Abstract
Robust stabilization and environmental disturbance attenuation are ubiquitous systematic properties that are observed in biological systems at many different levels. The underlying principles for robust stabilization and environmental disturbance attenuation are universal to both complex biological systems and sophisticated engineering systems. In many biological networks, network robustness should be large enough to confer: intrinsic robustness for tolerating intrinsic parameter fluctuations; genetic robustness for buffering genetic variations; and environmental robustness for resisting environmental disturbances. Network robustness is needed so phenotype stability of biological network can be maintained, guaranteeing phenotype robustness. Synthetic biology is foreseen to have important applications in biotechnology and medicine; it is expected to contribute significantly to a better understanding of functioning of complex biological systems. This paper presents a unifying mathematical framework for investigating the principles of both robust stabilization and environmental disturbance attenuation for synthetic gene networks in synthetic biology. Further, from the unifying mathematical framework, we found that the phenotype robustness criterion for synthetic gene networks is the following: if intrinsic robustness + genetic robustness + environmental robustness ≤ network robustness, then the phenotype robustness can be maintained in spite of intrinsic parameter fluctuations, genetic variations, and environmental disturbances. Therefore, the trade-offs between intrinsic robustness, genetic robustness, environmental robustness, and network robustness in synthetic biology can also be investigated through corresponding phenotype robustness criteria from the systematic point of view. Finally, a robust synthetic design that involves network evolution algorithms with desired behavior under intrinsic parameter fluctuations, genetic variations, and environmental disturbances, is also proposed, together with a simulation example.
Introduction
Synthetic biology is the design and construction of biological devices and systems for useful purposes. 1 Such purposes include a focus on finding how the synthetic gene or man-made DNA works in a living cell, how to use the genetic engineering to build new biological systems, and rebuilding natural systems as platforms to benefit human society. Real physical and biological systems inherently suffer from intrinsic perturbations and external disturbances. In the last two decades, robust stabilization and noise-filtering theories have been developed by control engineers both in order to achieve the robust stability needed for tolerating intrinsic perturbations and in order to obtain the ability to noise filter against external disturbance; these would in turn improve the reliability and performance of control systems.2–6 Their applications are also extended from control systems2–5 to signal processing systems, 6 communication systems, 7 and biological systems.8,9 Since both engineered systems and biological systems need robust stabilization and noise-filtering abilities to tolerate intrinsic perturbations and resist external disturbances (or noises) so as to maintain their desired function or performance, there should exist some common schemes for robust stabilization and disturbance attenuation for these two kinds of systems. In previous studies,10,11 network robustness, genetic robustness, intrinsic robustness, and environmental robustness–-and their trade-off on phenotype robustness–-have been discussed in gene regulatory networks, evolutionary networks, and ecological networks. Since synthetic biology combines engineering methodology and biological technology in order to design artificial biological systems in host cells–-which suffer from intrinsic parameter fluctuations, genetic mutations, and environmental disturbances in vivo–-the network robustness and phenotype robustness criteria of synthetic biology are to be discussed in this study.
In general, there are several methods that have been proposed to design or construct a gene circuit for synthetic biology; these involve (1) DNA modification, (2) transcription regulation, (3) post-transcriptional regulation, and (4) post-translational regulation. 12 The goal is to create biological modules that can perform several different basic functions, including the design and engineering of new biological functions and systems by BioBrick-assembled gene circuits, which have been widely used in synthetic biology.13,14 These gene circuits are reengineered with different promoters, ribosome binding sites (RBS), protein or RNA regions, and transcriptional terminators. Despite recent efforts targeting engineering at the genome level, most synthetic biology constructs are engineered at the level of genetic circuits encoded on plasmids, from a very simple device like a repressilator 15 to the more complicated devices like a counter 16 or a switch.12,17 This relatively new field has provided insights into the mechanisms of natural gene networks, 18 engineered multicellular pattern formations, 19 bacterial photography, 20 tumor-target bacterial, 21 feed-forward network-based concentration sensors, 22 robust and tunable oscillators, 23 genetic networks that count, 16 genetic sequential logic circuit such as Push-on Push-off switch, 17 and the more complicated synchronous toggle switch. 12 Furthermore, entire metabolic pathways have been engineered on the genome level to produce an anti-malaria compound, 24 biofuels from plant biomass,25,26 lycopene through automated genome engineering and accelerated evolution, 27 and a synthetic chromosome 28 translated into a host bacterium. 29
However, a wide variety of loss-of-function mutations or genetic variations are observed in BioBrick-assembled genetic circuits, including point mutations, inversions, and deletions. 13 One problem with engineered genetic circuits in synthetic biology is their robust stability under genetic variations and environmental disturbances. 30 Therefore, how to design a robust synthetic gene network that can tolerate genetic variations due to point mutations, inversions or deletions, resist environmental disturbances, and also function properly in a host cell, is an important topic of synthetic biology.13,31–39 In systems biology, a core mechanism has been found showing that there exists an interplay between system disorder and robustness of gene networks. 40 Additionally, in the evolutionary process there exists some interplay in evolutionary gene networks between the ability to evolve and network robustness. 41 According to our previous studies, we have found that there exists a unifying mathematical framework in systems, evolutionary, and ecological networks.10,11,41 Here, we would like to propose a robust gene network design which can ensure the specified network function (phenotype robustness) according to our previous works to see if there still exists a similar mathematical framework in synthetic biology. This design also provides an insight into how to design a robust synthetic gene network based on phenotype robustness criteria. In synthetic biology, it does not matter whether gene circuits are simple devices such as a repressilator or complicated devices such as a toggle switch, the dynamics of these biological systems are represented by ordinary differential equations (ODEs) and thus can be analyzed using our proposed phenotype robustness theory from the system perspective.
Recently, Kuepter et al
42
have developed an approach based on semi-definite programming that identifies a consistent steady state for polynomial equations and inequality constraints this allows one to certify the feasible and infeasible regions of parameter space. In synthetic biology, ‘feasible region’ means the parameter set for a desired steady state in a synthetic gene network; the design of synthetic gene networks is difficult because of the uncertainties in molecular concentrations and parameter values. Batt and his colleague
43
provide an approach to develop efficient algorithms for solving robustness analyses and tuning problems. This approach was implemented in a publicly available tool RoVerGeNe; its practical applicability has been demonstrated on a synthetic transcriptional cascade in
In general, synthetic biologists always had to select a set of design parameters using conventional trial-and-error methods to achieve the desired behavior of a synthetic gene network. However, when the scale of a synthetic gene network becomes large and the desired behavior becomes complex, how to select a set of adequate design parameters becomes an important topic for synthetic biology. Natural genetic circuits have evolved and have been optimized for their functions by means of natural selection, while at the same time being robust enough to handle genetic variation and environmental changes. Can we use the same principle to select an adequate parameter set for engineering a synthetic gene circuit? Genetic algorithm (GA) based design methods have been employed to mimic natural selection in the evolution process and to select adequate kinetic parameters for designing robust genetic oscillators with prescribed amplitude, period, and phase, under genetic variations and environmental disturbances. 47 GA is a stochastic optimization algorithm, which was inspired by the mechanisms of natural selection and evolution genetics 48 to optimally track a desired target. The proposed GA based design method can search for design parameters from feasible ranges to achieve fitness maximization, which is equivalent to the optimal robust tracking of a desired oscillation behavior under the effect of intrinsic and extrinsic perturbations on the host cell. The robust tracking problem is equivalent to the robust stabilization problem for the desired oscillatory behavior, whose tracking error is inversely proportional to the fitness function of synthetic oscillator to the oscillation, ie, the minimum tracking error is equivalent to the maximum fitness function. Therefore, a GA based design method is to efficiently search for an adequate set of design kinetic parameters in order to maximize the fitness function and achieve the optimal (minimum error) tracking of the desired oscillation behavior, which is done by mimicking natural selection in the evolutionary process. 47
Based on previous studies,10,11 the framework of phenotype robustness criteria has been constructed for systems, evolutionary, and ecological biology. We found that if network robustness can confer intrinsic robustness for tolerating intrinsic parameter fluctuations, genetic robustness for buffering genetic variations and environmental robustness for resisting environmental stimuli, then the phenotypes of these biological networks will be maintained under intrinsic parameter fluctuations, genetic variations, and environmental stimuli. Moreover, the trade-off between intrinsic robustness, genetic robustness, environmental robustness, and network robustness can be revealed by the phenotype robustness criteria for biological networks at different levels from gene regulatory networks, evolutionary networks, and ecological networks. In the following sections, intrinsic robustness, genetic robustness, environmental robustness, network robustness, and their trade-off on phenotype robustness in systems, evolutionary, and ecological networks are to be extended to synthetic biology in sequence.
Trade-off between Genetic Robustness, Environmental Robustness and Network Robustness in Synthetic Biology
Network robustness analysis of synthetic gene network
Synthetic biology can be seen as the application of engineering to biological molecular systems, with a particular purpose. By transformative innovation, it is possible to build living machines from off-the-shelf chemical parts using similar strategies to what electrical engineers employ to design computer chips.15,18,19 The ultimate goal of synthetic biology is to design and construct biological systems with desired behaviors.8,49 Drawing upon the automated synthesis of DNA molecules and their assembly into genes and microbial genomes, it is predictable for the more efficient redesign of natural biological systems as well as the construction of functional “genetic circuit” and metabolic pathways for practical purposes.16,20–29 Synthetic biology is foreseen to have important applications in biotechnology and medicine, as well as in revolutionizing how we conceptualize and approach the engineering of biological systems.
33
However, although the construction of gene regulatory networks has demonstrated the feasibility of synthetic biology,13,30–34 the design of gene networks is still a difficult problem and the most newly designed gene networks cannot function properly.
15
These design failures are due mainly to both intrinsic perturbations–-such as intrinsic parameter fluctuations due to random molecular fluctuations (gene expression noise), genetic variations due to mutation, splicing, evolution from the fast metagenesis of host cell (
For gene network design in synthetic biology, we first architect a gene circuit system for some functions, then specify the gene components and synthesize these gene components into a gene circuit, and finally send the engineered gene network into the host cell
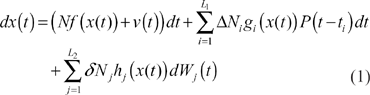
where the state vector
The design purpose of the synthetic gene network in (1) is to specify the kinetic parameters in the stoichiometric matrix 
In order to achieve the desired steady state 

Proposition 1
If some design kinetic parameters and decay rates in the stoichiometric matrix 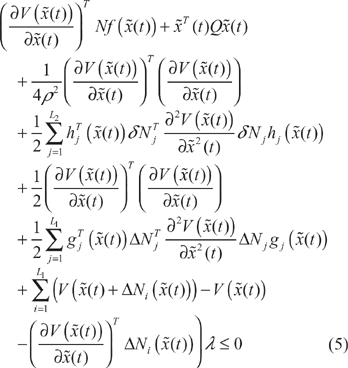
Proof
For the complete proof, see Appendix A. By considering the case of the corresponding uncertainty, the phenotype robustness criterion of the synthetic gene network in (5) can also be rewritten as follows:

Remarks
Previous robust synthetic designs45,50,51 considered only random molecular fluctuations. In Proposition 1, both random molecular fluctuations and discontinuous genetic variations from generation to generation are considered in the design procedure of a synthetic gene network that is intended to mimic the real synthetic gene network in the host cell. This is because yeast and
In general, it is very difficult to specify the stoichiometric matrix 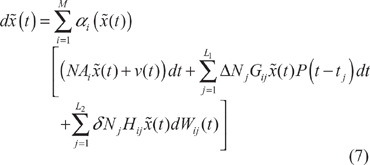
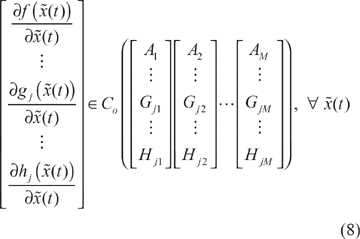
Proposition 2
Assume that some design kinetic parameters and decay rates in the stoichiometric matrix 
Remarks
The disturbance filtering level 
The optimal filtering level (ie, network sensitivity) 
Robust Synthetic Gene Network Design via Network Evolution
In the above robust synthetic gene network design, we need to select a set of design parameters in the stoichiometric matrix

The block diagram of the proposed optimal tracking scheme for a synthetic gene network via network evolution algorithm.
Consider using the nonlinear stochastic gene network in (1) to describe a synthetic gene network in the host cell as follows:
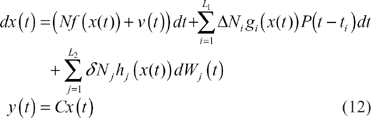
Let us denote tracking errors between the output of a synthetic gene network and the output of some desired systems of interest as follows (see Fig. 1):


In general, it is not easy to directly solve the constrained optimization tracking problem in (14) by using the conventional optimal method and tuning the design parameters in the stoichiometric matrix 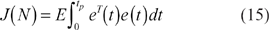

That is, a small mean square tracking error would imply a large fitness and vice versa. If we adapt the kinetic parameters (chromosomes) in the stoichiometric matrix
Therefore, the robust synthetic gene network design with desired output behavior 
The network evolution algorithm is employed to solve the above fitness maximization problem via genetic operators such as selection, crossover, and mutation, and to mimic natural selection in the evolutionary process for tuning the kinetic parameters in the stoichiometric matrix

The flow chart of network evolution algorithm for solving the best fit
Remark
Unlike natural selection, the above intelligent network evolution algorithm can be incorporated with an elitist strategy to enhance the convergence of network evolution algorithms. This strategy copies the best chromosomes from the old population into the next population so as to prevent losing the best solutions in succeeding iterations. It has been shown that the elitist strategy can improve the performance of network evolution algorithm.
The proposed network evolution algorithm is a powerful search algorithm for kinetic parameter selection in synthetic gene network design procedures involving natural genetics. The algorithm evaluates many points in the parameter space of the stoichiometric matrix
Robust Synthetic Gene Network Design via Library-Based Network Evolution
Many standard parts (devices) have recently been constructed for BioBrick,50,51 from which some adequate parts will be selected by synthetic biologists to be physically assembled into a multi-component synthetic gene network. In order to completely implement a transcription unit in a synthetic gene network, one needs the promoter, ribosome binding site (RBS), protein coding region, and a terminator, which can be obtained from the BioBrick. Therefore, several promoter libraries and promoter-RBS libraries have been constructed for the design of synthetic gene networks according to the kinetic activities of promoters and protein expression abilities of promoter-RBS, respectively. In other words, if we select only an adequate set of promoter devices or promoter-RBS devices from the corresponding libraries, then we could engineer a robust synthetic gene network with a desired behavior.50,51
Let us denote promoter libraries and promoter-RBS libraries as
Proposition 3
Assume we select a set of promoter devices or a set of promoter-RBS devices from the corresponding libraries, ie,
Similarly, the optimal disturbance filtering (network sensitivity) design of synthetic gene network in (10) can be changed by the library-based evolutionary search method, as follows:

When the number of promoter devices or promoter-RBS devices of libraries becomes very large, it is not easy to select an adequate set of promoter devices or promoter-RBS devices from the corresponding promoter libraries or promoter-RBS libraries. In this situation, the library-based network evolution algorithm can be used to search a set of promoter devices or a set of promoter-RBS devices from the corresponding libraries, in order to let the synthetic gene network achieve a desired behavior. Therefore, selecting corresponding design parameters from the feasible range of the stoichiometric matrix 
Computer Simulation Example
Here, an in silico design example is introduced to both verify the robust synthetic gene network design and to confirm the robust stabilization and disturbance attenuation performance by the network evolution algorithm, using the GA searching method. Although the synthetic gene circuits may be either simple devices or more complex ones, their dynamic profiles in vivo can be represented by nonlinear stochastic ODEs, which are suitable for the proposed systematic design method. In this example, we want to synthesize a cascade loop of transcriptional inhibitions built in 




Synthetic transcriptional cascade loop in silico design example.
In reality the gene network in the host cell (ie, 
The robust synthetic gene network requires designing the parameters The standard deviations of intrinsic parameter fluctuations and external noises to be tolerated are given as:
The desired steady state The prescribed disturbance filtering (attenuation) level of external disturbances is specified by

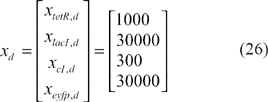
Based on the design procedure, we first shift the desired steady state 
In order to confirm the performance of the proposed robust synthetic gene network, we design the synthetic gene network with a set of kinetic parameters 
In Figure 4, by choosing design parameters in (28) which are within the specified range in (27), the desired steady states (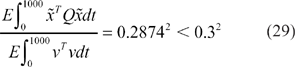
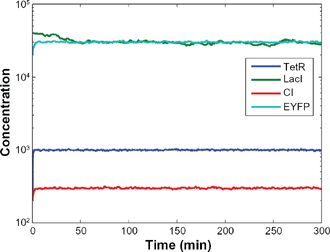
Within the specified parameter range in (27), by choosing the design parameters κ

As evidence to the contrary, if the design parameters are outside the specified range in (27), eg, with (
Clearly, the prescribed disturbance filtering level is achieved by the proposed method. In the simulation stresults, it is obvious that the synthetic gene network applying the proposed design method has both the robust stability to tolerate intrinsic parameter fluctuations and the filtering ability needed to attenuate the external disturbances and achieve the desired steady states. In practical application, if the synthetic gene network has enough robust stability and filtering ability, then the gene network could work with proper functions under intrinsic fluctuations and external molecular noises on the host cell. Furthermore, according to the network evolution algorithm, the design ranges of kinetic parameters and decay rates can easily be solved by fine tuning parameters; this fine tuning can be done via GA-based searching methods in design procedures found in Matlab's LMI Toolbox. Using the proposed design procedure, the synthetic gene network can be guaranteed to meet the three design specifications (i)–(iii) with enough robust stability and noise filtering to achieve the desired steady state (ie, to maintain the proper network function). Here, the primary goal is to confirm that GA-based search methods in the design procedure can be used to efficiently find the suitable system parameters needed for synthetic gene circuit design. Based on the proposed design procedure, the robust synthetic gene circuit can function properly to achieve some steady state (
Discussion
The simulation example in (24) consists of a negative circuit made from the sequence inhibitions of three repressor genes and a fluorescent output; it is designed to reveal the gene expression state in vivo. In general, negative feedback can reduce noises and improve stability,
63
but it can also generate oscillations with a long time delay.15,64 In this example, the kinetic parameters
The study and design of gene networks has become a hot topic in systems and synthetic biology. At the nano-scale, a gene regulatory network inherently experiences random intrinsic parameter fluctuations, genetic variations, and environmental disturbances. A nascent field, synthetic biology, has recently been anticipated to have important applications in biotechnology and medicine and to contribute significantly to better understanding of the functioning of complex biological systems. At present, the development of a gene network under some prescribed design specifications is still difficult; most newly created gene networks are non-functional due to intrinsic parameter fluctuations, genetic variations, and environmental disturbances. How to design a robust synthetic gene network with desired behavior under some specified (or allowable) intrinsic parameter fluctuations, genetic variations, and environmental disturbances in vivo is an important issue for robust synthetic gene networks. This is also the most important feature of a biological network–-to maintain its phenotype in the face of genetic variations and changing environmental conditions in the evolutionary process.
To achieve this consistency, biological networks strike a balance between robustness and evolvability, ie, between resisting and allowing changes in their own internal states. In a series of studies,10,11 we have developed a single unifying mathematical framework for encompassing such diverse examples of stochastic biological networks to discuss intrinsic robustness, genetic robustness, environmental robustness, and network robustness, as well as their trade-off in systems, synthetic, ecological, and evolutionary biology. According to our analyses, the phenotype robustness criteria of stochastic biological networks in systems, synthetic, ecological, and evolutionary biology have a similar mathematical framework. The biological networks in systems, synthetic, ecological, and evolutionary biology can be modeled as nonlinear stochastic systems with intrinsic parameter fluctuations, genetic variations, and environmental disturbances. In these systems intrinsic parameter fluctuations are described by the Wiener (Brownian) process, environmental disturbances are described by the Gaussian white noise, and genetic variations are described by the Poisson point process. Therefore, the interplay of network robustness, intrinsic robustness, genetic robustness, and environmental robustness can be analyzed by the nonlinear stochastic system theory. This interplay can also therefore be analyzed by linear stochastic system theory to get more insight into the network robustness mechanism, when the global linearization technique is employed in interpolating several local linear stochastic systems so as to approximate the nonlinear stochastic system.
From the system theory perspective, the phenotype robustness of nonlinear stochastic gene networks in synthetic biology needs to obey a similar phenotype robustness criterion, ie, “intrinsic robustness + genetic robustness + environmental robustness ≤ network robustness”, which means network robustness needs to be strong enough to tolerate either heritable perturbations (genetic variations) or non-heritable perturbations (ie, random molecular fluctuations and environmental disturbances). This robustness is needed so that the phenotype of gene networks can be maintained in systems, synthetic, and evolutionary biology with a similar mathematical framework. The phenotype robustness of the stochastic gene network is completely consistent with the idea of canalization of development and inheritance of acquired characters reported in previous studies. 68 According to these phenotype robustness criteria, the correlation among intrinsic robustness, genetic robustness, environmental robustness, and network robustness by recent genomic experiments in yeast–-ie, genes conferring similar intrinsic, genetic, and environmental robustness to maintain phenotypic robustness–-can be rationally explained from the systematic perspective. 69 In other words, if the network robustness of gene network is large enough, genetic perturbations or environmental disturbances can then be taken over respectively or simultaneously to maintain the functional phenotype in systems, synthetic, and evolutionary biology.
Rather than desirable features of biological networks, genetic, environmental, and phenotypic random variations are inevitably noisy processes in systems, synthetic and, evolutionary biology.40,70 These stochastic processes arise from the complexity and evolutionary process of biological gene networks. However, there is evidence of high fidelity and minimal noise, including the proof editing of DNA replication and protein translation, in systems and evolutionary biology. Enzymes have evolved toward high specificity. Gene expression is regulated by elaborate mechanisms; random variations seem to have been minimized in systems, synthetic, and evolutionary biology. For instance, gene regulatory networks within bacterial operons may have evolved to minimize genetic and phenotypic noises and infidelity. However, chemophysical constraints on the specificity and fidelity of biological networks are costly so that there is a trade-off between the benefits of fidelity and its cost. Therefore, if biological networks want to retain enough network robustness to confer intrinsic robustness for tolerating intrinsic parameter fluctuations, genetic robustness for buffering genetic variations, and environmental robustness for resisting environmental disturbances, much effort has to be taken and a high cost must be paid.
In general, it is very difficult to solve the HJI in (5) for the phenotype robustness criterion in synthetic gene networks. With the global linearization technique, the HJI problem for robust stabilization of nonlinear stochastic biological networks is reduced to solving an equivalent set of Riccati-like inequalities in (9), looking for the robust stabilization of each local linearized synthetic gene network. We also found that if the network robustness of each local linearized synthetic gene network can take on the local intrinsic robustness, genetic robustness, and environmental robustness of each local linearized synthetic gene network, then the phenotype of the nonlinear stochastic synthetic gene network could also be maintained. Furthermore, from the robust linear system theory point of view, the global linearization method can be used to solve the robust design problem of nonlinear synthetic gene networks, according to the phenotypic robustness criterion in (11).
In synthetic biology, all biological devices have their own system characteristics chosen within their biological allowable range. Thus, to find decisions that satisfy given design constraints and that meet a specific goal at its optimal value (from a huge search space), genetic algorithms (GA) are used to manipulate the problem of searching many parameters simultaneously. A GA has several advantages. It can quickly scan a vast solution set. The GA works by its own internal rules, which means that it doesn't have to know any rules of the problem; this is very useful for complex or loosely defined optimization problems. Due to the parallelism that allows them to implicitly evaluate many schemas at once, GAs are particularly well-suited to solving optimal problems where the space of all potential solutions is truly huge, primarily because they can produce multiple, equally good solutions, for the same optimal problem. However, GAs also have some drawbacks. Although GAs find solutions through evolution, this is their biggest disadvantage. Evolution is inductive and thus it evolves
Conclusion
In this study, based on the nonlinear stochastic synthetic gene network in vivo, we developed a phenotype robustness criterion for synthetic gene networks; we also proposed a robust synthetic gene network design, using network evolution algorithm, to find out adequate system parameters within feasible ranges for the gene circuit design. Because gene circuit design is similar to electronic chip design, some intrinsic parameter fluctuations (eg, genetic mutations) and external disturbances (eg, molecular noises) may cause the synthetic gene circuits to lose function after a single life-time or a number of generations. Based on our previous studies, similar phenotype robustness criterion is also found in synthetic gene networks. Thus, we also proposed a library-based search method using Genetic Algorithms; this method was used to choose suitable biological devices for use in gene circuit designs that ensure functional performance, in spite of adverse effects from random parameter fluctuations, genetic mutations, and external noises.
This series of studies (Part I to Part III) presents a unifying mathematical framework that describes different levels of stochastic biological networks under intrinsic parameter fluctuations, genetic variations, and environmental disturbances. According to the unifying stochastic biological systems, the phenotype robustness criteria of biological networks in systems, synthetic, ecological, and evolutionary biology are also investigated from the robust stabilization and disturbance sensitivity perspective. It was found that if intrinsic robustness + genetic robustness + environmental robustness ≤ network robustness (ie, network robustness can confer intrinsic robustness for tolerating intrinsic parameter fluctuations, genetic robustness for buffering genetic variations, and environmental robustness for resisting the environmental disturbances), then the phenotype will be robust in biological networks at different levels of systems, synthetic, ecological, or evolutionary biology. Using the global linearization method, we also found that if the network robustness of each local linearized system is greater than the total sum of intrinsic robustness, genetic robustness, and environmental robustness of each local linear system, then the phenotype of the biological network is also maintained, regardless of intrinsic parameter fluctuations, genetic variations, and environmental disturbances. Furthermore, network evolution algorithms are also introduced to mimic genetic evolution under natural selection, to search for adequate design parameters from feasible parameter regions, and to select a set of adequate devices from promoter libraries or promoter-RBS libraries for the purpose of engineering robust synthetic gene network; by robust we mean those networks which can perform reliably under intrinsic parameter fluctuations, genetic variations, and environmental disturbances, in vivo, for a long time. The network evolution-based design method may have significant potential for application in the synthetic design of biological networks when synthetic gene networks become more complex.
Footnotes
Author Contributions
Conceived and designed the experiments: BSC. Analysed the data: BSC. Wrote the first draft of the manuscript: BSC. Contributed to the writing of the manuscript: BSC, YPL. Agree with manuscript results and conclusions: BSC. Jointly developed the structure and arguments for the paper: BSC, YPL. Made critical revisions and approved final version: BSC, YPL. All authors reviewed and approved of the final manuscript.
Funding
The work was supported by the National Science Council of Taiwan under grant NSC 100-2745-E-007-001-ASP and NSC 101-2745-E-007-001-ASP.
Competing Interests
Author(s) disclose no potential conflicts of interest.
Disclosures and Ethics
As a requirement of publication author(s) have provided to the publisher signed confirmation of compliance with legal and ethical obligations including but not limited to the following: authorship and contributorship, conflicts of interest, privacy and confidentiality and (where applicable) protection of human and animal research subjects. The authors have read and confirmed their agreement with the ICMJE authorship and conflict of interest criteria. The authors have also confirmed that this article is unique and not under consideration or published in any other publication, and that they have permission from rights holders to reproduce any copyrighted material. Any disclosures are made in this section. The external blind peer reviewers report no conflicts of interest.
