not associated to a QTL.
An advantage of probes derived from mRNA is that they can be generated from different tissues at various development stages and therefore are highly effective for identifying genes that are differentially expressed during the life-cycle of an organism.40,41
Of course, a method is needed for selecting and mapping candidate loci associated with particular ESTs. Amplified fragment length polymorphism (AFLP) 42 combined with cDNA libraries can be applied to yield highly informative transcript-derived fragments (TDF) for mapping traits whose expression is time-dependent.
In a first step, Suárez et al
25
introduced the sequencing of ESTs from such TDFs and their locations within a genetic map from cassava. Later on, Quin et al
26
showed how to detect AFLP bands from the pattern of restriction of ESTs. This technique offers the advantage of allowing the exploitation of existing EST resources. More simply, SNPs can be investigated by
The use of the techniques just outlined should be effective in assisting breeding programs for the selection of qualitative as well as QTLs. 46 Here, we showed that quantitative real-time PCR allows the detection of genes that are significantly over-expressed in favorable tissues such as is the case of geranyl pyrophosphate synthase that is over-expressed by a factor ~25 in forming seeds and that is, therefore, a good candidate to tag a QTL associated to phorbol synthesis.
The chemical composition of
Fatty acids differ according to three characteristics: (i) the size of carbon chain, (ii) the unsaturation number and (iii) chemical moeities. The larger the size of the carbon chain, the larger the cetane number and the lubricity, but the higher the viscosity and risk of injector choking. On the other hand, the greater the degree of unsaturation, the larger the cetane number, but also the molecule instability and therefore the risk of polymerization and choking. However, unsaturation promotes soot emission and the abatement of soot emission due to oxygen (~10%) naturally present in the biodiesel might be counterbalanced by the rate of alkyl ester unsaturation (double bonds).
54
A large fraction of plants oils have fatty acid compositions similar to that of
Map-based cloning of a diacylglycerol acyltransferase (DGAT) that catalyzes the final step in the glycerol biosynthetic pathway allowed the selection of a new protein variant affecting oil content and composition in maize seeds.
28
QTLs for C16:0, C18:0, C18:1, C18:2, C18:3, C20:1 and C22:1 were also described in rapeseed.
57
There is no reason why such approaches could not be applied to other aspects of this study. However, not all gene functions detected in this work are expected to be useful therefore screening and evaluation will be necessary to guide future studies. However, such screening can be based on some speculation concerning gene function, which is not the case with blinded probes. Among the interesting features of this investigation, we found several ESTs associated with ECs from the biosynthesis pathway of terpenes. An example of this is geranyl pyrophosphate synthase that is a key enzyme upstream the pathway of terpene biosynthesis and that we found over-expressed by a factor ~25 in forming seeds. This over-expression occurs at a fruit stage where terpene precursors are, indeed, expected to form. One may expect that selecting accessions of
Author Contributions
KAG did the cDNA library and cDNA sequencing under the supervision of ASG and JCMC. KAG did the qRT-PCR experiments under the supervision of MAVS. IPL did the oil characterization under the supervision of RSC. NC wrote the project, did the EST processing and managed the project with JCMC. All the authors participated to the project.
Disclosures
This manuscript has been read and approved by all authors. This paper is unique and is not under consideration by any other publication and has not been published elsewhere. The authors and peer reviewers of this paper report no conflicts of interest. The authors confirm that they have permission to reproduce any copyrighted material.
Footnotes
Acknowledgements
K. Gomes is grateful to Fundação de Amparo à Pesquisa do Estado da Bahia (FAPESB) for providing a student fellowship. N. Carels is grateful to Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES) and Fundação Oswaldo Cruz (FIOCRUZ) for providing a research fellowship from the Centro de Desenvolvimento Tecnológico em Saúde (CDTS). This work received financial support from Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq), Brazil (no. 471214/2006-0). We thank Dominique Garcia for helping with plant material and cDNA library preparation as well as Fernanda Amatto Gaiotto for help with microsatellite analysis. We regret the unexpected death of Prof. Julio Cascardo while he was at the top of his career. We are grateful for his continuous dedication to what he thought to be the best for everybody.
Supplementary Tables
ESTs that tag for enzymatic functions putatively involved in hormone biosynthesis in J. curcas.
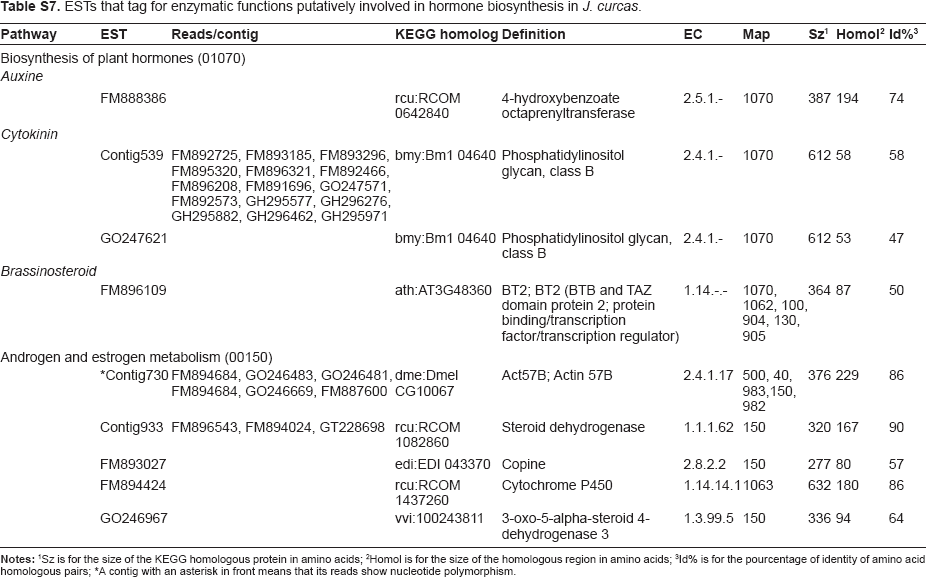
| Pathway | EST | Reads/contig | KEGG homolog | Definition | EC | Map | Sz 1 | Homol 2 | Id% 3 |
|---|---|---|---|---|---|---|---|---|---|
| Biosynthesis of plant hormones (01070) | |||||||||
|
|
|||||||||
| FM888386 | rcu:RCOM 0642840 | 4-hydroxybenzoate octaprenyltransferase | 2.5.1.- | 1070 | 387 | 194 | 74 | ||
|
|
|||||||||
| Contig539 | FM892725, FM893185, FM893296, FM895320, FM896321, FM892466, FM896208, FM891696, G0247571. FM892573, GH295577, GH296276. GH295882, GH296462, GH295971 | bmy:Bm1 04640 | Phosphatidylinositol glycan, class B | 2.4.1.- | 1070 | 612 | 58 | 58 | |
| G0247621 | bmy:Bm1 04640 | Phosphatidylinositol glycan, class B | 2.4.1.- | 1070 | 612 | 53 | 47 | ||
|
|
|||||||||
| FM896109 | ath:AT3G48360 | BT2; BT2 (BTB and TAZ domain protein 2; protein binding/transcription factor/transcription regulator) | 1.14.-.- | 1070, 1062, 100 904, 130, 905 | 364 | 87 | 50 | ||
| Androgen and estrogen metabolism (00150) | |||||||||
| * Contig730 | FM894684, G0246483, G0246481 FM894684, G0246669, FM887600 | ,dme:Dmel CG10067 | Act57B; Actin 57B | 2.4.1.17 | 500, 40, 983, 150, 982 | 376 | 229 | 86 | |
| Contig933 | FM896543, FM894024, GT228698 | rcu:RCOM 1082860 | Steroid dehydrogenase | 1.1.1.62 | 150 | 320 | 167 | 90 | |
| FM893027 | edi:EDI 043370 | Copine | 2.8.2.2 | 150 | 277 | 80 | 57 | ||
| FM894424 | rcu:RCOM 1437260 | Cytochrome P450 | 1.14.14.1 | 1063 | 632 | 180 | 86 | ||
| GO246967 | vvi:100243811 | 3-oxo-5-alpha-steroid 4-dehydrogenase 3 | 1.3.99.5 | 150 | 336 | 94 | 64 | ||
Sz is for the size of the KEGG homologous protein in amino acids
Homol is for the size of the homologous region in amino acids
Id% is for the pourcentage of identity of amino acid homologous pairs
A contig with an asterisk in front means that its reads show nucleotide polymorphism.
