Abstract
Background
Influenza has been shown to increase the risk for severe bacterial infection, in the tropics the seasonality of influenza epidemics is less marked, and this may not be the case. Dengue is often followed by prolonged asthenia and some physicians hypothesized increased susceptibility to infections based on anecdotal observations.
Design and methods
Time series of influenza and dengue surveillance were confronted bacterial sepsis admissions to test the hypotheses. Monthly surveillance data on influenza and dengue and aggregated sepsis data in Cayenne hospital were matched between 24/10/2007 and 27/09/2016. An ARIMA (1,0,1) model was used.
Results
The series of the number of monthly cases of sepsis was positively associated with the monthly number of cases of influenza at time t (β=0.001, p=0.0359). Forecasts were imperfectly correlated with sepsis since influenza is not the only risk factor for sepsis. None of the ARIMA models showed a significant link between the dengue series and the sepsis series.
Conclusions
There was thus no link between dengue epidemics and sepsis, but it was estimated that for every 1,000 cases of flu there was one additional case of sepsis. In this tropical setting, influenza was highly seasonal, and improved vaccination coverage could have benefits on sepsis.
Introduction
Significance for public health
Simultaneous infections may have complex consequences ranging from synergistic, neutral or antagonistic. Dengue fever and influenza cause repeated epidemics and may have consequences on the host's immune response. Hypothesizing that an infection by dengue or influenza could increase the risk of bacterial infection we showed that there was no relation between dengue and sepsis, but that there was a relation between influenza and sepsis. Despite the tropical setting of French Guiana, the highly seasonal pattern of influenza suggests the vaccine reimbursement window should be extended.
Coinfections by different pathogens lead to agonistic or antagonistic consequences depending on the agents involved. Epidemiological and experimental evidence show that prior respiratory viruses pave the way for invasive bacterial infections. Prior viral infections may enhance adherence, colonization and invasion by bacterial pathogens in many different ways: Physical damage to the respiratory epithelium and function, ciliary clearance of bacteria, and bacterial adherence to injured tissues. 1 Immunopathology also facilitates bacterial disease: Measles, for example, reduces the ability to mount an effective immune response against bacterial pathogens; in vitro, this translates in alterations of lymphoproliferative responses, cytokine profiles, and antigen-presentation; in vivo this translates in lymphopenia, suppressed Mantoux responses, and impaired vaccine efficacy. 2 Influenza leads to neutrophil dysfunction, IL10-mediated immunosuppression, a pulmonary NK cell response, an IFN gamma-mediated macrophage function depression and macrophage desensitization to bacterial ligand-triggered Toll-like receptors (TLR) signaling. These examples illustrate the complexity of the mechanisms underpinning the adverse outcomes of successive or simultaneous infections. The temporal sequence of the respective infections may also modulate the nature and intensity of the immune response, and promote severe immunopathology leading to sepsis.
Dengue fever is an arthropod borne viral disease which leads to large epidemics that often overwhelm health services. In a small fraction of cases, life-threatening severe forms of dengue occur with bleeding manifestations or capillary leakage. 3 Although acute symptoms often recede within a week, a large number of persons however complain of post dengue exhaustion which may persist for weeks after the acute episode. The pathophysiology of dengue is still incompletely understood but it leads to complex immune phenomena. 4
In the present study, we hypothesized that dengue may lead to transient immunosuppression which could increase the risk of subsequent bacterial infections. Several studies have shown that influenza increases the risk for severe bacterial infection. 5 These studies mostly took place in temperate zones. In the tropics the seasonality of influenza epidemics is less marked, the air temperature and virus aerosolization may not be followed by the same consequences, notably in terms of severe bacterial pneumonia.5-7
Therefore, we confronted time series of influenza surveillance, Dengue surveillance and admissions in Cayenne hospital for bacterial sepsis to look for any statistical relationship between these viral epidemics and subsequent bacterial sepsis.
Design and Methods
The study was retrospective and used data sets collected between 24/10/2007 and 27/09/2016.
Influenza dengue data
The surveillance data was obtained from the CIRE Antilles Guyane (Interregional cell for epidemiology) which is responsible for the monitoring of epidemic diseases and records suspected and confirmed cases on a weekly basis from reference laboratories and a sentinel network of clinicians. The CIRE Antilles Guyane is part of the Institut National de Santé Publique France and covers all the territory of French Guiana. Surveillance data included laboratory confirmed data and also a sentinel network of physicians who report clinical cases in an epidemic context (ARAVEG, Centres Délocalisés de Prévention et de Soins). The epidemic is defined by confirmed cases, but during an epidemic the sentinel clinical data has very good predictive value. This surveillance data is used for the health authorities’ response and allows them to declare a state of epidemic and unfold appropriate measures. Aggregated sepsis data in Cayenne hospital were collected from the hospital medical information system (PMSI, CORA software). The database was queried using the sepsis ICD codes A390, A392, A400, A401, A402, A403, A408, A409, A410, A411, A412, A413, A414, A415, A418, A419 (Meningococcal meningitis, Acute meningococcemia, Streptococcal sepsis, other sepsis (sepsis due to Staphylococcus aureus, sepsis due to other specified staphylococcus, sepsis due to unspecified staphylococcus, sepsis due to Hemophilus influenzae, sepsis due to anaerobes, sepsis due to other Gram-negative organisms, Gram-negative sepsis, unspecified)).
Sepsis data
These codes were pooled in a single “sepsis” variable, which was present or absent, and was a single value event if some patients could in theory have 2 distinct ICD codes. Cayenne hospital is the French Guyana's reference hospital, the only one with an Intensive Care Unit.
Data analysis
The collection of influenza, dengue, and sepsis data at monthly intervals is by definition a time series. Time series were first plotted to look for macroscopic trends. Since there can be seasonal variations, and increasing and decreasing trends that can lead to erroneous interpretations of correlations, the analysis first removes such variations, hence obtaining stationary data from which it can try to accurately model the explanatory variables and forecast future values. Then Auto Regressive Integrated Moving Average (ARIMA) models [Box and Jenkins, 1976] were used to look for a temporal relation between influenza and dengue epidemics, and the number of cases of sepsis. ARIMA models allow modeling time series taking into account potential residual autocorrelations under the hypothesis that each measure may be correlated to the previous ones. Thus ARIMA models allow combining three types of temporal processes, the contribution of each being specified in the notation ARIMA(p,d,q), where p is the order of the autoregressive process AR(p), d the integrated part corresponding to the degree of differencing I(d), and q the order of the moving average model MA(q). Multiplicative seasonal ARIMA is applied to series of a similar phenomenon that is repeated through time. In addition to the initial parameters, seasonal parameters for a specified lag must thus be estimated. As for simple ARIMA they are: seasonal autoregressive (ps), seasonal differencing (ds), and seasonal moving average (qs) parameters. Thus an ARIMA model including a seasonal trend is specified as ARIMA(p,d,q)(ps,ds,qs).
Model fitting
Identification stage for the sepsis series - Based on a descriptive analysis (plots) of the monthly number of sepsis cases and of the series’ autocorrelation (ACF) and partial autocorrelation functions (PACF), different ARIMA models (p,d,q)(ps,ds,qs)were tested for the sepsis series.
Estimation and diagnostic checking stage for the sepsis series - Models with significant parameters at the 10% were selected for the final stage. The Ljung Box Q test was used to check the nonautocorrelation of residuals of the selected models. Normality was examined graphically based on the histogram of residuals and on the normality plot.
Final stage - Lags of 0 and 1 month of the series of monthly values of influenza was subsequently incorporated in the previous selected models in order to identify an eventual impact of influenza on the number of sepsis cases. Potential final models included significant parameters at the 10% level. The Ljung-Box Q test, the residuals histogram and the normality plot were used to confirm that the residual series of the models were Gaussian white noise. Akaike's Information Criterion (AIC), the principle of parsimony, the control of the model variance and the quality of previsions were the performance criteria used to ultimately decide for the final model. A similar reasoning was used for dengue.
Results
Description of the different series
The monthly number of suspected influenza cases and sepsis over the study period are shown in Figure 1, which shows apparent, but nonsystematic simultaneous peaks of influenza and sepsis. Overall, 24/10/2007 and 27/09/2016, there were 163,767 cases of influenza and 68,901 cases of dengue reported by the sentinel network, and 1,808 cases of sepsis. Data show that except for year 2008, influenza outbreak occurred every year. The magnitude of the epidemics was highly variable across years: a major peak was observed in year 2009 with up to 7000 cases in September. Years 2012 and 2014 also recorded important epidemics. The outbreaks appeared more moderate for other years. The period of influenza activity was also variable across years; thus, though in most cases an intense activity is observed during the first months of the years. Time series of sepsis showed a mean number of 14 admissions in hospital for sepsis per month, varying from 2 to 37 monthly cases.
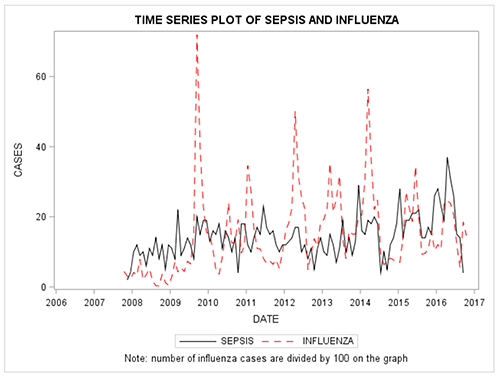
Time series plot of influenza and sepsis showing apparent, but nonsystematic simultaneous peaks of influenza and sepsis.
The monthly number of suspected dengue cases and sepsis over the study period are shown in Figure 2, highlighting the 3 dengue epidemics of 2009, 2010 and 2013, with up to 2,600, 1,500 and 2,800 monthly cases identified, respectively. The series description does not reveal any trend for a correlation between dengue outbreaks and a subsequent increase in the number of sepsis cases.
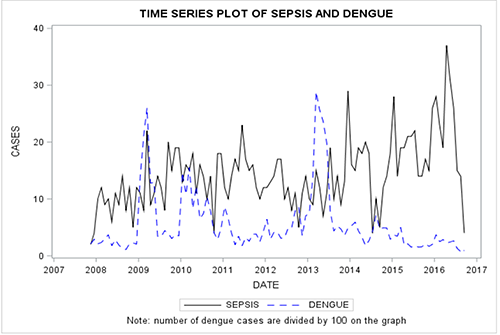
Monthly number of suspected dengue cases and sepsis.
ARIMA modeling of sepsis
The identification stage allowed testing different models among which two were retained following the estimation and diagnostic stage: ARIMA (2,0,0) and ARIMA (1,0,1). The fit of the two selected models was correct with parameters that were all significant below 5%, uncorrelated normally distributed residuals. These two models were thus used to test influenza and dengue as explanatory variables of sepsis.
Sepsis and influenza
Figure 3 shows the cross-correlations between sepsis and influenza series with significant correlations at lags 0 and 1 month, which was judged to be clinically pertinent given the local mucous membrane alterations caused by influenza. 8 From each of the two previously detailed models of the sepsis series, two new models were declined: one including influenza at time t as input series and one including influenza at time t-1. In all four models influenza was significantly associated with sepsis at the 5% significance level, confirming a correlation between the influenza time series and the sepsis time series. The final model selected was an ARIMA(1,0,1)+ influenza(t). Estimates of the model are shown in Table 1. The series of the number of monthly cases of sepsis was positively associated with the monthly number of cases of influenza at time t (β=0.001, p=0.0359). The β coefficient of the influenza series thus translated in an additional case of sepsis out of every 1000 cases of influenza.
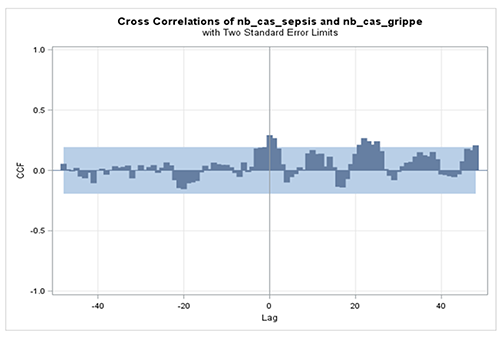
Cross-correlations between sepsis and influenza series with significant correlations at lags 0 and 1 month.
Estimates of the ARIMA(1,0,1) model + influenza(t).

Figure 4 shows the observed number of sepsis cases in regard to the predicted number of sepsis cases according to the ARIMA model with influenza. Forecasts were imperfectly correlated with sepsis.

Number of sepsis cases in regards to the predicted number of sepsis cases according to the ARIMA model with influenza.
Sepsis and dengue
We integrated the dengue series to the four initial sepsis models testing lags 0, 1 and 3. None of the models showed a significant link between the dengue series and the sepsis series.
Figure 5 shows the cross-correlations between the dengue and the sepsis time series. The only significant correlations appeared for very long lags that did not seem biologically plausible in terms of incubation of diseases and their temporal overlap. Thus, both the descriptive, cross-correlation and modeling approaches led to coherent results which did not show a correlation between the two series.

Cross-correlations between the dengue and the sepsis time series.
Discussion
In interpandemic periods, there are an estimated 1 billion cases of flu, 3-5 million cases of severe illness, and 300,000-500,000 deaths, each year, worldwide. 9 A number of studies have shown that a large proportion of influenza pneumonia in fact results from a synergy between the influenza virus and S. pneumoniae.8,10 The influenza virus increases colonization of the respiratory tract by Streptococcus pneumonia which then facilitates its invasion and the development of pneumonia. This may also occur for other bacteria such as H. influenza or Staphylococcus aureus.1,11-14 Influenza A infections can also have a significant influence on the risk of subsequent meningococcal meningitis15-19 with variable strength for different influenza subtypes.
Here we observed significant temporal relationships between influenza epidemics and the number of cases of sepsis hospitalized in the reference hospital of French Guiana. Despite being close to the equator, influenza seasonality was marked, presumably because of the close link with mainland France where influenza epidemics often affect air travelers to French Guiana who then propagate the epidemic locally. 20 Although the population of French Guiana is small, the output ARIMA model estimates approximately one additional case of sepsis out of every 1,000 cases of influenza which during an epidemic is not negligible. Therefore, vaccination against seasonal influenza could have non negligible benefits in reducing the number of cases of sepsis. The vaccine for the northern hemisphere is effective in French Guiana, however the timing and the shape of the epidemic are different from what is observed in mainland France. 20 On average influenza epidemics in French Guiana last 22 weeks whereas in mainland France they last 9 weeks. 20 The onset of the epidemic usually occurs 4 weeks after the beginning of the epidemic in mainland France. However, vaccine reimbursement policy only extends until December 31 of a given year. This is likely to limit the number of persons vaccinated and given the 1 sepsis per 1000 influenza cases observed here, extending reimbursement of the vaccine after December may allow benefits on sepsis. 14 For dengue, after observing a case of histoplasmosis after an episode of dengue fever, the initial hypothesis put forward from the bedside was that some patients developed other infections after dengue. This hypothesis was not supported by the available data. Therefore, post dengue “immunosuppression” does not seem to have any consequences on subsequent sepsis. The limitations of the present study were possibly the insufficient time span and low patient numbers given the small population of French Guiana which could have masked less obvious interactions. In addition, the sepsis definition was broad, notably because of the small patient numbers when looking at specific diagnoses. The influenza and dengue cases were clinical suspicions which may not have been always accurate. However, they originated from a sentinel network working in close relation to the Health authorities and French Guiana's Pasteur Institutes National Reference Laboratory for arboviruses and influenza. The predicted positive value of the criteria for defining suspected cases by the sentinel network in a proven epidemic context is high, therefore we are confident that the time series reflected the evolution of the studied epidemics. 21
In conclusion, contrarily to our hypothesis there was no link between dengue epidemics and sepsis. As observed elsewhere there was a significant link between flu epidemics and sepsis. The flu epidemics were highly seasonal, and it was estimated that for every 1,000 cases of flu there was one additional case of sepsis. In this tropical setting, influenza was highly seasonal, and improved vaccination coverage could have benefits on sepsis.
Footnotes
Acknowledgements
Antoine Enfissi from Pasteur institute and Vanessa Ardillon from the CIRE Antilles Guyane for surveillance data.
