Abstract
Background
MammaPrint was developed as a diagnostic tool to predict risk of breast cancer metastasis using the expression of 70 genes. To better understand the tumor biology assessed by MammaPrint, we interpreted the biological functions of the 70-genes and showed how the genes reflect the six hallmarks of cancer as defined by Hanahan and Weinberg.
Results
We used a bottom-up system biology approach to elucidate how the cellular processes reflected by the 70-genes work together to regulate tumor activities and progression. The biological functions of the genes were analyzed using literature research and several bioinformatics tools. Protein-protein interaction network analyses indicated that the 70-genes form highly interconnected networks and that their expression levels are regulated by key tumorigenesis related genes such as
Conclusion
Genes in the MammaPrint gene signature comprehensively measure the six hallmarks of cancer-related biology. This finding establishes a link between a molecular signature and the underlying molecular mechanisms of tumor cell progression and metastasis.
Background
The MammaPrint assay was developed as a prognostic tool to predict the recurrence risk of breast cancer.1–3 It has been validated in almost 1,600 patients (reviewed in4,5). The 70 genes that make up the MammaPrint signature were selected from genome-wide expression data using a data-driven approach. This means that the genes were selected in an unbiased fashion; there were no predefined assumptions as to whether certain genes were more likely to be involved in the risk of development of distant metastases in patients with early stage breast cancer. This resulted in a set of 70 genes that was able to predict the risk of recurrence with a high sensitivity.
Functional annotation of the human genome has been greatly facilitated by the availability of new functional genetic approaches to study genes.
6
To date, most of the 70 genes in the MammaPrint profile have well-described biological functions. For the remaining genes, we can identify tentative functions and can only speculate about their respective roles in tumor progression and metastasis. It is reasonable to assume that MammaPrint genes whose function is not currently known will, nevertheless, be shown to have important roles in breast cancer biology. One such example is the gene
The fact that gene expression of the primary tumor can predict whether the tumor may metastasize indicates that the metastatic potential is likely hardwired into the tumor cells at a relatively early stage during tumorigenesis and is preserved throughout tumor formation and metastasis development.8–10 This process can be captured in the six steps that are referred to as the “hallmarks of cancer”, which reflect the acquired characteristics of a cancer cell. How and in what order tumor cells acquire these characteristics can differ, but ultimately all cancer cells need these characteristics to successfully metastasize and proliferate. 11
To address if the MammaPrint genes indeed capture these six steps, we examined the biological functions of each of the 70 genes. Traditionally, annotation methods focus only on statistically enriched functional categories in gene signatures. Although these categories can reveal major biological processes, they do not always result in a systematic understanding on how all genes in a molecular signature work together. In this study, we used a bottom-up system biology approach that aims to reveal cellular processes reflected by each single gene in MammaPrint and discover how the 70-genes work together to regulate tumor activities. With this knowledge, we investigated the underlying molecular mechanisms linking their biological function with the molecular biology of breast cancer metastasis.
Results and Discussion
The 70-gene profile covers the six hallmarks of cancer
The six hallmarks of cancer describe the acquired characteristics of a cancer cell that collectively dictate malignant growth and thus reflect the potential of a tumor to metastasize. 11 These six capabilities are shared by most (and perhaps all) types of human cancer. In breast cancer, the 70 MammaPrint genes are predictive of metastasis development, and we set out to ask whether the biological properties of these genes are correlated with the six hallmarks of cancer (as illustrated in Fig. 1):

Depicted is how the genes in 70-gene tumor expression profile are involved in the six well-defined hallmarks of cancer, in tumor progression and metastasis related biological processes, as well as epithelial-mesenchymal transition. Adapted from Cell, 100, Hanahan D, Weinberg RA., The Hallmarks of Cancer, 57–70, Copyright (2000) with permission from Elsevier.
Evading apoptosis
Self-sufficiency in growth signals
Insensitivity to anti-growth signals
Limitless replicative potential
Tissue invasion and metastasis
Sustained angiogenesis
The hallmark avoiding apoptosis confers resistance towards programmed cell death. The major converging point of diverse apoptotic signals that a tumor cell may receive are the mitochondria. Mitochondrial death signals are governed by the BCL2 family of proteins that release cytochrome C and, in turn, activates caspases.
12
Tumor cells can resist apoptosis amongst others by altering expression level of BCL2 or caspases associated proteins. This mechanism is represented by two of the MammaPrint genes (
Biological function of MammaPrint genes and cancer hallmarks. MammaPrint genes are involved in all tumor progression and metastasis-related biological processes, and cover the six well-defined hallmarks of cancer.

The hallmark self-sufficiency in growth signals, refers to tumor cells' reduced dependence on exogenous growth stimulations by generation of their own growth signals. This can be achieved by manipulating the level of growth factors and their receptors or by mutation/altered expression of signal transduction molecules. This characteristic behavior of tumor cells is captured by six growth factor associated genes in the MammaPrint profile (
Equally important, is the hallmark labeled insensitivity to anti-growth signals. This defines the capability of tumor cells to disrupt responses to antiproliferative signaling. A well-studied example is the disruption of growth inhibiting effect of TGFß during tumorigenesis. 13 This pathway is represented by the TGFB3 gene in the MammaPrint profile.
The three hallmarks, evading apoptosis, self-sufficiency in growth signals and insensitivity to anti-growth signals, all lead to growth and proliferation of tumor cells, regardless of the types of exogenous signals received from the tumor microenvironment.
11
Although the biological processes by which normal cells acquire these three capabilities can be quite diverse, the biological features of proliferation and oncogenic transformation are shared among malignant tumor cells (see Fig. 1). These shared characteristic behaviors are captured by 12 proliferation or oncogenic transformation-related genes (
A fourth hallmark of the tumor cell is its limitless replicative potential. Tumor cells bypass two built-in checkpoints that limit the replicative potential of normal cells, the p53 and RB-dependent M1 senescence checkpoint and the telomerase-dependent M2 checkpoint.
14
Fifteen MammaPrint genes are cell cycle genes, and can be assigned to this important feature of tumor cells (
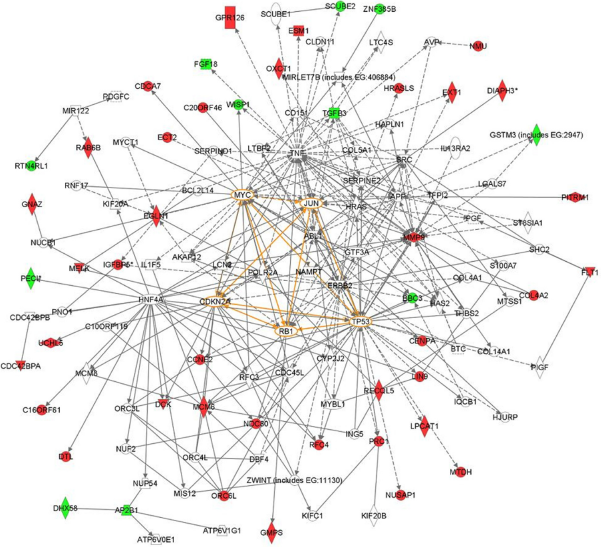
Protein-protein interaction network analyses indicate that the 70 genes form highly interconnected networks centered on known cancer-related transcription regulators such as
The hallmark of tissue invasion and metastasis is a fifth critical step that involves local invasion of the tumor cells into surrounding tissue, escape from the primary tumor site, entry of metastatic tumor cells into the vasculature (intravasation), transportation and survival into the circulation, and arrest and exit of metastatic tumor cells from the vasculature into distant organs (extravasation).
15
During the process of local invasion, tumor cells lose adhesion proteins, remodel extracellular matrix, gain motility by changes in their cytoskeleton and invade adjacent tissue.
11
Five of the MammaPrint genes encode adhesion molecules, extracellular matrix constituents and proteins involved in the breakdown of extracellular matrix (
The last hallmark of cancer is sustained angiogenesis.
17
The survival and growth of tumor cells depends on an adequate supply of oxygen and nutrients through blood vessels and by diffusion through the surrounding tissue. Existing vasculature and passive diffusion are sufficient for oxygen supply to tumors of a limited size. However, aggressive solid tumors often grow to a size that can no longer be sustained by the existing tissue vasculature. Tumor cells enhance their glycolytic capability to ensure an energetically efficient metabolism and proliferation rate under hypoxic conditions. These altered metabolic pathways are preserved by the tumor even when the oxygen concentration is sufficient.
18
Seven genes implicated in this altered metabolism are represented in the MammaPrint profile (
It should be emphasized that the biological processes associated with the six hallmarks such as proliferation, cell-cell adhesion, angiogenesis and invasion are intrinsically linked. That is, a gene known to play a dominant and critical role in one hallmark might also indirectly be involved in other hallmarks. To better understand interactions between the 70 MammaPrint genes and their transcription regulation, we performed protein-protein interaction network analyses. The networks showed that the 70 genes are highly interconnected and center around known cancer-related transcription regulators such as
To summarize, MammaPrint has been developed using a data-driven approach and results in a gene profile that has comprehensive coverage of the six hallmarks of cancer, as well as tumor progression and metastasis related biological processes (Table 1, Fig. 1). In addition, protein-protein interaction network analyses presented here, indicate that the 70-genes form highly interconnected networks and that their expression levels are regulated by key tumorigenesis related genes such as
The biological model of acquisition of metastatic competence through epithelial-mesenchymal transition and the 70-Gene Profile
In the previous section, we have shown that malignancy and metastatic competence of tumor cells at the primary tumor site are measured by the expression level of genes in the 70-gene MammaPrint profile. However, this has not provided an answer as to how tumor cells at the primary tumor site initially acquire their metastatic capability. A biological model that is increasingly gaining acceptance is that tumor cells at the primary site might acquire their metastatic capacity through a process similar to epithelelial-mesenchymal transition (EMT): a key epigenetic program that cells undergo during early embryonic development.
8
During EMT, epithelial cells lose cell adhesion molecules, reorganize their cytoskeleton, gain increased motility and migrate from an epithelial sheet-like structure to an irregular structure of mesenchyme.
19
This change in cellular phenotype is similar to the process that tumor cells undergo to initiate metastasis. Evidence suggests that tumor cells might initiate EMT by turning on or off some of the same transcription factors that are used in early embryonic development.
20
These transcription factors regulate the expression of genes that allow tumor cells to lose adhesion, remodel the surrounding extracellular matrix, acquire enhanced motility to enable cellular migration, resist apoptotic signals, and adapt to an unfamiliar microenvironment at the distant site. The biological model based on the assumption that EMT processes are involved in breast cancer metastasis is consistent with the biological functions of the genes in the MammaPrint 70-gene profile identified here (Table 1). A substantial number (ie, 14 genes) of the 70 gene profile encode for proteins that are known to play an role in early embryonic development and are likely involved in EMT (
Conclusions
The MammaPrint 70-gene tumor expression profile was developed using an unbiased data-driven approach without making assumptions that certain genes are more likely to be involved in tumor metastasis. In this study, we have demonstrated how this approach results in a tumor expression profile that contains many genes actually involved in the six well-defined hallmarks of cancer, in tumor progression and metastasis-related biological processes, as well as in epithelial-mesenchymal transition.
It is equally interesting to investigate which genes are not among the 70 genes in the prognostic signature. Most notably, neither
This study shows how the 70 genes in the MammaPrint gene signature comprehensively measure the six hallmarks of cancer-related biology. This finding establishes a link between a molecular signature and the underlying molecular mechanism of tumor cell progression and metastasis.
Methods
To correlate the probes of the 70 MammaPrint genes 2 with the latest NCBI Human genomic plus transcript database (database updated through 18 March 2009), a BLAST search was performed using the corresponding probes at the NCBI BLAST website (http://blast.ncbi.nlm.nih.gov/Blast.cgi 22 ).
After retrieving the genes, they were translated into protein sequences. The transcribed proteins perform functions through conserved functional domains, related small functional site motifs and preserved 3D-structural features. These features were used to investigate the biological function of each of the 70 genes that make up the MammaPrint breast cancer gene expression profile:
For the annotation of functional domain architecture of individual genes, genes were translated into protein sequences, and the workflow for functional annotation of proteins implemented on the SMART web server 25 was followed:
To identify subcellular localization of a protein, the presence of sorting signals and/or cleavage sites was predicted by the bioinformatics tool SignalP. 24
To identify transmembrane regions of a protein, the TMHMM 25 algorithm was used. Regions of the protein separated by transmembrane regions were analyzed separately.
To identify conserved functional domains (eg, serine/threonine kinase domains, HLH transcription factor domains, epidermal growth factor- like domains), the bioinformatics tool HMMER 26 was run against the Pfam 27 and SMART 23 databases.
To analyze the segments that are not covered by highly conserved functional domains and low complexity regions, homologies to other proteins were retrieved with BLAST search. 28
For the interpretation of the biological functions of the individual genes within the cellular context (signal transduction pathway, metabolism pathway and protein-protein interaction), Ingenuity Pathways Analysis (Ingenuity® Systems, www.ingenuity.com) was used and a detailed literature review was performed.
Authors' contributions
ST carried the functional analysis and drafted the manuscript. PR participated in its design and helped to draft the manuscript. RB and LvV participated in the design of the study and helped to draft the manuscript. FdS conceived of the study, participated in its design and helped to draft the manuscript. AMG conceived of the study, participated in its design and coordination and helped to draft the manuscript. All authors read and approved the final manuscript.
Disclosure
This manuscript has been read and approved by all authors. This paper is unique and is not under consideration by any other publication and has not been published elsewhere. The authors are employed by Agendia BV. The authors confirm that they have permission to reproduce any copyrighted material.
Footnotes
Acknowledgements
The study was supported by a grant from TIPharma project T3–108 The authors would like to thank Dr R Bender and Dr B Chan for critically reading the manuscript.
