Abstract
BACKGROUND
DNA methylation of the differentially methylated regions (DMRs) of imprinted genes is relevant to neurodevelopment.
METHODS
DNA methylation status of the DMRs of nine imprinted genes in umbilical cord blood leukocytes was analyzed in relation to infant behaviors and temperament (n = 158).
RESULTS
CONCLUSION
While the small sample size limits inference, these pilot data support gene-specific associations between epigenetic differences in regulatory regions of imprinted domains at birth and later infant temperament.
Introduction
Early childhood social–emotional functioning and temperament reflect stable, biologically based individual differences in behavioral tendencies. 1 These behavioral tendencies have been linked to subsequent childhood externalizing, internalizing, and emotional problems.2–5 Prenatal environmental influences, such as smoking, dietary factors, and maternal stress, have all been linked to early childhood social–emotional functioning and temperament.6–8 Many of these environmental influences have also been related to variation in epigenetic processes.9–13 While variation in epigenetic processes is hypothesized as a potential biological mechanism explaining associations between prenatal exposures and early childhood temperament and social–emotional functioning, 14 it is not clear to what extent epigenetic processes relate to these early behavioral tendencies. Understanding the degree to which epigenetic processes relate to these behavioral tendencies is important as it may help inform the etiology of more complex childhood neurodevelopmental and psychiatric outcomes. 15
Epigenetic studies of behavioral and mental health have largely focused on DNA methylation in promoter regions of candidate genes that regulate the hypothalamic–pituitary axis implicated in stress modulation, as well as genes involved in dopaminergic and serotonergic systems that modulate reward and emotion.16–18 Imprinted genes, however, may also be relevant. Many imprinted genes are highly expressed in the brain and are critical in regulating neurodevelopment.19–27 The parent-of-origin-dependent manner in which this subset of genes is expressed is tightly regulated through epigenetic processes, including DNA methylation at differentially methylated regions (DMRs), which directly affects the genes' expression.
28
The ways in which imprinted gene regulation might affect brain development could be either by disrupting regulation of nutrient acquisition, hormones, or fetal growth, or, more directly, by influencing neuronal growth and pruning, or axonal sprouting and interconnections.
29
DNA methylation and altered expression of imprinted genes have been associated with clinical neurodevelopmental disorders in children, such as the Rett, Angelman, and Beckwith-Wiedemann syndromes, 27 but studies of quantitative behavioral phenotypes are limited. Some exceptions are recent studies of neurological functioning in newborns. In an initial study, researchers demonstrated that altered expression of 22 imprinted genes in the placenta was correlated with neurological outcomes shortly after birth (ie, prior to hospital discharge) using the Neonatal Intensive Care Unit Network Neurobehavioral Scale (NNNS). 32 In a follow-up study with this same cohort, this same group found that higher mean DNA methylation of the serotonin receptor 2A (HTR2A), which may also be subject to genomic imprinting, 33 was associated with a less desirable score on the NNNS measurement of infant quality of movement and a more desirable score on measurement of infant attention. 34
Identifying the extent to which DNA methylation is related to early childhood temperament and social–emotional functioning could point to potential biological markers useful in risk assessment or as targets for interventions that aim to ameliorate exposure-induced methylation changes. Considering previous findings linking neurological outcomes shortly after birth to imprinted gene expression, as well as the relevance of imprinted genes to prenatal growth and brain development, we investigated the associations between early childhood temperament and the DNA methylation status of umbilical cord blood leukocytes with reference to the nine DMRs in the imprinted genes that are important for embryonic growth regulation (
Methods
Cohort and study sample
Participants were from a population-based birth cohort in the southeastern United States (the Newborn Epigenetics STudy [NEST]). 37 This research complied with the principles of the Declaration of Helsinki. All participants provided informed consent and the study was approved by Duke University's institutional review board.13,37 Pregnant women were recruited from prenatal clinics serving Duke University Hospital and Durham Regional Hospital Obstetrics facilities from April 2005 to June 2011. The analyses focused on participants recruited between 2009 and 2011 when measures of infant temperament were added to the survey and administered at one year after the child's birth. Eligibility criteria were as follows: age ≥18 years, English speaking, pregnant, and intention to use one of the two obstetrics facilities for the index pregnancy to enable collection of umbilical cord blood.
Of the 2,548 women approached, 1,700 (66.6%) were enrolled. Of the 1,700 women, 347 were withdrawn due to miscarriage (n = 117), infant death (n = 3), or refusal for further participation prior to delivery or at the follow-up at age one year (n = 227). Of the remaining 1,353 eligible women, 605 (46%) completed a one-year follow-up survey that included assessment of temperament.
Bisulfite pyrosequencing for determining DNA methylation was conducted using umbilical cord blood samples in the cohort. The analyses were limited to participants with singleton births for whom both DNA methylation had been measured and data on child outcomes between 12 months and 24 months had been reported. This ranged from 158 to 198 individuals depending on the particular DMR being evaluated.
The analysis cohort differed from the enrollment cohort with respect to educational attainment
Measures
The Infant–Toddler Social–Emotional Adjustment (ITSEA) questionnaire was used to assess infant social–emotional behavior and temperament. 38 The ITSEA instrument is a parent report questionnaire that measures four internalizing domains (General Anxiety, Separation Distress, Depression–Withdrawal, and Inhibition to Novelty) and three externalizing domains (Peer Aggression, Aggression–Defiance, and Activity–Impulsivity). All questions are rated on a three-point scale: zero = not true or rarely, one = somewhat true or sometimes, and two = very true or often. Scales of the ITSEA questionnaire have demonstrated acceptable test–retest reliability and interrater reliability. 39 Cronbach's alpha coefficients for the internalizing and externalizing domains were 0.80 and 0.86, respectively.
The very short form of the Early Childhood Behavior Questionnaire (ECBQ) 40 was used to assess three broad factors of temperament related to reactivity and self-regulation, including negative affectivity, effortful control, and surgency. The ECBQ has been shown to have good test–retest reliability, is internally consistent, and demonstrates satisfactory interrater agreement. 40 Indicators such as low frustration tolerance, sadness, fearfulness, and low soothability mark the negative affectivity factor. The effortful control factor is marked by indicators such as attention, inhibitory control, and low-intensity pleasure. The surgency factor is marked by indicators such as impulsivity, high activity level, and high-intensity pleasure. Cronbach's alpha coefficients for negative affectivity, effortful control, and surgency were 0.75, 0.75, and 0.80, respectively.
Aother variables
At enrollment (median gestation age: ∼12 weeks), a self- or interviewer-administered questionnaire solicited information about maternal age, race, marital status, educational status, psychiatric history (ever diagnosed or treated for depression or anxiety), parity, and smoking history (whether they smoked prior to or during pregnancy). Birth weight, gestational weeks, and infant gender were obtained from medical records.
DNA methylation
Umbilical cord blood was collected via umbilical vein puncture into ethylenediaminetetraacetic acid-containing vacutainer tubes. Genomic DNA (500–800 ng) was treated with sodium bisulfite using the EZ DNA Methylation Kit as per the manufacturer's instructions (Zymo Research). Bisulfite-converted DNA (∼40 ng, assuming complete recovery after bisulfite modification) was amplified by polymerase chain reaction (PCR) using the PyroMark PCR Kit (Qiagen). PCR and pyrosequencing primers, genomic coordinates, PCR amplification conditions, and assay validation experiments have been described in detail elsewhere.41–43 Assay validation experiments included duplicate DNA methylation measurements of all DMRs with a methylation profile ± two standard deviations (SDs), as well as gene expression studies in support of the functional significance of the identified methylation marks. Supplementary Table 1 and Supplementary Figure 1 display details of the function and location of the select genes.
Characteristics of the samples.

Statistical analyses
Multivariate multiple regression models were used to test the association between methylation status and the five temperament outcomes, controlling for birth weight, gestational weeks, infant age at follow-up, gender, maternal race, maternal education level, smoking status during pregnancy, maternal age at delivery, parity, and maternal reported history of anxiety or depression, as these factors have been associated with DNA methylation at this or other genomic regions and infant temperament. Analyses utilized the mean methylation value for each of the nine DMRs in nine separate models to predict the response measures for the five temperament domains: two primary domains of the ITSEA (internalizing and externalizing) and the three factors of the ECBQ (negative affectivity, effortful control, and surgency). Mean methylation values less than or greater than three SDs s from the mean were treated as outliers and removed. Cronbach's alphas for individual CpGs within each DMRs were all ≥0.88. Supplementary Table 2 displays the mean methylation values for the nine DMRs in the analysis sample in relation to the sample excluded from analyses because of lack of temperament data. Potential moderation by gender was examined by including an interaction term (gender × DMR) in the models. Secondary analyses examined each subdomain of the behavioral or temperament measures when a significant association between a DMR and one of the temperament outcomes was observed. This was performed to determine the subdomain of temperament likely to contribute to the association observed in the primary analyses. To facilitate interpretation of effect size, standardized regression coefficients are presented.
Results
The distribution of the study sample characteristics is presented in Table 1. Mean maternal age at delivery was 29 years; 44% of the study sample had a college degree or higher; non-Hispanic Blacks, non-Hispanic whites, and Hispanics comprised 27%, 39%, and 30%, respectively. Table 2 displays means and SDs for the ITSEA summary and subscales and ECBQ scale.
Mean values and standard deviations (SDs) for the ITSEA summary scales and subscales, as well as ECBQ summary scales.

We evaluated the association between DNA methylation at the regulatory DMRs for nine imprinted genes. The regression residuals for the nine models were normally distributed. Results of multivariate multiple regression indicated significant associations between ITSEA and ECBQ domains and the DRMs regulating
The results of the regression analyses (Table 3) further indicated significant associations between
The interaction term (gender × DMR) for gender and
The relation between the
Results of multivariate multiple regression for the effects of DNA methylation on infant temperament.

In three separate regression models, maternal self-reported smoking was not significantly associated with
Discussion
In this study, we found that (1) higher DNA methylation at the intragenic
Investigating epigenetic processes, especially in DMRs of imprinted genes, may offer insights into the genesis of childhood temperament and social–emotional development.
44
Epigenetic regulation of imprinted genes has been associated with clinical neurodevelopmental disorders,
20
but to our knowledge, this is the first study to show a relationship between prenatal epigenetic regulation of imprinted genes and infant temperament. Using a different methodological approach and developmental stage (newborn functioning), others have shown that altered placental tissue DNA methylation in a cluster of imprinted genes, including
DNA methylation in the DMRs of imprinted genes could serve as biosensors reflecting a response to a range of environmental exposures.
45
In these data, we did not find that maternal self-reported smoking was significantly related to any of the DRMs that were also related to neurobehavioral outcomes. However, in an epigenome-wide association study, prenatal exposure to tobacco smoke, which is linked to neurodevelopmental differences in attention and impulsivity in children, has been found to be related to methylation of regions in the
Notable findings in this study were the relationships observed between temperament domains and DNA methylation of
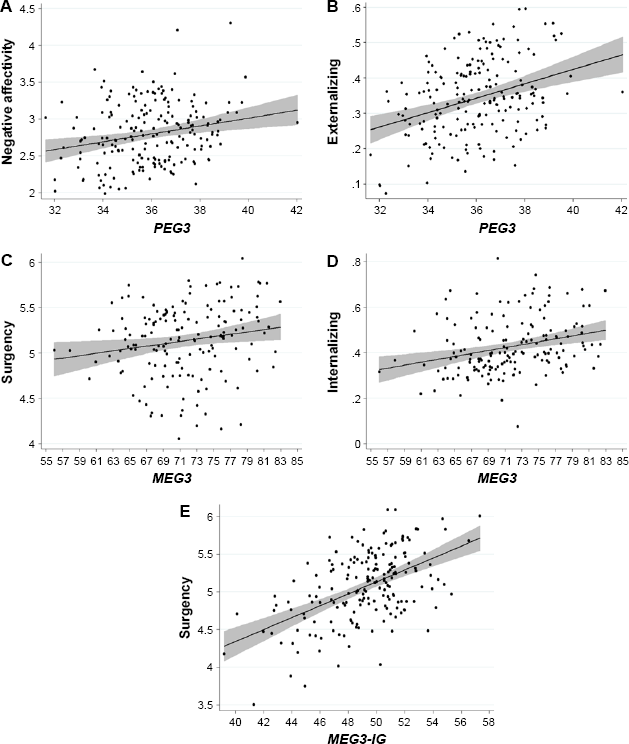
Fitted regression lines and 95% confidence intervals (shaded) for relationship between
It is widely known that for most regions, DNA methylation is a tissue- and cell-specific process. Because studies involving otherwise-healthy humans must assess DNA methylation using accessible biological specimens, such as peripheral blood leukocytes, as a surrogate for the brain, mechanistic links between DNA methylation and neurobehavioral outcomes cannot be inferred with certainty. Nonetheless, recent studies do suggest that there are significant correlations in DNA methylation at some imprinted genes between peripheral blood leukocytes and fetal brain tissue. 41 Further, methylation marks are shown to be stable over time at some genomic regions. 41 Thus, DNA methylation from peripheral tissue may have utility in epidemiologic studies of cognitive and neurobehavioral phenotypes in children, but results from such studies should be interpreted within the context of these caveats. 45
It is also important to note that the overall effect of the association between DNA methylation and temperament outcomes in this study was small. The significant standardized regression coefficients for the multivariate models ranged from 0.15 to 0.28, which indicate that these accounted for a small proportion of the variance in temperament domains (∼0.02%–0.08%). Notably, however, small effect sizes are often observed in epigenetic studies in otherwise-healthy human populations, and the effect sizes observed in these data are comparable to those reported in other studies.60–62
A limitation of this study was the use of parent report to assess childhood outcomes. Although, this is standard practice in clinical assessment, future studies may benefit from the inclusion of direct observation. Another limitation is the modest sample size. Replication in larger samples is warranted to more definitively confirm these findings. Finally, we tested a multivariate response set of temperament and social–emotional functioning in relation to mean DNA methylation in nine gene DMRs and the likelihood of finding a significant association may be inflated by multiple testing. Correction for multiple testing would render the effects we observed here nonsignificant, with the exception of the association between
In sum, these early data support gene-specific associations between epigenetic differences in the regulatory regions of two imprinted domains at birth and later infant temperament, further justifying follow-up work on the role of imprinted genes in neurodevelopmental outcomes in children. Investigating imprinted genes in relation to neurodevelopmental outcomes may be a useful approach to understanding the role of DNA methylation on neurodevelopment because it is known that this set of genes is tightly regulated through epigenetic mechanisms, and environmentally induced effects can have a significant impact on gene expression. Further, identifying epigenetic marks associated with temperament could be useful in risk assessment, thus providing a window of opportunity for prevention. Epigenetic signatures are potentially reversible. Therefore, it may also be possible to discover ways to correct DNA methylation profiles, restore gene function, and optimize neurodevelopmental growth. Although such strategies are possible, they require significant advancement in our understanding of the epigenetic programming involved in children's neurodevelopment.
Author Contributions
Conceived the study: BFF, CH, and SKM. Designed the original cohort from which the follow-up data were collected: CH, JMS, SKM, and APM. Directed the methods for recruitment and collection of cord blood samples: APM. Directed the DNA methylation assays and experiments: SKM. Prepared data sets and performed preliminary analyses: C-TL, ZH, and ESI. Performed final analyses and led the writing of the paper with input from RLJ, CH, AS, and SKM: BFF. All authors reviewed and approved of the final manuscript.
Footnotes
Supplementary table 1
Chromosomal location, expressed allele, methylated allele, and relevant function for imprinted genes included in this study.
Acknowledgments
We would like to thank the women and children in the Newborn Epigenetics STudy (NEST) cohort for their participation in this study. The contents are solely the responsibility of the authors and do not necessarily represent the official views of the National Institutes of Health or the US Environmental Protection Agency (USEPA). Further, USEPA does not endorse the purchase of any commercial products or services mentioned in the publication.
