Abstract
Human umbilical cord vein endothelial cells (HUVECs) secrete a number of factors that greatly impact the proliferation and differentiation of hematopoietic stem and progenitor cells (HSPCs). These factors remain largely unknown. Here, we report on the most comprehensive proteomic profiling of the HUVEC secretome and identified 827 different secreted proteins. Two hundred and thirty-one proteins were found in all conditions, whereas 369 proteins were identified only under proinflammatory conditions following IL-1β, IL-3, and IL-6 stimulation. Thirteen proteins including complement factor b (CFb) were identified only under IL-1β and IL-3 conditions and may potentially represent HSPC proliferation factors. The combination of bioinformatics and gene ontology annotations indicates the role of the complement system and its activation. Furthermore, CFb was found to be transcriptionally strongly upregulated. Addition of complement component 5b-9 (C5b-9) monoclonal antibody to the stem cell expansion assay was capable of significantly reducing their proliferation. This study suggests a complement-mediated cross-talk between endothelial cells and HSPCs under proinflammatory conditions.
Keywords
Introduction
Hematopoietic stem and progenitor cells (HSPCs) are defined by their ability to self-renew and differentiate into all cell types of the hematopoietic system. They form the basis of cell therapy for the treatment of numerous malignant and nonmalignant hematopoietic diseases (14,22). Furthermore, they represent a promising gene therapy target that can potentially treat a broad variety of human diseases (36). Major limitations of the current use of HSC transplantation include insufficient numbers and engraftment potential (37). Cultivation and ex vivo expansion of HSPCs have been proposed as a measure to increase the engraftment potential of hematopoietic grafts with insufficient stem cell numbers. Coculture with feeder cells is a promising tool for the ex vivo expansion and maintenance of HSPCs. For this purpose, numerous cell types have been used including mesenchymal stem cells (MSCs) from the bone marrow, MSC-like cells from the human placenta, endothelial cells (ECs) from porcine and human brain, as well as human umbilical cord vein endothelial cells (HUVECs) (9,24,26,31,42).
Endothelial cells and, in particular, HUVECs have been demonstrated to express interleukin-1 (IL-1), IL-5, IL-6, IL-8, IL-11, IL-15, numerous colony stimulating factors (CSF), granulocyte-CSF (G-CSF), macrophage CSF (M-CSF), and granulocyte-macrophage CSF (GM-CSF), as well as several chemokines, monocyte chemotactic protein-1 (MCP-1), chemokine (C-C motif) ligand 5 (CCL5), and growth-related oncogene protein-α (GRO-α) (4,6,19,20,28). Thus, ECs have been identified to play an important role in the proliferation and differentiation of HSPCs and may provide conditions that are superior to cytokine-supplemented liquid culture (49).
We and others have demonstrated that ECs can be successfully used to maintain and increase the number of HSPCs (5,31). Secretion of hematopoietic factors by ECs and direct cellular contact, which permits the interaction of adhesion molecules and surface-bound cytokines or chemokines with their corresponding receptors, may contribute to the maintenance of HSPCs. Furthermore, we previously demonstrated that ECs stimulated with tumor necrosis factor-α (TNF-α) induce the generation of dendritic cells from CD34+ stem cells, whereas IL-1β and IL-3 were capable of inducing the proliferation of hematopoietic and myeloid progenitors (31,32). We recently reported on the transcriptomic profiling of the hematopoietic support of interleukin-stimulated HUVECs using microarray and thereby identified several novel factors such as spermine, spermine-N′-acytyltransferase, prostaglandin E2, extracellular matrix components, micro-RNA 21, and endothelial microparticle-associated proteins that may play a crucial role in determining stem cell fate and proliferation (2). A recent study reported that ECs, posing as feeder cells, are capable of supporting the self-renewal and expansion of embryonic stem cells (ESCs) and neural stem cells (NSCs) (45,50).
The secretome is defined as a subset of the proteome, which contains all proteins actively exported out of a cell following a particular period of time and under distinct conditions. The types of proteins secreted by the cells strictly depend on the cell type and cellular state, revealing a great deal on the actual mechanisms within the cell (1). Moreover, the secreted protein can play a direct autocrine and/or paracrine role in the control and regulation of numerous biological processes such as cell proliferation, development, signaling, adhesion, and apoptosis (18,25).
Proteomic profiling of the secretome using liquid chromatography-mass spectrometry (LC-MS) is a common technique applied in the identification of secreted proteins from cell culture supernatants. Shotgun proteomics allows for the identification of proteins in a complex mixture using a combination of high-performance liquid chromatography (HPLC), and MS or tandem MS (MS/MS). In order to identify the secretome in cell culture, specific conditions must be considered, which include cell cultivation in serum- and protein-free medium, and the avoidance of interfering factors such as large amounts of bovine serum albumin (BSA), which may influence mass spectroscopic analysis.
Here, we describe and characterize the secretorial response of endothelial feeder cells on the proinflammatory cytokines IL-1β, IL-3, IL-6, with a comparison to the secretome of nonstimulated HUVECs. In order to increase our knowledge of the hematopoietic support of IL-stimulated HUVECs, proteomic profiling of the IL-stimulated HUVEC secretomes were performed via two complementary proteomic approaches.
Materials and Methods
Cord Blood, HSPC Isolation
Cord blood specimens were collected in heparin-coated syringes from full-term delivered neonates, following written consent from the mothers. Mononuclear cell fractions were isolated by gradient centrifugation, and CD34+ HSPCs were immune magnetically separated as previously described (11). The purity of isolated CD34+ cells was analyzed using flow cytometry and purity above 95% was observed.
Isolation and Culture of Endothelial Cells
HUVECs were obtained from freshly collected umbilical cords by flushing umbilical veins with 0.1% collagenase (Sigma-Aldrich, Munich, Germany), as previously described (17). Cells were cultured in Earle's M199 medium (Biochrom AG, Berlin, Germany) supplemented with 16% fetal bovine serum (FBS, Hyclone, South Logan, UT, USA), 4% human serum from healthy volunteers, 2 mM l-glutamine (Biochrom AG), 0.15 mg/ml endothelial growth factor (Intracel, Rockville, MD, USA), 0.015 mg/ml heparin (Sigma-Aldrich), and 1% fungicide (Invitrogen, Darmstadt, Germany).
Stimulation of HUVECs and Secretome Extraction
In order to prevent contamination of the HUVEC secretome with human and/or bovine serum proteins that were utilized for EC growth medium supplementation, confluent monolayer ECs from the third passage were rinsed twice with PBS (Biochrom, Berlin, Germany), followed by incubation with 10 ml human endothelial serum-free medium (GIBCO, Invitrogen, Darmstadt, Germany). Following a 3-h incubation, the serum-free medium was removed and cells were washed twice with PBS. Endothelial cells were then stimulated with 100 U/ml IL-1β, IL-3, IL-6 (all from PreproTech, Hamburg, Germany), or medium alone as a negative control (NC), in Earle's M199 medium supplemented with 0.15 mg/ml endothelial growth factor. Cell culture supernatants were collected following stimulation for 16 h. Possible cellular contamination was removed from the supernatant via a soft centrifugation step of 500×g for 10 min and sterile filtration (0.02 μm filter from Sarstedt, Nümbrecht, Germany).
In-Solution Digestion and Mass Spectrometry
Peptide mixtures were analyzed by liquid chromatography coupled to a LTQ-FT-ICR mass spectrometer (Hybrid-2D-Linear Quadrupole IonTrap-Fourier Transform Ion Cyclotron Resonance Mass Spectrometer, Thermo Electron Corp., Waltham, MA, USA). Prior to analysis, proteins were subjected to standard “in-solution digestion” procedure in which proteins were reduced with 100 mM dithiothreitol (DTT; 30 min at 56°C; Sigma-Aldrich), alkylated with iodoacetamide [45 min at room temperature (RT) in the dark; Sigma-Aldrich] and digested overnight with trypsin (sequencing Grade Modified Trypsin-Promega V5111, Madison, WI, USA). The peptide mixture was applied to an RP-18 precolumn (LC Packings, Sunnyvale, CA, USA) using water containing 0.1% trifluoroacetic acid (TFA; Sigma-Aldrich) as a mobile phase and then transferred to a nano-HPLC RP-18 column (nanoACQUITY UPLC BEHC18-Waters 186003545, Milford, MA, USA) using an acetonitrile gradient (0–60% AcN in 30 min; Merck KGaA, Darmstadt, Germany) with a flow rate of 150 nl/min and in the presence of 0.05% formic acid (Sigma-Aldrich). The column outlet was directly coupled to an ion source of LTQ-FT-MS working in the regime of data-dependent MS to MS/MS switch. Each sample was measured three times with varying m/z ranges: 300–600, 500–800, and 700–2,000. A blank run ensuring avoidance of cross-contamination from the previous sample was conducted prior to analysis of the subsequent sample.
In-Gel Digestion and Mass Spectrometry
Four milliliters of the HUVEC secretomes were concentrated via 10 kDa MWCO Amicon filters (Millipore, Billerica, MA, USA). The concentrated protein extracts were diluted in 3× Laemmli sample buffer (Bio-Rad, Munich, Germany), heated for 10 min at 70°C, and then separated on a continuous gel (4–12% Bis-Tris gradient gels, Invitrogen). Prior to digestion, each lane was split into 20 sections and gel bands were washed in 50 mM ammonium bicarbonate (Sigma-Aldrich) and 5% acetonitrile, followed by a second wash step with 50 mM ammonium bicarbonate and 50% acetonitrile at RT for 15 min. Both steps were performed twice. Following washing, the bands were dehydrated with 100% acetonitrile for 10 min. Tryptic digestion was performed in-gel with 10 μg trypsin in 50 mM ammonium bicarbonate for 3 h at 37°C. Proteins were initially subjected to electrospray ionization (ESI)-MS. For this purpose, tryptic fragments were analyzed by an Easy-nlC (Proxeon, Odense, Denmark) directly coupled to ESI-MS/MS LCQ Deca XP IonTrap MS (Thermo Finnigan, Waltham, MA, USA). Tryptic fragments of 23 μl were desalted via a NS-MP-10 Biosphere C18 (5 μm, 120 Å, 75 μm I.D.×360 μm O.D.; L = 20 mm; nanoSeparations, Nieuwkoop, Netherlands) using 0.1% v/v TFA with a flow rate of 8 μl/min. The elution gradient was established by mixing 0.1% v/v formic acid (solvent A) and 0.1% v/v formic acid in acetonitrile (solvent B) with a flow rate of 200 nl/min. The gradient was initiated with 5% v/v solvent B and increased linearly to 95% v/v of solvent B following 40 min. Data acquisition from the ESI-MS was performed throughout the LC run.
Dynamic mass was enabled, and exclusion mass with low/high was set to 3.00 m/z. Raw data were extracted by the TurboSEQUEST algorithm (Thermo Electron Corp.), and trypsin autolytic fragments and known keratin peptides were subsequently filtered. All DTA files generated by BioWorks version 3.1 (Thermo Scientific, Waltham, MA, USA) were merged and converted to Mascot generic format files (mgf).
Spectral Identification with Hybrid Approaches
Spectral identifications were carried out using hybrid approaches as previously reviewed by Nesvizhskii and Aebersold (33,34) that were combined with the “de novo sequencing of spectrums” using Peaks Studio 5.2 (www.bioinformaticssolutions.com) and a database search using Mascot and OMSSA algorithms for each mgf file. The following parameters were used for de novo sequencing: Precursor ions and product ions mass error tolerances were ±40 ppm and ±0.005 Da, respectively. Only those peptides with at least four amino acids with a confidence interval of 99.5 were considered as de novo sequenced peptides. These sequences were used in a second step for a database search versus unipro_sprot Database. Only proteins with false discovery rates (FDR) < 0.2 were accepted as identification. In addition to de novo sequencing, MS/MS ion database searches were performed with the following set of parameters: (i) database: Uniprot_sprot, (ii) taxonomy: Homo sapiens, (iii) proteolytic enzyme: trypsin, (iv) maximum of accepted missed cleavages: 1, (v) mass value: monoisotopic, (vi) peptide mass tolerance: ±0.8 Da, (vii) fragment mass tolerance: ±0.8 Da, and (viii) variable modifications: oxidation of methionine and acrylamide adducts (propionamide) on cystein. No fixed modifications were considered except the mgf files of in solution digested tryptic fragmets, for which fixed modifications of cysteine through carbamidomethylation and variable methionine oxidation were allowed. In the Mascot search, only proteins identified with scores corresponding to p < 0.05 were used. A threshold of E value of 0.05 was applied for OMSSA search. Identified proteins were verified by endothelial proteome entities of some public databases, and/or our previously revealed microarray data, and/or using quantitative real time PCR (qPCR).
Gene Ontology Analysis
Gene ontology (GO) analysis and illustration of the GO categories in respect to molecular function, biological processes, and cellular components was performed utilizing ProteinCenter” database (ProXEON, Thermo Scientific).
Quantitative Real-Time PCR
Total RNA was isolated from stimulated HUVECs obtained at the various time points via a Qiagen RNeasy Mini Kit in accordance to the manufacturer's instructions (Qiagen, Hilden, Germany). cDNA was synthesized from 2 μg of total RNA using the high-capacity RNA-to-cDNA master mix (Applied Biosystems, Foster City, CA, USA) according to the manufacturer's protocol. The cDNAs were subjected to quantitative PCR using iQ SYBR Green supermix (Bio-Rad) and detected on the iCycler iQ5 (Bio-Rad). For quantification, ΔΔCT algorithm was applied utilizing β-2 microglobulin (β2M) as a housekeeping gene. Primer sequences are presented in Supplement 1 (ftp://suppltrans:suppltrans@ftp.charite.de/Secretomics/).
Validation of CD34+ Cell Expansion From Supernatants of Interleukin-Stimulated HUVECs
In total, 1.5 ml of stem cell medium (IMDM; Invitrogen) supplemented with 20% FBS, 200 mM of l-glutamine, 50 μg/ml gentamycine (Biochrom AG), and 10 mM β-mercaptoethanol (Sigma Aldrich) containing 5×104 to 1 × 105 purified CD34+ HSPCs were cultured with 1.5 ml of each IL-stimulated HUVEC culture supernatant. HSPCs were expanded in six-well culture plates (Sarstedt) for a period of 2 weeks.
Flow Cytometry Analysis of C3d
Approximately 5×104–105 purified CD34-positive cells were expanded in 3 ml stem cell medium alone or in 1.5 ml stem cell medium supplemented with 1.5 ml of the IL-1β-or IL-3-stimulated HUVEC supernatants for 2 weeks. The cells were harvested from the six-well plates and were washed twice with PBS/BSA. Cells were incubated with anticomplement component 3d-fluorescein isothiocyanate (C3d-FITC; DAKO, Clone F_0323, Hamburg, Germany), cluster of differentiation 45-phycoerythrin (CD45-PE; BD Bioscience, Heidelberg, Germany), and corresponding isotype controls (DAKO and BD Bioscience, respectively) for 30 min at 4°C. Cells were washed with PBS/BSA. These experiments were repeated in triplicates. Flow cytometry was performed on a BD FACS Calibur with CELL QUEST software (BD Bioscience). Flow cytometric data analysis was performed using the FlowJo 7.2.5 (Tree Star, Ashland, OR, USA).
Functional Assay to Identify the Role of the Complement System on HSPC Expansion
In order to identify the potential role of the complement system in respect to its influence on HSPC expansion, two different antibodies were analyzed. Briefly, 2.5 μg/ml of monoclonal anti-C5b-9 antibody (Clone 979/96, provided by Dr. S. Bhakdi from Johannes Gutenberg-University Mainz, Germany) and 5 μg/ml of anti-C5 antibody (eculizumab, Lonza, Basel, Switzerland) were supplemented onto the cell culture expansion experiments. The expansion rates were determined for IL-1β- and IL-3-stimulated HUVEC supernatant with or without antibodies following a 2-week period. Each experiment was repeated thrice.
Statistical Analysis and Ethics
Student's t tests for paired samples were performed using Microsoft Excel 2007 (Redmond, WA, USA). Values of p < 0.05 were termed as significant. The study was approved by the ethical review board of the Charité, with the registration number EA1/012/08. All spectral raw data that were used in this study are provided on our permanent server (ftp://suppltrans:suppltrans@ftp.charite.de/Secretomics/). Proteomics identification by Mascot were also uploaded on the Pride proteomic identification database (http://www.ebi.ac.uk/pride) with the following accession numbers: 16832–16839.
Results
Identification of Secreted Proteins of IL-1β, IL-3, IL-6, and Unstimulated HUVEC Supernatants
Factors secreted by IL-stimulated HUVECs were profiled using two different proteomic approaches (in-gel and in-solution digestion) and three different bioinformatic algorithms (de novo, Mascot, OMSSA), in order to ensure robust protein identifications. Only those peptides that were termed as “confident” were used and added to a complementary list for each condition.
A total of 397 different proteins were identified in the IL-1β-stimulated HUVEC secretome via in-solution digestion (301 proteins by Mascot, 291 by OMSSA and 219 by de novo). In comparison, only 118 different proteins were identified using in-gel digestion. In total, 462 proteins were identified via both in-solution and in-gel digestion approaches for the IL-1β secretome (Supplement 2).
In IL-3-stimulated HUVECs, a total of 382 different proteins were identified in the secretome via in-solution digestion (299 by Mascot, 254 by OMSSA, and 254 by de novo). Using in-gel digestion, 127 different proteins were identified. In summary, a combination of both approaches reveals a total of 453 proteins (Supplement 3).
Via in-solution digestion, 393 proteins were identified by Mascot search, 302 proteins by OMSSA, and 231 proteins by de novo in the IL-6 secretome, resulting in a total of 397 different proteins. One hundred and forty-seven different proteins were identified using in-gel digestion. A combination of both approaches resulted in the identification of 512 proteins in the IL-6 secretome (Supplement 4).
Within the native HUVEC secretome that was used as a negative control, a total of 382 different proteins were identified using in-solution digestion (303 by Mascot, 256 by OMSSA, and 256 by de novo), whereas in-gel digestion identified 126 proteins. By combining the results from both proteomic approaches, 459 proteins were identified (Supplement 5).
Characterization of the Identified Proteins From the HUVEC Secretomes
A comparison of the native HUVEC secretome with the secretomes under proinflammatory conditions is depicted in Figure 1. From all four conditions, 231 common elements were identified, indicating a robust and reproducible identification of proteins (Supplement 6). It is well known that even the native HUVEC secretome has beneficial effects on stem cells in regards to cell proliferation, maintenance, and differentiation. The gene ontology (GO) analysis of these elements revealed a differentiation into biological processes, molecular functions, and cellular components (Fig. 2) and revealed the identification of 36 proteins associated with cell proliferation and 84 proteins associated with cell differentiation. However, within our proteomic profiling strategies, we cannot determine quantitative differences of these elements under the different conditions. Up- and downregulation of some of these proteins or posttranslational modifications might potentially play a role in stem cell expansion.
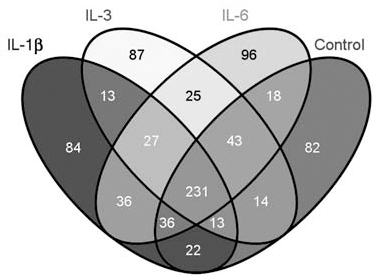
Comparative Venn analysis of IL-1β-, IL-3-, and IL-6-stimulated HUVEC secretomes with an unstimulated HUVEC secretome. Comparison of the unstimulated human umbilical cord vein endothelial cell (HUVEC) secretome with HUVEC secretomes under proinflammatory conditions via Venn analysis. In total, 231 proteins were identified in all four conditions. Eighty-four unique proteins were found in the interleukin (IL)-1β, 87 unique proteins in the IL-3, and 96 unique proteins in the IL-6 HUVEC secretome. In respect to the highest hematopoietic stem progenitor cell (HSPC) expansion, we identified 13 common elements that were only found in the IL-1β and IL-3 secretome.

GO annotations of common elements of all four stimulation conditions. GO annotation of the HUVEC secretome for molecular functions, biological processes, and cell components of 231 common elements from all four examined conditions.
The domain annotation of the 231 proteins revealed the identification of 51 secreted proteins with a signal peptide (see Table 1). Several proteins were overrepresented in the categories “cell proliferation” and “cell differentiation,” including endothelial growth factor (EGF)-containing fibulin-like extracellular matrix protein 1 (EFEMP1), granulins, thrombospondin-1, calreticulin, clusterin, and basement membrane-specific heparan sulfate proteoglycan core protein (HSPG2). However, due to exocytosis via exosomes or microparticles, proteins without a signal peptide can also be identified in the HUVEC secretome.
List of 51 Secreted Proteins Identified in All Four Conditions to Possess a Signal Peptide

Proteomic data are available in Pride Database (http://www.ebi.ac.uk/pride/) with the following accession numbers: 16832-16839. Cov., coverage in (%); UniPep number of unique peptides.
Under proinflammatory conditions, we identified 368 proteins (Supplement 7). Figure 3 illustrates the GO analysis of these proteins. Eighty-two secreted proteins were identified to possess a signal peptide, whereas only those with at least one unique peptide (n = 57) are presented in Table 2. Within this group, 17 proteins were recognized as being related to cell proliferation and 32 proteins to cell differentiation, which includes immunoglobulin superfamily member 10, jagged-1, angiotensin-converting enzyme isoform 1, and tumor necrosis factor receptor superfamily member 11A.

GO annotation of proteins identified only under proinflammatory conditions. Gene ontology (GOP) analysis of the HUVEC secretome for molecular functions, biological processes, and cell components of the 368 proteins that were identified only under proinflammatory conditions.
List of 57 Secreted Proteins With a Signal Peptide Identified Only Under Proinflammatory Conditions

Proteomic data are available in Pride Database (http://www.ebi.ac.uk/pride/) with the following accession numbers: 16832-16839. Cov., coverage in (%); UniPep, number of unique peptides.
As previously demonstrated, the cell culture supernatants from HUVECs exhibit great hematopoietic support for HSPCs. The beneficial effects and the highest HSPC expansion has been found in IL-1β- and IL-3-stimulated HUVEC secretomes. From this perspective, 13 common elements under both conditions that were observed to be highly interesting are presented in Table 3.
List of 13 Common Elements Identified in IL-1β- and IL-3-Stimulated HUVEC Secretomes
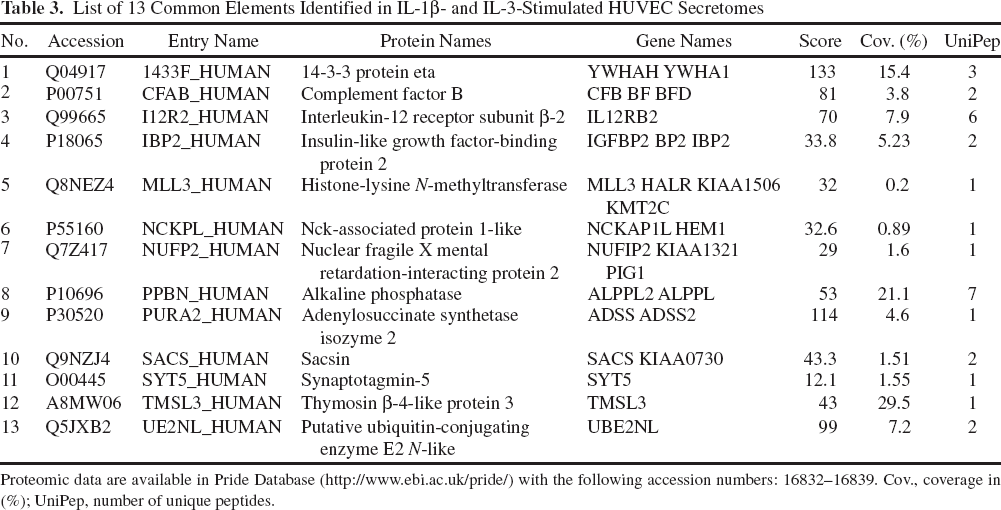

Proteomic data are available in Pride Database (http://www.ebi.ac.uk/pride/) with the following accession numbers: 16832–16839. Cov., coverage in (%); UniPep, number of unique peptides.
Quantitative Gene Expression Profiles
Figure 4 demonstrates gene expression profiles of several factors (n = 8) that we assumed may play crucial roles in HSPC support. Some have been identified as common elements of the IL-1β and IL-3 secretomes. These include complement factor B (CFB), insulin-like growth factor binding protein 2 (IGFBP2), myeloid/lymphoid or mixed-lineage leukemia 3 (MLL3), nuclear fragile X mental retardation protein interacting protein 2 (NUFIP-2), and thymosin β-4-like protein 3 (TMSL3). Jagged-1 was identified in all pro-inflammatory conditions, whereas complement factor C3 was found in all secretomes.
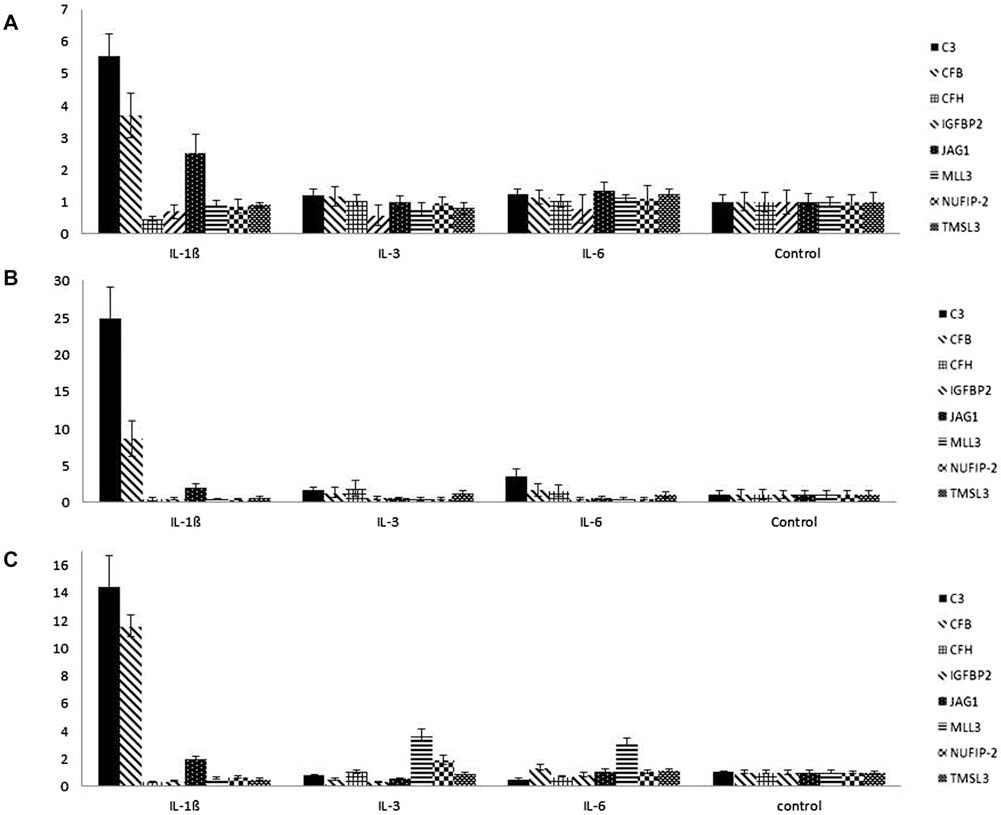
Gene expression profiles of eight genes following 4, 8, and 16 h of IL stimulation. Gene expression profiles of complement factor C3, complement factors B and H (CFB and CFH), insulin-like growth factor-binding protein 2 (IGFBP2), Jagged-1 (JAG1), myeloid/lymphoid or mixed-lineage leukemia 3 (MLL3), nuclear fragile X mental retardation protein interacting protein 2 (NUFIP2), and thymosin β-4-like protein 3 (TMSL3) are presented here over a time course of (A) 4 h, (B) 8 h, and (C) 16 h following IL stimulation in comparison to unstimulated HUVECs at each time point as a corresponding control. Upregulation of C3, CFB, and Jagged-1 were detected in IL-1β-stimulated HUVECs at each time point, but not in IL-3- or IL-6-stimulated HUVECs. MLL-3 was found to be upregulated in IL-3- and IL-6-stimulated HUVECs following 16 h of stimulation, but not in IL-1β-stimulated HUVECs.
We detected a strong upregulation of CFB in IL-1β-stimulated HUVECs in comparison to the corresponding control over the time course of 4, 8, and 16 h. Interestingly, although CFB is a common element of the IL-1β and IL-3 secretomes, no upregulation of this factor can be observed in IL-3-stimulated HUVECs. Complement factor C3 is the most important factor of the complement system in the formation of both classical and alternative C3 and C5 convertases. Correspondingly to CFB, we identified the upregulation of C3 in IL-1β-stimulated HUVECs but not in IL-3, IL-6, or in the unstimulated HUVECs. In order to address whether complement activation may be inhibited by complement factor H (CFH), we also investigated the gene expression profiles of CFH. However, no upregulation of CFH was observed.
Jagged-1 was identified in IL-1β, IL-3, and IL-6 secretomes, but not in native HUVECs. However, Jagged-1 was only upregulated in IL-1β-stimulated HUVECs. Other genes investigated via qPCR demonstrated no differential gene expression.
The Role of the Complement System
The gene expression profiles of IL-1β-stimulated HUVECs in respect to complement factors and the identification of different complement proteins in the HUVEC secretomes raised much interest and resulted in further analysis of the role of the complement system. C3d is a marker for previously occurring complement activation. For this reason, we investigated surface bound C3d on expanded HSPCs. As depicted in Figure 5, HSPCs, expanded for 2 weeks in stem cell medium supplemented with equal volumes of IL-1β or IL-3-stimulated HUVEC supernatants, revealed a strong increase of C3d fragment on their surface (87.97 ± 4.75% and 86.17 ± 9.44%, respectively). These data suggest that complement activation and thus the formation of a sublytic membrane attack complex (MAC, C5b-9) may play a crucial role in respect to HSPC expansion.

Flow cytometry analysis of surface C3d on expanded HSPCs. The level of C3d expression on expanded HSPCs analyzed by flow cytometry is depicted here. HSPCs expanded with both the IL-1β- and IL-3-stimulated HUVEC supernatants revealed a strong increase of C3d expression of 87.97 ± 4.75% and 86.17 ± 9.44%, respectively.
We further investigated a potential role of the complement system in regards to HSPC expansion, via supplementation of HSPC with two different antibodies C5 and C5b-9. Interestingly, anti-C5b-9 revealed an inhibition of HSPC expansion (Fig. 6). A 1.42-fold decrease of HSPC expansion in IL-1β and a 1.98-fold decrease in IL-3, respectively, can be observed due to the supplementation of anti-C5b-9 (p = 0.0229 and p = 0.0433, respectively). However, no significant differences were observed via anti-C5 antibody supplementation.
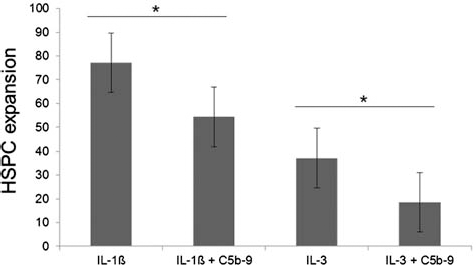
Anti-C5b-9 significantly inhibits HSPC expansion. The expansion rates of HSPCs cultured with both IL-1β- and IL-3-stimulated HUVEC supernatants are demonstrated here. The monoclonal antibody directed to C5b-9 revealed the inhibition of HSPC expansion in IL-1β- and IL-3-stimulated HUVEC supernatants (p = 0.0229 and p = 0.043, respectively). These data strongly suggest that the complement system and, in particular, a sublytic C5b-9 play a crucial role in HSPC expansion. Error bars representing SEM, as SD is unrepresentative due to deviations of the different biological replicates (n = 3) used for this experiment.
Discussion
Endothelial cells represent an instructive vascular niche that determines HSPC fate via the release of specific paracrine growth factors called angiocrine factors (8,16). With the main focus on clinical scale expansion to improve the engraftment and reconstruction of hematopoiesis for HSPC transplantation, we and several other groups used ECs as feeder cells in coculture with HSPCs to effect their proliferation and differentiation (16,31,42,49,50). Gene expression profiling of ECs using microarray technology has been used as an indirect means to identify EC-derived angiocrine factors (2,9). Due to the genomic complexity of ECs, it was difficult to elucidate exact information concerning the mechanisms and factors that are responsible for the intercellular communication between ECs and HSPCs. Therefore, we performed qualitative proteomic profiling of the EC secretome in order to directly identify the secreted angiocrine factors.
We identified 231 commonly secreted proteins from all four EC secretomes (Supplement 6). The detection of these proteins in all four conditions indicates a robust and confident protein identification. However, quantitative differences of these proteins may differ between the individual conditions but cannot be detected with our proteomic approach. As the secretome is defined as the subset of proteins actively secreted from the cells under certain conditions, it would be expected that most of the proteins contain a signal peptide according to genomic annotation. However, only 51 of the 231 proteins are secreted proteins (Table 1), and the majority of the proteins were assigned to be intracellular compartments. This phenomenon can be explained with multigranular exocytose of Weibel–Pallade bodies (WPBs), also called caveolae. They can be intracellularly fused with secretory pods and nanovesicles in ECs (12,47) and contain a large number of signaling molecules, enzymes, and receptors and release their cargo constitutively and in response to physiological stimuli such as trauma, hypoxia, and mediators of inflammation (30). In coherence with this unconventional secretion pathway, we identified 48 proteins, also in the nonstimulated EC secretome, which were enriched under the GO class “cytoplasmic vesicle.” These include structural protein of vesicular transport such as actin, cofilin-1 and -2, annexin A2, adaptor protein 2 (AP-2), complex subunits (α-1, -2 and b), heat shock protein 90 (HSP-90) and HSP-60, 14-3-3 protein, and α-actinin-1 and -4.
Recently, the mitogen-activated protein kinase (MAPK) and v-akt murine thymoma viral oncogene homolog (AKT; protein kinase Ba) signaling pathways in ECs were identified as crucial mechanisms in secreting factors that regulate both the expansion and differentiation of HSPCs (8). The activation state of ECs determines the balance between HSPC self-renewal and differentiation within the vascular endothelial niche (3). Previously, we demonstrated that the proinflammatory cytokines IL-1β- and IL-3-stimulated EC secretomes induce the proliferation and differentiation of myeloid precursor cells from HSPCs more effectively, while IL-6-stimulated ECs only demonstrate a benefit in HSPC survival (31). Both IL-1β and IL-3 activate the MAPK and AKT signaling pathways (21,27,41). In this respect, secreted proteins from ECs under proinflammatory conditions are optimal in identifying factors and mechanisms that are responsible for the regulation of the expansion and differentiation of HSPCs.
Within this context, we identified 368 proteins secreted only under proinflammatory conditions (Supplement 7). In comparison to the unstimulated EC secretome, numerous proteins were identified to be closely classified under vesicular transport. These include coatomer subunits (COP a, b, and g), clathrin, dynein, AP-2, AP-3, arfaptin-2, synaptotagmin-5, CD109, and CD93, which indicate a differentially regulated shedding of endothelial vesicles under proinflammatory conditions. These findings are in accordance with the literature (23). Several groups demonstrated that HSPCs can be reprogrammed with the horizontal transfer of RNAs and transcription factors (TFs) using a microvesicle (MV)-mediated pathway (38). In coherence with these studies, we also identified TFs in the EC-secretome such as transcription factor 7-like 2 (T-cell specific, HMG-box) (TF7L2), transcription factor Dp-1 (TFDP1), zinc finger protein 281, human immunodeficiency virus type I enhancer-binding protein 3 (HIVEP3), homeodomain protein Iroquois (IRXA1), and mesoderm posterior protein 1. Although 39 of the 368 identified proteins demonstrated an RNA binding function in accordance to GO analysis, which may be responsible in the preservation of RNA in the secretome. Pilot studies are currently underway analyzing the influence of endothelial MVs on the proliferation and differentiation of HSPCs.
As IL-1β- and IL-3-stimulated EC secretomes demonstrated the highest expansion rate of HSPCs, commonly secreted proteins under these conditions are of great importance in revealing both the responsible factors and associated mechanisms of hematopoietic support. We identified 13 proteins that were commonly secreted only under these conditions (Table 3). In order to evaluate quantitative differences between the different conditions, quantitative real-time PCR (qPCR) was performed and revealed gene expression profiles of several of these proteins (Fig. 4). Most of these genes were not found to be differentially expressed between the two conditions. This phenomenon may be explained by an alternate means of posttranscriptional regulation, in which these factors are translated and secreted by ECs only following IL-1β and IL-3 stimulation. On the other hand, it cannot be excluded that these factors may also be secreted from IL-6 or nonstimulated ECs nor that their detection may be prevented via high-abundant proteins.
The complement system belongs to the innate immune system and is primarily a defense mechanism against invading pathogens. In this study, several proteins in the EC secretome were identified as belonging to the complement system. These include complement component 1q receptor (C1qR), complement C4-A, Igk chain C region, Igλ-1-3, and 6 chain C regions, clusterin and CD59, also termed as MIRL (membrane inhibitor of reactive lysis). Gene expression analysis of complement factor B (CFB) and complement factor 3 (C3) revealed their upregulation over the time course of 4, 8, and 16 h following IL-1β stimulation. CFB is a common element of the IL-1β and IL-3 secretome, whereas C3 was identified in all four conditions. C3 plays a central role in complement activation and is a part of the complement convertases. CFB is able to activate the complement system via the alternative pathway. We further investigated the gene expression profile of complement factor H (CFH), an inhibitor of the alternative pathway. However, no upregulation was observed. The identification of complement proteins in the EC secretomes and the upregulation of CFB and C3 indicate that the complement system and its activation might be associated with hematopoietic support.
A number of studies have reported on the involvement of the complement system in early development, stem cell commitment, differentiation, and homing (29). HSPCs express the classical receptor for C3 cleavage fragments such as C3a receptor (C3aR) and complement receptor 3 (CR3), which provides evidence that a cross-talk between ECs and HSPCs via the complement system and/or its activation state is possible (39,40,48).
In order to control the proximal activation of the complement system, we analyzed the presence of C3d on the surface of expanded HSPCs. C3d, in contrast to C3a, is a C3 cleavage fragment with a long half-life. A strong increase of C3d molecules was identified on their surface, suggesting an activation of the complement system under proinflammatory conditions. A previous study demonstrated the release of stable C3 convertase in the extracellular matrix by IL-1β-stimulated HUVECs, showing that activation of distal complement components only require extracellular matrix (ECM) exposure independently from ECs (15). To further investigate the role of distal complement activation in the hematopoietic support of the EC secretome, an Eculizimab (anti-C5) and a monoclonal antibody directed to C5b-9 was supplemented during HSPC expansion. A decrease in HSPC expansion in IL-1β and IL-3 was observed due to anti-C5b-9 but not with Eculizumab. Stem cells were cultivated in a medium containing FBS. Eculizumab is human specific and a proliferation inhibiting effect was only observed with anti-C5b-9. However, this reversely indicates that both a sublytic C5b-9 is present and has beneficial effects on HSPC expansion. We hypothesize that the generation of active C3 convertases via IL stimulation of HUVECs—which was subsequently transferred into another expansion medium containing FBS—results in a distal complement activation leading to the generation of a sublytic C5b-9 complex, which increases HSPC proliferation. The C5b-9 deposition by IL-1β-stimulated HUVECs has been demonstrated elsewhere (10). This HSPC expansion is significantly reduced by anti-C5b-9. In coherence with our results, the impact of the sublytic C5b-9 complement complex in relation to cell proliferation has been demonstrated by Rus and colleagues (43). Moreover, sublytic C5b-9 has been shown to trigger several signal transduction pathways such as janus kinase 1 (JAK1) and signal transducer and activator of transcription 3 (STAT3), analogously to those induced by growth factors and cytokines (35). Further studies investigating the role of the complement system in hematopoetic support are required.
There are various reports on the investigation of the EC secretome in other biological contexts such as shear stress, atherosclerosis, and senescence (7,13,44,46). Recently, Tunica and coworkers identified 314 proteins from nonstimulated ECs using nano-LC-MS/MS following free flow electrophoretic fractionation (46). Similarly, Burghof and Schrader identified 390 proteins under nonstimulated conditions, whereas 327 and 507 proteins were identified under laminar and oscillatory shear stress, respectively (7). In accordance with these reports, we identified a total of 827 different proteins from three proinflammatory conditions and an additional nonstimulated control. Our study represents the most comprehensive profiling of proteins, which were also identified in other studies, validating our proteomic strategy and robust bioinformatic processing.
In conclusion, we present here a detailed proteomic profile of the hematopoietic support of the endothelial secretome using two different proteomic approaches. Angiocrine factors of the vascular niche, which are responsible for the beneficial effects associated with expansion and differentiation, have not yet been reported. This is the first study representing potential factors and/or mechanisms of the vascular niche, which may determine stem cell fate. Possible factors and mechanisms such as microvesicle and/or complement mediated cross-talk between ECs and HSPCs have been presented and discussed here. However, the evaluation of individual factors that may increase HSPC expansion cannot recapitulate the complexity of in vivo hematopoiesis.
Footnotes
Acknowledgments
This study was partially funded by European Social Fund (ESF). G.B. designed the study, initiated the experiments, performed the bioinformatic analysis, interpreted data and wrote the manuscript; J.K.-M. performed cell culture and flow cytometry experiments and wrote the manuscript; J.D. and O.K. performed MS analysis; V.S. and M.A.-S. provided experimental assistance; S.K.-M. provided experimental assistance and edited the manuscript; A.S. conceived and helped in the design and coordination of the study and drafted the manuscript. The authors declare no conflicts of interest.
