Abstract
Prostate cancer is the second most common cancer and the second leading cause of cancer death in men. However, prostate cancer can be effectively treated and cured, if it is diagnosed in its early stages when the tumor is still confined to the prostate. Combined with the digital rectal examination, the PSA test has been widely used to detect prostate cancer. But, the PSA screening method for early detection of prostate cancer is not reliable due to the high prevalence of false positive and false negative results. Epigenetic alterations including hypermethylation of gene promoters are believed to be the early events in neoplastic progression and thus these methylated genes can serve as biomarkers for the detection of cancer from clinical specimens. This review discusses DNA methylation of several gene promoters during prostate carcinogenesis and evaluates the usefulness of monitoring methylated DNA sequences, such as
Introduction
Prostate cancer (PCa) is the second most common cancer in men after skin cancer and the second leading cause of cancer death in men after lung cancer. In the United States, there are over 300,000 newly diagnosed cases each year, and about 40,000 patients die of the disease. 1 Approximately 90% of patients with advanced prostate cancer develop osseous metastases, which are difficult to eradicate. Patients with osseous metastases have a mean survival time of nine months to one year. 1 PCa can be effectively treated and cured, however, if it is diagnosed in its early stages (i.e. in stage I and II), when the tumor is still confined to the prostate. Combined with the digital rectal examination, the prostate specific antigen (PSA) test has been widely used to detect PCa (Cancer Facts, National CancerInstitute; http://cis.nci.nih.gov/fact/5_29.htm). This test measures the serum levels of PSA, an enzyme that is produced by the prostate and released into the bloodstream, reaching concentrations no more than 3–4 ng/ml in healthy individuals. PSA levels above that value are considered as an indication of possible PCa. However, PSA is specific for prostate tissues, but not for PCa. Various non-cancerous conditions such as benign prostatic hyperplasia (BPH), prostatitis, prostatic ischemia or infarction can cause elevated levels of PSA (http://cis.nci.nih.gov/fact/5_29.htm). Further, serum PSA levels are not always a sensitive indicator for PCa, as these may be normal despite the presence of the disease. 2 Thus, the PSA screening method for early detection of PCa is not reliable due to the high prevalence of false positive and false negative results (sensitivity 90%; specificity 10%–31%). 3 Consequently, only 25 to 30 percent of men who have prostate biopsies based on elevated PSA levels are diagnosed with PCa. 4 Recently, assays based on the detection of the specific serum marker EPCA-2 (sensitivity 94%, specificity 92%) 5 and overexpression of telomerase (sensitivity 58%, specificity 100%), DD3 gene (sensitivity 67%, specificity 83%), 3 and prostate cancer gene 3 (PCA3) (sensitivity 58%, specificity 72%) 6 have been established and bear great promise for PCa diagnosis, and may reduce the number of unnecessary biopsies.
Epigenetic alterations, including hypermethylation of CpG islands in the gene promoters are believed to be early events in neoplastic progression.7–12 However, recent findings in prostate carcinogenesis provide evidence that DNA hypomethylation changes occur subsequent to CpG island hypermethylation in later stages of carcinogenesis. 13 Hypermethylation of tumor suppressor gene promoters contributes to their silencing during the neoplastic process. 14 Thus, methylated gene promoters can serve as markers for the detection of cancer from clinical specimens such as tissue biopsies or body fluids. 15 Compared to tests that measure cancer-related proteins or RNAs, tests that measure gene alterations at the DNA level have several advantages for the early detection of cancer. DNA is stable in many of the conditions under which clinical specimens are collected and stored. Many DNA modifications can be reliably detected by PCR-based techniques,16,17 meaning that very small amounts of DNA are needed for such tests. PCR amplification-based tests also allow detection of as few as one cancer cell (or genome copy) in a background of thousands of normal cells, thereby permitting detection of a cancer before it can be visualized by imaging or traditional pathology. Moreover, DNA alterations can be measured qualitatively, as well as quantitatively. Finally, assays based on the DNA alterations can be both diagnostic and prognostic. Therefore, methylated DNA sequences can form the basis of a sensitive and specific, robust and informative test for the detection of cancer. 17
Alterations of DNA Methylation during Carcinogenesis: Hypomethylation in the Introns and Hypermethylation in the Promoter
DNA methylation refers to the covalent binding of a methyl group specifically to the carbon-5 position of cytosine residues of the dinucleotide CpG (Fig. 1). This is catalyzed by a family of enzymes, the DNA methyltransferases (DNMTs). Two types of DNA methylation alterations have been demonstrated in human cancers. The first refers to global hypomethylation in which the genomes of cancer cells show decreased methylation compared to normal cells.18–20 The hypomethylation is primarily due to the loss of methylation in repetitive elements and other non-transcribed regions of the genome. This genome-wide hypomethylation potentially leads to loss of imprinting, chromosomal instability, cellular hyperproliferation, and activation of oncogenes 21 such as K-ras and PU.1.22–25
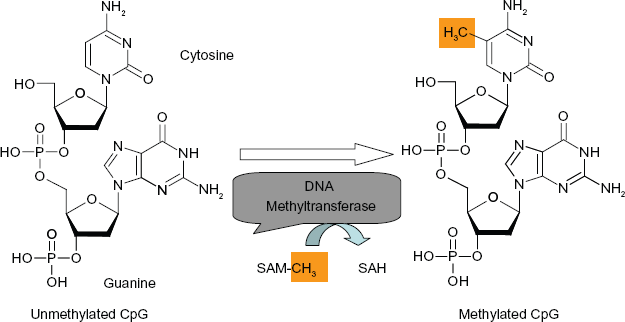
DNA methylation catalyzed by DNA methyltransferase. DNA methyltransferase transfers methyl group from S-adenosyl methionine (SAM-CH3) to cytosine yielding S-adenosyl homocysteine (SAH) and 5-methylcytosine.
The second type of methylation alteration in cancer cells is the hypermethylation of CpG islands in the promoter regions of tumor suppressor and other regulatory genes that are normally unmethylated. The promoter regions of these genes may be inactivated by methylation, which silences their expression (Fig. 2). However, differential methylation is not a general mechanism for regulating gene expression, because most inactive promoters remained unmethylated. 26 It is thought that DNA methylation alters chromosome structure and defines regions for transcriptional regulation. Clusters of CpG sites are found dispersed around the genome and are referred to as CpG islands. 27 These islands are found in the promoter region of about 60% of genes, and in exons, introns, and repetitive elements of most genes. In normal cells, most CpG islands in the promoter regions are unmethylated whereas CpG islands in intronic regions and repetitive elements are heavily methylated, perhaps to help the cell identify regions for gene transcription.
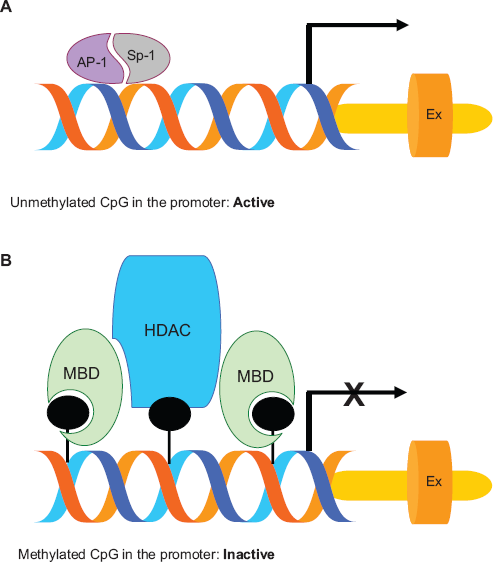
Simplified cartoon showing gene transcription by unmethylated promoter (A) and gene silencing by the methylated promoter (B).
Although the importance of CpG island methylation has been demonstrated in cancer, the mechanisms that lead to these changes in cancer are not yet understood. Of three members (DNMT1, DNMT3a, and DNMT3b) of the DNA methyltransferase family, DNMT1 is believed to be primarily involved in the maintenance of CpG methylation.28,29 However, other studies suggest that DNMT3b, independently or in cooperation with DNMT1, also contributes to hypermethylation.30–32 The suppression of transcription by DNA methylation may occur by either direct inhibition 33 or indirect inhibition 34 of transcription factor binding. For the latter, a family of proteins known as methyl binding domain (MBD) proteins is believed to specifically bind DNA containing methylated CpG sites. 34 At least three of the five known members of this family (MeCP2, MBD2 and MBD3) have been shown to be associated with large protein complexes containing histone deacetylase (HDAC1 and HDAC2) and chromatin-remodeling (Sin3a and mi-2) activities.35,36 Histone deacetylase (HDAC1 and 2) and chromatin remodeling activities (Mi-2 and Sin3a) produce alterations in chromatin structure that make it refractory to transcriptional activation. 37 In addition to the large protein complexes, the MBD proteins may associate with several other complexes involved in transcriptional repression. Recently, MeCP2 was shown to interact with at least two other proteins, c-ski and N-CoR, known to be involved in transcriptional repression. 38 However, Ohm et al recently hypothesize that the stem cell-like chromatin pattern may predispose tumor suppressor genes to DNA hypermethylation and heritable gene silencing during tumor initiation and progression. 39
As mentioned above, cancer cells exhibit two apparently opposing changes in the DNA methylation pattern: a decrease of DNA methylation in the intronic CpG islands and an increase of DNA methylation in the promoter CpG islands. Recent studies suggest that both changes may play important roles in the tumorigenic process. However, the increased methylation at the promoter CpG islands has been by far the most studied and has a much clearer role in carcinogenesis. Increased CpG island methylation can result in inactivation of many well-characterized tumor suppressor genes (e.g.
Hypermethylated Genes in Prostate Cancer
In prostate cancer, a large number of genes (e.g. DNA damage repair genes, tumor-suppressors, cell cycle control genes, cell adhesion molecules, and signal transduction genes) contribute to initiation and progression of the disease and expression of these genes is correlated with the pathological grade. 12 , 15 , 41–45
Interestingly, expression of some of these genes is regulated by DNA methylation at their promoters. In general, promoter hypermethylation is associated with the loss of expression of these genes. A spectrum of methods is available for DNA methylation. These include cytosine deamination PCR, semi-quantitative and quantitative methylation-specific PCR (MSP), differential methylation hybridization (DMH), restriction landmark genomic scanning (RLGS), single-nucleotide primer extension (SNuPE), pyrosequencing, and methylation microarray for large-scale genome analysis.15,46 However, MSP is a simple and sensitive method, and is the most commonly employed method for methylation analysis.
15
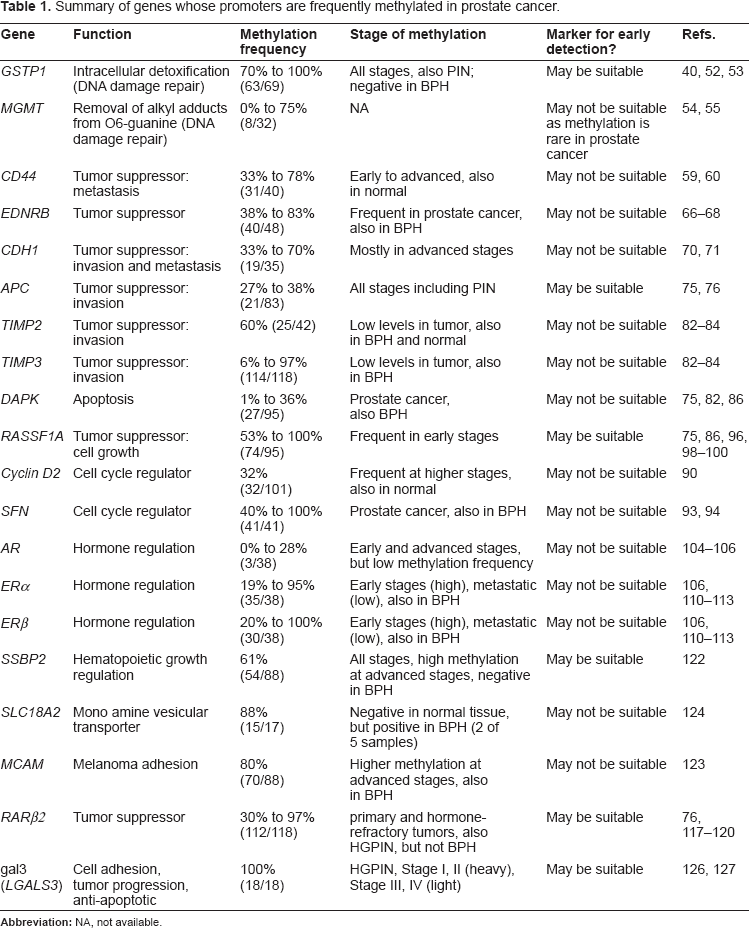
Table 1 shows some common genes that are hypermethylated in the prostate cancer and are tested for the development of early detection. These genes participate in DNA damage repair (
Summary of genes whose promoters are frequently methylated in prostate cancer.
The glutathione S transferases (GSTs) are a family of enzymes involved in protecting cells from DNA damage, and thence cancer initiation. GSTs carry out intracellular detoxification of xenobiotics and carcinogens by covalent bonding to glutathione via a thiol link.
47
In humans, six cytosolic isoforms of GST (alpha, mu, pi, sigma, theta, and omega) and one membrane isoform of GST have been described- of which the pi isoform (
Another DNA repair gene O
6
-Methylguanine-DNA-Methyltransferase (
CD44, an integral membrane glycoprotein, plays a role in cell adhesion and cell-matrix interactions as a receptor for hyaluronic acid and osteopontin. 58 In prostate cancer, CD44, acts as a metastasis suppressor gene, and its down-regulation is associated with tumor progression and metastasis. Hypermethylation of CpG islands in the promoter region of CD44 results in decreased expression.59,60 In a recent study, methylation of CD44 was evaluated among men who develop biochemical PCa recurrence after receiving radical prostatectomy. 61 The methylation profile of CD44 was found to be an independent predictor of biochemical recurrence (associated with 9-fold increased risk). This finding, if validated in larger studies, may identify patients with aggressive cancer.
The endothelin peptides consisting of three iso-types, ET-1, ET-2, and ET-3 have potent vasoconstrictive properties and are differentially expressed in various cells and tissues.
62
Two receptors for endothelin peptides (ETA and ETB) have been identified in various cells and tissues. Belonging to a family of hyptohelical G-protein-coupled receptors, they are differentially expressed during prostate cancer progression and also differ in binding the enothelin isotypes.63,64 ETA binds to two isotypes ET-1 and ET-2 only, whereas ETB binds to all three isotypes ET-1, ET-2, and ET-3.
63
In prostate cancer, expression of ETA is increased, whereas expression of ETB is reduced.
65
Moreover, the ETB gene (
E-cadherin, a transmembrane glycoprotein and a member of the cadherin family of cell adhesion molecules, may function as a tumor suppressor gene in invasion and metastasis by mediating cell-cell adhesion via calcium-dependent interactions.
69
In prostate cancer, expression of E-cadherin is decreased during tumor progression and this decreased expression has been correlated with hypermethylation of the promoter in patients biopsies.70,71 However, in metastatic prostate cancer cells in bone, E-cadherin expression is increased.
72
Interestingly, the promoter of E-cadherin gene
The adenomatous polyposis coli (APC) is a multifunctional protein that acts as a tumor suppressor gene in familial adenomatous polyposis.
73
It plays a role in the Wnt signaling pathway, cell migration, cell adhesion, and mitosis.
74
In prostate cancer development,
Tissue inhibitors of metalloproteinases (TIMPs) are known to control the activity of matrix metalloproteinases (MMPs)
79
in several biological processes such as cell growth, apoptosis, invasion, metastasis and angiogenesis.80,81 Four members of TIMPs have been identified and are known to be down-regulated in prostate cancer.82,83 Down-regulation of TIMP proteins is associated with hypermethylation of the corresponding gene promoters.
84
In particular, low-level methylation of
Death-associated protein kinase (DAPK), a member of the pro-apoptotic calcium regulated serine/threonine kinases, is expressed in all tissues.
85
Its inactivation leads to the loss of this important apoptotic pathway. Although different mechanisms may affect
Cell cycle genes such as retinoblastoma protein (RB), cyclins, cyclin dependent kinases (CDKs), and CDK inhibitors (CDKIs) are very important in regulation of the cell cycle. In cancer, the efficacy of cell cycle checkpoints is often affected, especially control of the G1/S transition. 87 CDKIs are negative regulators of the cell cycle and considered to be tumor suppressor genes. CDKIs are categorized into two families, the INK4 family and the CIP/KIP (kinase inhibitor protein) family. The INK4 family is composed of four members CDKN2A or p16, CDKN2B or p15, CDKN2C or p18, and CDKN2D or p19. 88 The CIP/KIP family includes CDKN1A or p21, CDKN1B or p27, and CDKN1C or p57. 89 While the INK4 family specifically inhibits CDKs 4 and 6, the CIP/KIP family inhibits most CDKs.88,89
In prostate cancer, cell cycle checkpoint genes can be inactivated by a number of mechanisms such as deletion, point mutation, and hypermethylation. For example, cyclin D2 promoter methylation has been detected in prostate cancer and correlated with disease progression.
90
However, other cell cycle genes such as
RAS proteins are involved in extra-cellular signal transduction and regulate cell growth, survival and differentiation.
95
A new family of genes encoding RAS-binding proteins, RAS association domain family 1 gene (
Androgens such as testosterone and 5α-dihydrotestosterone are the main steroid hormones in the prostate and act through the androgen receptor (AR). 101 The expression of the AR gene and androgen dependence is consistent with the early stages of prostate cancer.102,103 However, AR expression and androgen dependency is lost in the terminal stages of metastatic prostate cancer and the loss of AR expression appears to be regulated by DNA methylation in its promoter.104–106 Moreover, methylation appears to be more prevalent in hormone-refractory tumors than in primary tumors. 105
Estrogens are believed to play an important role in prostate carcinogenesis by acting through intracellular receptors, ER-α and ER-β.107,108 These receptors are expressed in a cell and tissue specific manner, and involved in the regulation of the normal function of reproductive tissues. 109 However, several studies have reported the loss or down-regulation of these receptors during prostate cancer development110,111 and the DNA methylation in their promoters has been associated with decreased or loss of expression of these two genes in prostate cancer.106,112 Moreover, a high frequency of methylation in the promoter region of the ER-β has been observed at the early stages of the disease, whereas methylation declined in metastatic tumors. 112 Promoter methylation of ER-α and ER-β in BPH has also been reported to a lesser extent than in prostate cancer tumors.112,113
Single-stranded DNA-binding protein 2 (SSBP2), a novel regulator of hematopoietic growth and differentiation,
121
has recently been shown to be hypermethylated in prostate cancer.
122
In a quantitative MSP assay, the
The melanoma cell adhesion molecule (
The vesicular monoamine transporter 2 gene,
The galectin-3 gene
Methylated Genes Suitable for Early Detection of Prostate Cancer
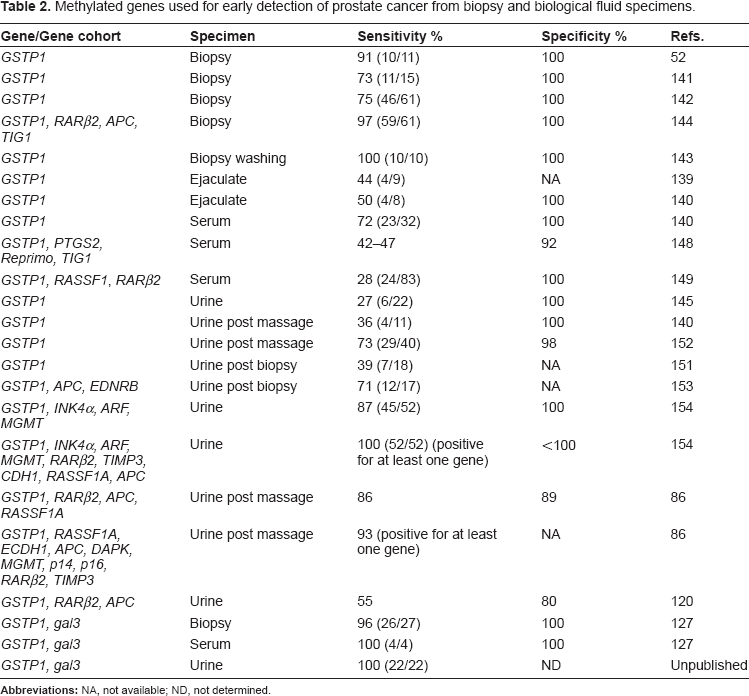
Although many genes are observed to be methylated in prostate cancer, a few genes have been investigated as targets for early detection (Table 2). Most have insufficient methylation frequency to provide the needed sensitivity, while other methylated genes are also present in the BPH, making them non-specific. The most suitable gene appears to be
Methylated genes used for early detection of prostate cancer from biopsy and biological fluid specimens.
The finding that the gal3 gene promoter is completely methylated in stage I and II PCa makes the gal3 gene
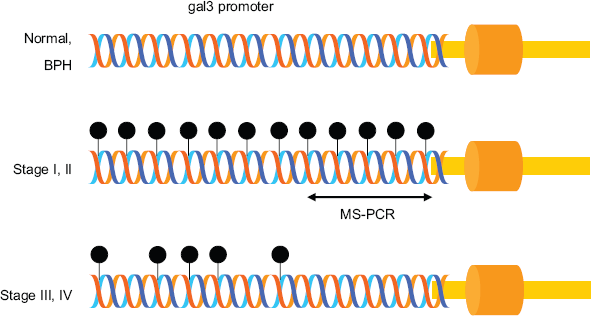
Schematic representation of MS-PCR. In normal and BPH prostate tissues, the gal3 promoter is unmethylated, whereas in stage I and II, it is methylated heavily. However, gal3 promoter is lightly methylated in stage III and IV. Stage-specific cytosine methylation of the gal3 promoter enabled the development of MS-PCR for the detection of stage I and II PCa.
Early Detection of Prostate Cancer in the Biological Fluids such as Serum and Urine
Several studies suggest that prostate cancer can be detected in serum and urine.15,17,120,145 Identification of methylated DNA in urine would be a critical milestone in the development of a non-invasive diagnostic for early stages of prostate cancer. DNA passes into urine and blood through three main routes. 146 The first occurs when prostate cells are directly released into the urethra through prostatic ducts. DNA can also pass into urine by phagocytosis, in which macrophages engulf DNA from necrotic tumor tissue then the macrophages themselves appear in both urine and blood. Lastly, when cell proliferation is accelerated, cellular DNA content can overwhelm phagocytes and directly spill into circulation and urine.
The first study of body fluid-based detection involves the analysis of plasma samples of prostate cancer for the detection of
As described above, prostate cancers shed neoplastic cells or debris amenable to DNA analysis. Cairns et al
145
first analyzed voided urine samples from patients with prostate cancer and detected
Goessl et al
152
demonstrated the utility of
An additional study in which multiple gene analyses were evaluated is that of Hoque et al,
154
who monitored a 4-gene cohort (
Roupret et al
86
examined urine samples from 95 prostate cancer patients undergoing radical prostatectomy and from 38 control patients for aberrant methylation of 10 genes (
Vener et al
120
recently tested urine samples from 234 patients with PSA concentrations > or = 2.5 microg/L using 3 gene-cohort,
Our studies of gal3 MS-PCR in combination with
Concluding Remarks
Several studies suggest that assays based on the detection of DNA methylation may provide better specificity and sensitivity than the PSA test. Furthermore, the methylated genes themselves may serve as promising biomarkers for early detection of prostate cancer. However, it should be emphasized that most published studies have been in small scale. Large scale studies will be necessary before any of theses assays can be considered clinically useful. In addition to clinical validation, assays for methylated genes must be robust, simple, sensitive, specific, and made available at affordable costs.
Regarding specificity of the assay, the most important consideration is to select a gene or genes whose promoter methylation is specific to prostate cancer. For example,
Regarding sensitivity, most studies of single genes or combinations of multiple genes did not reach a sensitivity of more than 75% and most importantly, these markers failed to detect early stages of prostate cancer.120,144 Therefore, a reliable marker for early detection (stage I, II) of prostate cancer is yet to be identified. We have reported heavy promoter methylation of gal3 gene (LGALS3) in early stages of prostate cancer and have demonstrated the usefulness of
The specificity of gal3 MS-PCR to prostate cancer is in question as gal3 is also known to be aberrantly expressed in many cancers. In most cancers such as lung, liver, gastric, head and neck, thyroid, and bladder cancers, expression of gal3 is increased,160–165 while expression is decreased in prostate, kidney, and pituitary cancers.127,132,133,166 Although the loss of gal3 expression in pituitary tumor is partially due to promoter methylation, these methylation sites are distinct from those we observe in PCa by gal3 MS-PCR. Thus, by judicious primer design to target only the PCa-relevant regions the assay should be insensitive to pituitary cancer. Although it remains to be seen if the gal3 promoter is heavily methylated in kidney cancers, gal3 MS-PCR in a several urine specimens from kidney and bladder cancers yielded negative results (unpublished data) underscoring its specificity to prostate cancer. However, non-specificity of the assay can be substantially reduced by combining the assay with the conventional PSA test as explained below. The PSA test is complicated by a large number of false positive results (specificity 10%–31%) rather than the false negative results (sensitivity 90%). However, the PSA false positive data are due to non-cancerous conditions of the prostate such as BPH, prostatitis, prostatic ischemia or infarction, as the PSA is specific for prostate tissues. Thus combining the gal
The selectivity of the assay for indolent prostate cancer bears some consideration. Indolent cancer (associated with low Gleason grade and a low PSA value) rarely becomes symptomatic during a patient's lifetime, and patients with indolent cancer are offered active surveillance rather than treatment.167,168 However, a sensitive assay such as qMSP or “MethyLight” (detection limit 20 pg of DNA, equivalent to one cell or one genome copy) and early detection markers of prostate cancer identify a considerable proportion of cancers that are indolent. 167 There is a measurable risk, however, of over-treatment of patients with indolent disease, which should be avoided because of potential adverse psychological and somatic side effects. 168 Therefore, if the assay turns out to be positive with indolent cancer (samples with low PSA, Scenario 4 as described above), further tests (such as sarcosine marker) 169 should be performed to confirm the indolence behavior of the cancer prior to any treatment.
The prospect for improved detection of PCa with the gal3/
Conflict of Interest Statement
The author is one of the two inventors on a patent related to the application of galectin-3 promoter hypermethylation for early detection of prostate cancer.
Footnotes
Acknowledgments
The author is very grateful to Dr. Thambi Dorai (Saint Vincent Catholic Medical Centers, New York, NY) and Dr. Eric Schott (University of Maryland Biotechnology Institute, Baltimore, MD) for critically reading the manuscript and helpful suggestions. The work carried out in the author's laboratory has been supported by the UMBI Presidential Proof of Concept Award and the National Institute of Health Grant RO3 CA133935–01.
