Abstract
MicroRNAs (miRNAs, miRs) are small noncoding RNAs that regulate gene expression at the posttranscriptional level. We have previously identified a group of novel miRNAs in proximal sciatic nerve after sciatic nerve transection by Solexa sequencing, and miR-sc3 is a member of the group. In this study, we aimed to investigate the effects of miR-sc3 on phenotypic modulation of Schwann cells (SCs). miR-sc3 was highly expressed in the injured nerve after sciatic nerve transection. An increased and decreased expression of miR-sc3 promoted and reduced the proliferation and migration of primary SCs, respectively. miR-sc3 directly targeted astrotactin 1 (Astn1) and led to translational suppression of Astn1. There was an inverse association between the time-dependent expressions of miR-sc3 and Astn1 in proximal sciatic nerve after sciatic nerve transection. Overall, miR-sc3 affected SC proliferation and migration by targeting Astn1, thus playing the regulatory role in peripheral nerve regeneration.
Introduction
The peripheral nervous system (PNS), unlike the central nervous system (CNS), is capable of spontaneous regeneration following nerve injury. Schwann cells (SCs), the principal glial cells in the PNS, contribute to the establishment of a beneficial environment for peripheral nerve regeneration. After traumatic injury to peripheral nerves, SCs in the distal stump dedifferentiate into a progenitor/stem cell-like state, proliferate, and migrate to form bands of Bungner for guiding axonal regrowth in the proximal stump and then promote axon remyelination (12). Recently, the specific cellular behaviors of SCs during nerve injury and regeneration have been broadly studied, suggesting the new roles of SCs in eliminating ectopic sodium channels from the internodes, preventing the premature differentiation of sensory organs, fasciculating the nerve fibers, and preventing oligodendrocytes from exiting the spinal cord (3,21).
MicroRNAs (miRNAs) are a class of endogenous, short noncoding RNA molecules with a length of about 22 nucleotides. They are highly conserved across species (2,4) and can negatively regulate up to 60% of the total human genes through posttranscriptional repression (5). The regulatory roles of miRNAs in neural development, neurodegeneration, and nerve regeneration have attracted considerable interest (7,9,17), and many studies have investigated the involvement of miRNAs in peripheral nerve injury and regeneration (14,24,26,28). For example, it has been demonstrated that miRNAs regulate the expressions of many genes that are required for myelination and maintenance of axons by SCs (6,19,27).
To screen novel miRNAs in the injured peripheral nerve and investigate their possible regulation of peripheral nerve regeneration, a rat model of sciatic nerve transection is usually used. We have identified a group of novel miRNAs that are differentially expressed in proximal sciatic nerve after sciatic nerve transection by using Solexa sequencing (15,16). Considering the tissue and spatiotemporal expression specificity of most miRNAs, we speculate that these miRNAs may be tightly controlled in a similar manner during nerve regeneration and that they might play important regulatory roles in the repair process. In the current study, we focused on miR-sc3, a member of the above-mentioned group, and investigated the influences of miR-sc3 on phenotypic modulation of SCs. Our results showed that miR-sc3 was differentially expressed in the injured nerve after sciatic nerve injury, and an increased expression of miR-sc3 promoted proliferation and migration of SCs by targeting astrotactin 1 (Astn1).
Materials and Methods
Animal Surgery
Adult, male Sprague–Dawley (SD) rats were obtained from the Animal Experiment Center of Nantong University in China. The animals received surgery to induce sciatic nerve transection as described previously (16). The proximal nerve segments were harvested at 0, 1, 4, 7, and 14 days post-nerve injury. RNA and protein samples were extracted from the harvested nerves. All the experimental procedures were conducted in accordance with Institutional Animal Care guidelines of Nantong University and were ethically approved by the Administration Committee of Experimental Animals in Jiangsu Province, China.
Cell Culture and Transfection of Primary SCs
Primary SCs were isolated from the sciatic nerve of 1-day-old SD rats and purified by removing fibroblasts with anti-Thy1.1 antibody (1:1,000) and rabbit complement (Sigma-Aldrich, St. Louis, MO, USA) as previously described (14). The final cell preparation consisted of 98% SCs, as determined by immunocytochemistry with anti-S100 (1:200; Dako, Carpinteria, CA, USA), a specific SC marker. The obtained SCs were cultured in Dulbecco's modified Eagle medium (DMEM; Gibco, Grand Island, NY, USA) containing 10% fetal bovine serum (FBS; Gibco) in a humidified 5% CO2 incubator at 37°C. SCs were passaged for no more than three passages prior to use.
Primary cultured SCs were transfected with synthetic miR-sc3, miR-sc3 inhibitor, or Astn1 siRNA (Ribobio, Guangzhou, China), respectively, using Lipofectamine RNAiMAX transfection reagent (Invitrogen, Carlsbad, CA, USA) according to the manufacturer's instructions.
Cell Proliferation Assay
Primary SCs were resuspended in fresh prewarmed (37°C) complete medium and plated at a density of 2 × 105 cells/ml on poly-L-lysine-coated 96-well plates (Sigma-Aldrich). At 36 h after cell transfection, 50 μM 5-bromo-2′-deoxyuridine (BrdU; Ribobio, Guangzhou, China) was applied to the cell culture, and SCs were incubated for an additional 24 h. The cells were fixed with 4% paraformaldehyde (PFA; Xilong Scientific, Guangzhou, China) in phosphate-buffered saline (PBS; Gibco) for 30 min. The Cell-Light™ BrdU DNA Cell Proliferation Kit (Ribobio) was used to detect cell proliferation rate according to the manufacturer's protocol. The ratio of BrdU-positive cells to cell population was calculated from images of randomly selected fields obtained under a DMR fluorescence microscope (Leica Microsystems, Bensheim, Germany). Assays were performed three times using triplicate wells.
Transwell-Based Cell Migration Assay
The migration ability of SCs was examined by a Transwell-based assay as described previously (14). The 6.5-mm Transwell chambers with 8-μm pores (Costar, Cambridge, MA, USA) were used, and the bottom surface of each membrane was coated with 10 μg/ml fibronectin. Primary SCs (106 cells/ml, 100 μl) were resuspended in DMEM, transferred to the top chamber of each Transwell, and allowed to migrate in a humidified 5% CO2 incubator at 37°C with the complete medium (600 μl) being pipetted into the lower chamber. The upper surface of each membrane was cleaned with a cotton swab at indicated time points. The cells adhering to the bottom surface of each membrane were stained with 0.1% crystal violet (Beyotime, Shanghai, China), imaged, and counted under a DMR inverted microscope (Leica Microsystems). Assays were performed three times using triplicate wells.
Wound-Healing Assay
A wound-healing assay was performed as described previously (18,23) with modifications. SCs were seeded onto the bottom chamber of a 24-well plate, incubated until confluent, and transfected with synthetic miR-sc3, an miR-sc3 inhibitor, or nontargeting negative controls for 24 h. Transfected SCs were starved in DMEM supplemented with 0.5% FBS (Invitrogen) and 0.15 μg/ml mitomycin C (Sigma-Aldrich) for 12 h and then wounded by a sterile pipette tip to form a linear scratch. The wounded cell monolayers were imaged under a DMR inverted microscope (Leica) at 0 and 12 h after wounding. The relative wound closure at 12 h compared with 0 h was measured using Wimscratch Quantitative Wound Healing Image Analysis (Wimasis GmbH, Munich, Germany).
Luciferase Assay
The 3′ untranslated region (3′-UTR) of endothelin 1 (Edn1), Shroom, contactin 2 (Cntn2), Astn1, or Adam10 was amplified by polymerase chain reaction (PCR) using rat genomic DNA as a template. The PCR products were subcloned into the region directly downstream of the stop codon in the luciferase gene in the luciferase reporter vector to generate p-Luc-UTR reporter plasmid. An overlap PCR method was used to construct a 3′-UTR mutant reporter plasmid with a pair of mutant primers (CCTAAGTCT
Quantitative Real-Time RT-PCR
To determine miR-sc3 expression, a total amount of 50 ng RNA was reverse transcribed using a TaqMan® MicroRNA Reverse Transcription Kit (Applied Biosystems, Foster City, CA, USA) or Prime-Script reagent Kit (Takara, Dalian, China) according to manufacturer's instructions. To detect the mRNA level of Astn1, RNA samples were reverse transcribed to cDNA by using a Prime-Script reagent Kit (Takara) according to manufacturer's instructions. Quantitative real-time RT-PCR was performed using SYBR Green Premix Ex Taq (Takara) on an Applied Biosystems StepOne real-time PCR System. All reactions were run in triplicate. The relative expression was calculated using the comparative 2–ΔΔCt method.
Western Blots
Protein lysates were extracted from lesioned sciatic nerve tissues or cell cultures via direct homogenization and lysed in a Laemmli sample buffer (Beyotime) containing 2% sodium dodecyl sulfate (SDS; Beyotime), Tris-HCl (52.5 mM; Beyotime), and protein inhibitors (Roche, Mannheim, Germany). The protein concentrations were determined by the Micro BCA Protein Assay Kit (Pierce, Rockford, IL, USA). Extracted protein lysates were mixed with β-mercaptoethanol, glycerin, and bromophenol blue (all from China National Pharmaceutical Group Corporation), and samples were incubated at 95°C for 5 min. Equal amounts of protein from each sample were separated on 10% SDS-polyacrylamide (Beyotime) gels and transferred to polyvinylidene fluoride membranes (Millipore, Bedford, MA, USA). Membranes were blocked in 5% nonfat dry milk (China National Pharmaceutical Group Corporation) for 2 h at 4°C, incubated overnight at 4°C with primary Astn1 antibody (1:1,000; Abcam, Cambridge, MA, USA), and then incubated with horseradish peroxidase-conjugated second antibodies (1:1,000; Pierce). Membranes were developed with an enhanced chemiluminescence reagent (Cell Signaling, Beverly, MA, USA) and exposed to Kodak X-Omat Blue Film (NEN Life Sciences, Boston, MA, USA). Measurements of band signal intensity were conducted with Grab-it 2.5 and Gelwork software (UVP, Upland, CA, USA).
Statistical Analysis
Statistical analyses were conducted using SPSS 15.0 (IBM, Armonk, NY, USA). The Student's t-test was used for statistical analysis between two groups. A value of p < 0.05 was considered statistically significant. All data are expressed as means ± SD.
Results
miR-sc3 Promoted SC Proliferation
The use of large-scale sequencing enables the identification of a large number of novel miRNAs (15,16). Their biological functions, however, remain unclear. SC dedifferentiation and proliferation normally appear at 7 days following peripheral nerve injury (13). To identify novel miRNAs that may affect phenotype modulation of SCs, therefore, the miRNAs that are highly expressed at 7 days following peripheral nerve injury have received special attention. miR-sc3 was among those highly expressed novel miRNAs and was selected as the object of the current study.
Primary SCs were transfected with synthetic miR-sc3, miR-sc3 inhibitors, and nontargeting negative controls, and then the influence of miR-sc3 on SC proliferation was investigated with a BrdU cell proliferation assay. The proliferation rate of SCs transfected with synthetic miR-sc3 was significantly increased by nearly twofold compared with that of SCs transfected with the negative control (Fig. 1A). Conversely, SCs transfected with the miR-sc3 inhibitor showed a significantly decreased proliferation rate, which was less than 50% of the control value (Fig. 1B). The results suggested that miR-sc3 could promote SC proliferation.

Effect of miR-sc3 on SC proliferation. (A) Primary SCs were transfected with synthetic miR-sc3 (miR-sc3) or the control (Con). The proliferation rate of SCs transfected with miR-sc3 was significantly increased compared with that of the control. **p < 0.01 versus control. (B) Primary SCs were transfected with a miR-sc3 inhibitor (anti-miR-sc3) or inhibitor control (Con). The proliferation rate of SCs transfected with the miR-sc3 inhibitor was significantly decreased compared with that of control. **p < 0.01 versus control.
miR-sc3 Promoted SC Migration
Transwell-based migration assay and wound-healing assay were performed to determine the effect of novel miRNAs on SC migration. Transfection of primary SCs with miR-sc3 mimic significantly promoted the migration ability of cells by more than twofold compared to transfection of primary SCs with the negative control (Fig. 2A and C), while transfection with the miR-sc3 inhibitor dramatically suppressed SC migration (Fig. 2B and D), suggesting that miR-sc3 promoted SC migration.
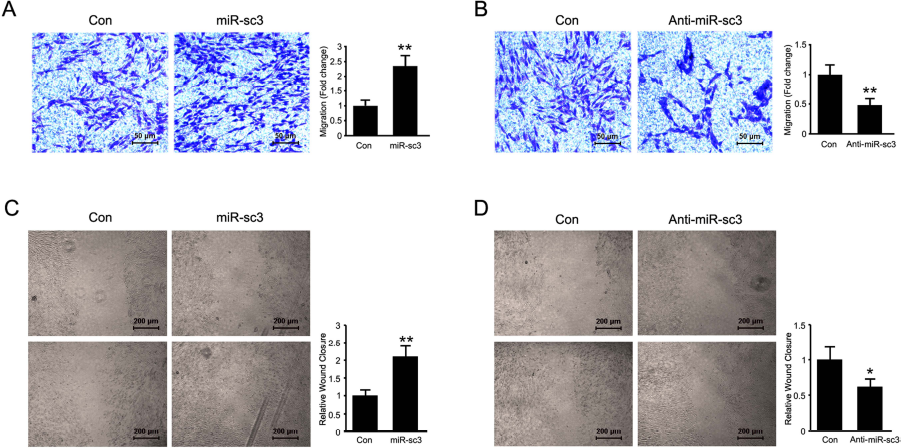
Effect of miR-sc3 on SC migration. Transwell assays (A, B) and wound-healing assays (C, D) both showed that miR-sc3 affected SC migration. (A, C) Primary SCs were transfected with synthetic miR-sc3 (miR-sc3) or the control (Con). The migration rate of SCs transfected with miR-sc3 was significantly increased compared with that of control. **p < 0.01 versus control. (B, D) Primary SCs were transfected with the miR-sc3 inhibitor (Anti-miR-sc3) or inhibitor control (Con). The migration rate of SCs transfected with the miR-sc3 inhibitor was significantly decreased compared with that of control. **p < 0.01 versus control.
miR-sc3 Negatively Regulated Astn1 by Directly Targeting Its 3′-UTR
miRNA binds to the 3′-UTR of target genes and negatively regulates the expression of target genes, thus executing its biological functions. Since miRsc3 regulates proliferation and migration of SCs, five potential target genes, Edn1, Astn1, Shroom, Cntn2, and Adam10, which were involved in cell migration and/or proliferation, attracted our special attention. The seed-matched sequences of miR-sc3 and the 3′-UTR of Edn1, Astn1, Shroom, Cntn2, and Adam10 were shown (Fig. 3A). The time-dependent expression profile of the five potential target genes indicated that the expression levels of Edn1, Shroom, Cntn2, Astn1, and Adam10 were decreased at all time points following nerve crush (Fig. 3B). This result was consistent with the finding that the expression miR-sc3 was increased after sciatic nerve transection.
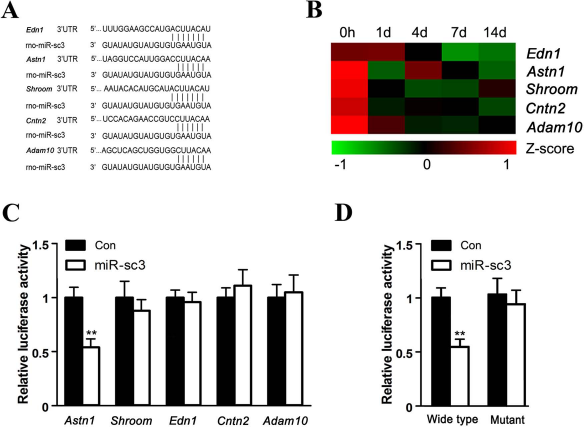
Astn1 was a direct target for miR-sc3. (A) Predicted target sites of miR-sc3 at the 3′-UTR of Edn1, Astn1, Shroom, Cntn2, and Adam10, five potential target genes for miR-sc3. (B) Heat map and cluster dendrogram of Edn1, Astn1, Shroom, Cntn2, and Adam10 from microarray in proximal sciatic nerve segment at 0, 1, 4, 7, and 14 days following nerve transection. Zero hours indicates immediately after transection. (C) The relative luciferase activity was analyzed after p-Luc-UTR vectors including 3′-UTR of Edn1, Astn1, Shroom, Cntn2, and Adam10 were cotransfected into 293T cells with synthetic miR-sc3 (miR-sc3) or the control (Con). Renilla luciferase vector was used as an internal control. **p < 0.01 versus control. (D) The relative luciferase activity was analyzed after p-Luc-UTR vectors including Astn1 3′-UTR of wild type and mutant were cotransfected into 293T cells with synthetic miR-sc3 (miR-sc3) or the control (Con). Renilla luciferase vector was used as an internal control. **p < 0.01 versus control.
The 3′-UTRs of Edn1, Astn1, Shroom, Cntn2, or Adam10 were inserted into the downstream region of the luciferase reporter gene to construct p-Luc-UTR reporter plasmids. Then, p-Luc-UTR plasmids of the five target genes, together with synthetic miR-sc3 or nontargeting negative control, were cotransfected into HEK 293T cells to identify the real target genes of miR-sc3. The greatest decrease in relative luciferase activity was detected when miR-sc3 and Astn1 Luc-3′-UTR were cotransfected into HEK 293T cells, while no significant difference was observed with miR-sc3 and the other four Luc-3′-UTR constructs (Fig. 3C). Accordingly, the mutant 3′-UTR reporter plasmid of Astn1 was constructed. The wild-type or mutant Astn1 Luc-3′-UTR construct was cotransfected with either synthetic miR-sc3 or nontargeting negative control into HEK 293T cells. When the wild-type 3′-UTR reporter plasmid of Astn1 was cotransfected with synthetic miR-sc3, the expression of luciferase was significantly decreased. The decrease in luciferase activity was abrogated in the mutant 3′-UTR reporter plasmid of Astn1 that was cotransfected with synthetic miR-sc3 (Fig. 3D). The results suggested that miR-sc3 targeted Astn1 through direct binding to 3′-UTR of Astn1.
miR-sc3 Inhibited the Translation of Astn1
To determine how miR-sc3 regulates the expression of Astn1, qPCR and Western blots were performed to determine the effect of miR-sc3 on the mRNA expression and the protein expression of Astn1, respectively. The mRNA expression of Astn1 in primary SCs was not significantly changed after the cells were transfected with synthetic miR-sc3 or the miR-sc3 inhibitor (Fig. 4A). In contrast, the protein expression of Astn1 was significantly suppressed by overexpression of miR-sc3, but was significantly enhanced by silencing of miR-sc3 (Fig. 4B, C). These results implied that miR-sc3 led to translational suppression of Astn1 rather than mRNA degradation of Astn1.

miR-sc3 inhibited the translation of Astn1. (A) The expression of Astn1 at mRNA level was analyzed by qPCR in SCs transfected with miR-sc3 mimic (miR-sc3) and miR-sc3 inhibitor (Anti-miR-sc3). GAPDH was used as an internal control. (B) The expression of Astn1 at protein level was analyzed by Western blot in SCs transfected with synthetic miR-sc3 (miR-sc3) and miR-sc3 inhibitor (anti-miR-sc3). GAPDH was used as an internal control. (C) Histogram showed relative quantitative comparisons of the Astn1 expression based on Western blots. *p < 0.05 versus respective controls (Con).
The Temporal Expression of Astn1 Was Inversely Associated with That of miR-sc3
Our previous Solexa sequencing showed that the expression of miR-sc3 was increased after sciatic nerve transection. To verify the sequencing data, in the current study, qPCR was performed to determine the expression profiles of miR-sc3 at 0, 1, 4, 7, and 14 days after sciatic nerve transection. The expression of miR-sc3 in the proximal nerve segment at 1, 4, 7, and 14 days after sciatic nerve transection was significantly increased, and the greatest expression occurred at 7 days after sciatic nerve transection compared with that immediately after sciatic nerve transection (Fig. 5A). The qPCR data were in agreement with Solexa sequencing results.
The mRNA and protein expressions of Astn1 after sciatic nerve transection were also measured. The mRNA expression of Astn1 at 1, 4, 7, and 14 days after sciatic nerve transection was significantly decreased compared with that at 0 h after sciatic nerve transection (Fig. 5B). Western blots demonstrated that the protein expression of Astn1 was significantly decreased after sciatic nerve transection (Fig. 5C, D). Importantly, the protein expression of Astn1 was decreased to the lowest value at 7 days after sciatic nerve transection, when the expression of miR-sc3 was increased to the highest value.

The temporal expression profile of Astn1 was inversely associated with that of miR-sc3. (A) The expression of miR-sc3 in the proximal nerve stump was analyzed by qPCR following sciatic nerve transection. U6 was used as an internal control. *p < 0.05, **p < 0.01 versus 0 h after nerve injury. Zero hour indicates immediately after transection. (B) The mRNA expression of Astn1 was decreased following sciatic nerve injury. **p < 0.01 versus 0 h after nerve injury. Zero hour indicates immediately after transection. (C) The protein expression of Astn1 in the proximal nerve stump was analyzed by Western blot following sciatic nerve transection. GAPDH was used as an internal control. (D) Histogram showed relative quantitative comparisons of the Astn1 expression at different time points based on Western blot. *p < 0.05, **p < 0.01 versus 0 h after nerve injury.
Astn1 Silencing Attenuated Suppressive Effects of the miR-sc3 Inhibitor on SC Proliferation and Migration
To further investigate whether the effect of miR-sc3 on SC proliferation and migration was mediated through downregulation of Astn1, stable knockdown of Astn1 was obtained by transfection of primary SCs with Astn1 siRNA (Fig. 6A), and then the cells were cotransfected with the miR-sc3 inhibitor. Consistent with the above observations (Figs. 1 and 2), the proliferation and migration of SCs transfected with the miR-sc3 inhibitor alone were significantly decreased compared to nontransfected SCs (Fig. 6B, C). Interestingly, the decreased proliferation and migration in cells transfected with the miR-sc3 inhibitor alone were significantly attenuated by cotransfection with Astn1 siRNA (Fig. 6B, C). The results suggested that Astn1 knockdown blocked the effect of the miR-sc3 inhibitor on SC proliferation and migration.

Astn1 knockdown attenuated the miR-sc3 inhibitor-induced decrease in SC proliferation and migration. (A) The mRNA expression of Astn1 in SCs transfected with Astn1 siRNA (si Astn1) was significantly decreased compared to that in SCs transfected with siRNA control (si Con). **p < 0.01 versus control. (B) The proliferation of SCs was significantly inhibited by miR-sc3 (anti-miR-sc3), but was then rescued by cotransfection with miR-sc3 inhibitors (anti-miR-sc3) and Astn1 siRNA (si Astn1). **p < 0.01. (B) The migration of SCs was significantly inhibited by miR-sc3 (anti-miR-sc3), but was then rescued by cotransfection with miR-sc3 inhibitors (anti-miR-sc3) and Astn1 siRNA (si Astn1). *p < 0.05.
Discussion
The PNS has an intrinsic capacity for self-repair and regeneration. For severe peripheral nerve injuries, however, the self-healing capacity is often insufficient for complete functional recovery. The development of clinical intervention for peripheral nerve repair is under way and would benefit from a better understanding of the cellular and molecular mechanisms underlying peripheral nerve regeneration.
Since the discovery of the regulatory roles of miRNAs in a diverse array of biological processes, increasing attention has been paid to the effects of different miRNAs on nerve regeneration. Our laboratory has identified a number of miRNAs that are differentially expressed in the dorsal root ganglia and the proximal nerve stump after sciatic nerve transection in rats (25,29). The majority of miRNAs are expressed in a tissue-specific and spatiotemporally specific manner (20). Solexa sequencing has enabled us to reveal the expression profile of a group of miRNAs in the injured nerve after sciatic nerve transection in rats (15,16). Out of this new group of miRNAs, miR-sc3 was chosen as the target of the current study.
Following sciatic nerve transection, an immediate stress response occurs immediately in the first 2 days, and then axonal regrowth is ready to happen. At that time, the active metabolism induces synthesis of numerous proteins and genes (12,13,30). Generally, miRNAs that are differentially expressed at 7 days post-nerve injury are usually associated with sciatic nerve regeneration. Meanwhile, SCs dedifferentiate and proliferate, migrate to form bands of Bünger, and promote axon myelination (12). Based on this knowledge, the current study focused on the effects of miR-sc3 on phenotype modulation of SCs. We found that overexpression (transfection with miR-sc3 mimic) and silencing (transfection with miR-sc3 inhibitor) of miR-sc3 displayed promoting and suppressing effects on proliferation and migration of primary SCs, respectively.
miRNAs negatively regulate the expression of target genes by mRNA degradation or translational repression (2,4). In the current study, we screened five potential target genes and eventually identified Astn1 as the real target. We also showed that miR-sc3 targeted Astn1 through direct binding to 3′-UTR of Astn1. The overexpression of miR-sc3 suppressed Astn1 protein expression, but failed to affect Astn1 mRNA expression, suggesting that miR-sc3 negatively regulated Astn1 by translational repression rather than mRNA degradation. It was noted that the mRNA or protein expression of Astn1 was decreased after sciatic nerve transection.
As is known, Astn1 has been extensively documented as a receptor for glial-guided neuronal migration (1,8,10). Astn1, together with Astn2 (a newly identified member of the astrotactin gene family), directly altered neuronal migration along glial fibers in the developing cerebellum (22). Accordingly, Astn1 is considered to be a susceptible gene in a variety of brain disorders, while the knowledge of Astn1-associated neuronal migration may be helpful in understanding these disorders (11). Our findings suggested that miR-sc3 regulated nerve regeneration through targeting Astn1.
Following sciatic nerve transection, the expression of miR-sc3 in the proximal nerve segment significantly increased with a peak value at 7 days after sciatic nerve transection. In contrast, the expression of Astn1, a putative target gene of miR-sc3, was significantly decreased at 1, 4, 7, and 14 days after nerve injury. In addition, the results showed that the protein expression profile of Astn1 does not mirror the mRNA expression profile of Astn1, suggesting Astn1 was posttranscriptionally regulated following sciatic nerve transection. The temporal expression of miR-sc3 was inversely associated with that of Astn1, providing further evidence that miR-sc3 negatively regulated the expression of Astn1 at the posttranscriptional level.
Considering the downregulation of Astn1 expression after sciatic nerve transection, we transfect primary SCs with a miR-sc3 inhibitor in the presence or absence of siRNA against Astn1 and observed that miR-sc3 inhibitor-induced decreases in SC proliferation and migration were significantly attenuated by cotransfection with Astn1 siRNA. This recapitulation of miR-sc3 activity by Astn1 knockdown further confirmed that Astn1 was a functional mediator of miR-sc3 in the regulation of SC phenotype modulation.
In summary, our data indicated that peripheral nerve injury induced upregulation of the expression of miR-sc3, a novel identified miRNA. Overexpression and silencing of miR-sc3 promoted and inhibited SC proliferation and migration, respectively, by targeting Astn1. The identification and functional analysis of miR-sc3 and Astn1 would contribute to understanding the biological processes following peripheral nerve injury.
Footnotes
Acknowledgments
This study was supported by National Key Basic Research Program of China (973 and 863 programs, 2014CB542202, 2012AA020502), National Natural Science Foundation of China (81130080, 31170946, 31300879), Jiangsu Provincial Natural Science Foundation (BK2012230), and Priority Academic Program Development of Jiangsu Higher Education Institutions (PAPD). The authors declare no conflicts of interest.
