Abstract
Rapid alkalinization factors (RALFs) are plant small peptides that could induce a rapid pH increase in the medium of plant cell suspension culture and play a critical role in plant development. The evolutionary process of the
Introduction
Peptide signaling is important for cell-to-cell communication and participates in a variety of developmental processes and environmental responses. A number of genes encoding small-secreted peptides have been identified in plants and a certain proportion of them are hormones.
1
These peptides play critical role in all aspects of the plant life cycle and have diverse functions. Such as, CLV3 (CLAV-ATA3) peptide regulates meristem size.
2
Peptide systemin induces the systemic defense response.
3
ENOD40 encodes two small peptides, both of which can affect the normal nodule development.
4
Defensins are involved in the innate immune system of plants.
5
PSK (phytosulfokine) peptide has been demonstrated to promote cellular proliferation and transdifferentiation.6,7 SCR peptide is the pollen self-incompatibility recognition factor in the
RALFs are a new type of plant peptide hormones that participate in diverse biological processes. Such as, activation of protein kinases, inhibition of root growth and development,18,21,27 regulation of fruit maturation,
20
nodule formation,
22
tissue expansion
10
and pollen development,24,26 and so on. Interestingly, the number of
Materials and Methods
Sequence identification and conserved motif analysis of RALF genes
To identify potential members of the
Alignment and phylogenetic analysis
To generate the alignment of the 91 RALF proteins from the
Estimation of the maximum number of gained and lost RALFs
To determine the degrees of gene family expansion in the analyzed plant lineages, we divided the phylogeny into ancestral clades (those containing at least one representative of monocots and eudicots), recent clades (monocot specific or eudicot specific) and species-specific clades. Nodes basal to the split among lineages denoted the most recent common ancestor (MRCA) and were labeled as N0 to N4. Gene duplication and loss events were inferred by reconciling the gene tree for each cluster/subcluster with the species tree using Notung v2.5. 36
Divergence levels analysis
To analyze positive or negative selection of the
Inference of duplication time
Pairwise alignment of nucleotide sequences of the
Codon bias analysis
Codon bias can reflect the degree of selective constraint in a gene. To measure the extent of codon bias, effective number of codons (ENC) and codon bias index (CBI) were estimated using DnaSP v.5.10.01. 43 The ENC values range from 20 to 61, meaning from the maximum codon bias (only one codon is used for each amino acid) to no codon bias (all synonymous codons for each amino acid are equally used). 44 The CBI values range from 0 to 1, meaning from uniform use of synonymous codons to maximum codon bias. 45 We also estimated some parameters related to codon bias, such as GC1,2 (the GC content at the first and second codon positions), GC3 (the GC content at the third codon positions) using DnaSP v.5.10.01. 43
Correlation analysis of expression data and protein subcellular localization
Expression profiling can provide useful clues to gene function. To examine the expression patterns of the
Results and Discussion
Identification, motif organization and phylogenetic analyses of the RALF genes
Number and motif structure of

Phylogenetic analyses can allow us to identify evolutionarily conservative and divergent of gene family. To achieve this goal, phylogenetic analyses of the 91
Contrasting changes in the numbers of RALF genes
To better understand how Evolutionary change in the number of 
Evolutionary patterns of RALF gene family
It has been suggested that the Evolution of the one subgroup of 
While segmental duplications were not the major factors that led to the expansion of the
Next, we also investigated the distributions of the unstable and stable Chromosomal locations of 
In addition, when distantly related species compared, the newly added genes tended to form species-specific clusters or sub-clusters in the eudicots. For example, seven
The phylogenetic tree topology revealed several pairs of
Since codon bias can provides some examples of weak selection at the molecular level. Moreover, several researches have verified that selection on synonymous sites is correlated with stability of mRNA secondary structure, translation efficiency and accuracy, ribosome traffic and protein folding.63–65 We also verified the codon usage bias of
Different signatures of selection in RALFs
To examine whether Divergence levels of 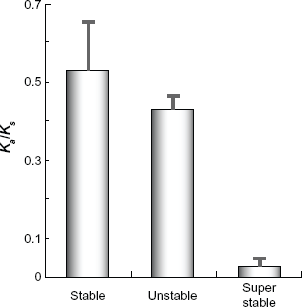
Different expression profiles of the RALFs in Arabidopsis
We also examined the expression patterns of the Arabidopsis Expression profiles of the 
Conclusion
This study explored the evolutionary process of
Author Contributions
Conceived and designed the experiments: JC. Analysed the data: FS, JC. Wrote the first draft of the manuscript: JC. Contributed to the writing of the manuscript: FS. Agree with manuscript: FS. Jointly developed the structure and arguments for the paper: JC. Made critical revisions and approved final version: JC, FS. All authors reviewed and approved of the final manuscript.
Funding
This project is supported by grants from the National Science Foundation of China (No. 31100923), the National Science Foundation of Jiangsu Province (BK2011467) and Jiangsu University Senior Personnel Research Grants (10JDG027) to JC.
Competing Interests
Authors disclose no potential conflicts of interest.
Footnotes
Supplementary Data
Likelihood values and parameter estimates for the

| Gene branches | Model | Log-likelihood | Positive selection sites | |
|---|---|---|---|---|
| Group I | M8 | 0.5206 | -4162.59 | Not found |
| M8a | 0.4827 | -4163.25 | Not found | |
| M7 | 0.5138 | -4162.07 | Not found | |
| M5 | 0.5723 | -4166.45 | 35,66,69,77,79,90,106 | |
| MEC | 0.6459 | -4096.26 | 39,43,66,69,77,79, | |
| Group II | M8 | 0.6145 | -1122.75 | 7,13,14,19,20,23,24,25,38,49,50,52,53,57,70,73,76 |
| M8a | 0.4435 | -1123.65 | Not found | |
| M7 | 0.4398 | -1123.63 | Not found | |
| M5 | 0.4846 | -1124.25 | 7,52,57,76 | |
| MEC | 0.7122 | -1115.32 | 4,7,13,14,20,23,24,25,30,38,49,52,53,57,61,73,76 | |
| Group III | M8 | 0.6031 | -1137.14 | 4,27,28,32,34,51,56,64,72,74,78 |
| M8a | 0.4139 | -1136.04 | Not found | |
| M7 | 0.4483 | -1136.32 | Not found | |
| M5 | 0.4737 | -1138 | 64,78 | |
| MEC | 0.6697 | -1131.68 | 4,10,25,27,28,32,34,36,51,54,55,56,58,64,72,74,77,78 | |
| Group IV | M8 | 0.4432 | -802.603 | Not found |
| M8a | 0.4913 | -803.102 | Not found | |
| M7 | 0.4271 | -802.584 | Not found | |
| M5 | 0.4917 | -803.795 | Not found | |
| MEC | 0.5384 | -797.179 | 48,58,60,68,72 | |
| Group V | M8 | 0.372 | -1587.53 | Not found |
| M8a | 0.4232 | -1588.03 | Not found | |
| M7 | 0.3795 | -1587.36 | Not found | |
| M5 | 0.4154 | -1587.73 | 8, | |
| MEC | 0.6176 | -1580.61 | 3,5,6,8,14,17,19,21,22,24,35,38,43,45,54,60,67,72,76, 78,86,88,93,101,111,121 | |
| Group VII | M8 | 0.4258 | -7178.9 | Not found |
| M8a | 0.3071 | -7200.06 | Not found | |
| M7 | – | – | – | |
| M5 | 0.3663 | -7211.88 | Not found | |
| MEC | 0.4086 | -7047.72 | 68,71,75,77, | |
| Group VIII | M8 | 0.4547 | -1471.73 | 3,14,15,17,18,30,31,36,48,52,55,61,109,112,123 |
| M8a | 0.2244 | -1466.05 | Not found | |
| M7 | 0.2281 | -1466.04 | Not found | |
| M5 | 0.2781 | -1467.72 | Not found | |
| MEC | 0.4315 | -1472.78 | 16,17,36,52,62,112 | |
| Group IX | M8 | 0.3449 | -2778.46 | Not found |
| M8a | 0.342 | -2776.4 | Not found | |
| M7 | 0.3533 | -2778.6 | Not found | |
| M5 | 0.3557 | -2779.98 | Not found | |
| MEC | 0.4434 | -2733.01 | 25,31,37,38,39,40,43,44,81,84 | |
| Group X | M8 | 0.3125 | -3496.94 | Not found |
| M8a | 0.2881 | -3500.02 | Not found | |
| M7 | 0.3155 | -3500.78 | Not found | |
| M5 | 0.3206 | -3504.9 | Not found | |
| MEC | 0.3524 | -3441.27 | 7,9,10,23,25 |
As a requirement of publication author(s) have provided to the publisher signed confirmation of compliance with legal and ethical obligations including but not limited to the following: authorship and contributorship, conflicts of interest, privacy and confidentiality and (where applicable) protection of human and animal research subjects. The authors have read and confirmed their agreement with the ICMJE authorship and conflict of interest criteria. The authors have also confirmed that this article is unique and not under consideration or published in any other publication, and that they have permission from rights holders to reproduce any copyrighted material. Any disclosures are made in this section. The external blind peer reviewers report no conflicts of interest.
