Abstract
Human amniotic mesenchymal cells (HAM cells) are known to contain somatic stem cells possessing the characteristics of pluripotency. However, little is known about the biology of these somatic cells because isolated HAM cells from amniotic membrane have a limited lifespan. To overcome this problem, we attempted to prolong the lifespan of HAM cells by infecting retrovirus encoding human papillomavirus type16E6 and E7 (HPV16E6E7), bmi-1, and/or human telomerase reverse transcriptase (hTERT) genes and investigated their characteristics as stem cells. We confirmed the immortalization of the four lines of cultured HAM cells for about 1 year. Immortalized human amnion mesenchymal cells (iHAM cells) have continued to proliferate over 200 population doublings (PDs). iHAM cells were positive for CD73, CD90, CD105, and CD44 and negative for CD34, CD14, CD45, and HLA-DR. They expressed stem cell markers such as Oct3/4, Sox2, Nanog, Klf4, SSEA4, c-myc, vimentin, and nestin. They showed adipogenic, osteogenic, and chondrogenic differentiation abilities after induction. These results suggested that immortalized cell lines with characteristics of stem cells can be established. iHAM cells with an extended lifespan can be used to produce good experimental models both in vitro and in vivo.
Introduction
The amnion is a fetal-origin tissue deriving from the inner cell mass of the blastocyst and is composed of a single layer of epithelial cells (human amnion epithelial cells, HAE cells) on a thicker basement membrane and a collagen spongy layer containing mesenchymal cells (human amnion mesenchymal cells, HAM cells). Thus, HAE cells and HAM cells have the potential to differentiate into various tissues (15, 21, 28).
Several studies have shown that mesenchymal stem/ progenitor cells derived from the amnion have the potential to differentiate into neurogenic (16, 17) and mesodermal lineages including osteogenic, chondrogenic, and adipogenic in vitro (2, 7, 8, 16, 24). In vivo studies have indicated that HAM cells are also capable of differentiation into cardiomyocytes (6, 30), neural precursors (3), and skeletal muscle (4). In addition to these properties, HAM cells can be obtained without ethical problems and invasive procedures because they are discarded after parturition and they express immunosuppression properties (24).
These properties are a potentially useful and noncontroversial source of cells for transplantation and regenerative medicine. However, HAM cells, which are usually isolated in the third trimester, undergo growth restriction and have a limited lifespan (16). The cells reach senescence because of DNA damage or shortened telomeres (23), implying that it would be difficult to obtain sufficient stable cells for cell transplantation therapy and regenerative medicine. To resolve these problems, we attempted to establish several strains of HAM cells without a lifespan limit by transfecting with retrovirus carrying the human telomerase reverse transcriptase (hTERT), B lymphoma Mo-MLV insertion region 1 homolog (Bmi-1), a polycomb-group gene, and human papilloma virus type 16 (HPV16) E6 and/or E7 genes (19, 20). Both retinoblastoma/cyclin-dependent kinase 4 inhibitor A (Rb/p16INK4a) inactivation with E7 and telomerase activation with E6 are required to extend the lifespan of human epithelial cells (10). Bmi-1, a c-myc cooperating oncogene in murine lymphomas, reduces expression of p16INK4a, stimulates cell proliferation (9), and is required for maintenance of self-renewing hematopoietic stem cells (12, 14). This method proved highly efficient in extending the lifespan of HAM cells.
In the present study we attempted to establish immortalized HAM (iHAM) cell lines and investigated those that had the characteristics of stem cells, those which expressed pluripotency markers and proliferation ability, as well as differentiation capabilities.
Materials and Methods
Isolation of HAM Cells
The study and the use of the amnion membrane were approved by the Research Ethics Committee of Toyama Medical and Pharmaceutical University Hospital. The amniotic membrane was peeled mechanically from the chorion of a placenta obtained with informed consent from a patient undergoing cesarean section. As previously described (24), HAM cells were isolated by sequential trypsin and collagenase digestion.
First, the epithelial cells were more than 98% removed from the amnion membrane using trypsin treatment four times. Mesenchymal cells were then isolated. Tissue pieces were placed in Dulbecco's modified Eagle's medium (DMEM; Sigma-Aldrich, St. Louis, MO) containing collagenase (0.75 μg/ml) and DNase (0.075 μg/ml) and were incubated at 37°C for 60 min with shaking (100 rpm). The dispersed mesenchymal cells were collected by filtration of the mixture through gauze and centrifugation. The yield of mesenchymal cells was approximately 1 × 106 cells/g tissue.
Infection of Retrovirus Constructs and Establishment of Cell Line
HAM cell lines were stably transfected with HPV16 E6/E7, human TERT, and/or human bmi-1 genes, as described previously (19). Construction of the destination vectors pDEST-CLXSN and pDEST-CMSCV puro and the expression vectors pCMSCVpuro-16E7 and pCLXSNhTERT was performed. The HPV16 E6E7 segment and human Bmi-1 cDNA were similarly cloned into the destination vectors. Briefly, after cloning segments of HPV16 E6E7 (16E6E7) and a splice donor site-mutant version of HPV16 E6 (16E6SD) (18) into pDONR201 (Invitrogen), these segments were recombined into retroviral vectors by LR reaction (Invitrogen) to generate pCMSCVpuro-16E6E7 and pCMSCVpuro-hBmi-1. Construction of the destination vector pDEST-CL-SI-MSCVpuro (designated as pSI-CMSCVpuroDEST previously), for retroviral expression of short hairpin RNA (shRNA), pCL-SIMSCVpuro-p53-shRNA (designated as pSI-CMSCVpuro-p53Ri previously), and the entry vector pENTR-H1R-stuffer were as described previously (20). To generate p16shRNA expression vectors pSI-CMSCVpuro-H1R-p16shRNA1, 6 and 8, 5′-AAC GCA CCG AAT AGT TAC G-3′, 5′-GGA CGA AGT TTG CAGGGG A-3′, and 5′-GCC CAA CGC ACC GAA TAG TTA CGGTC-3′, respectively, were chosen as the targeted sequences. Production of recombinant retro-viruses was as described earlier (10). Infected cells were selected in the presence of 0.5 μg/ml puromycin or 50–200 μg/ml G418. For combinations of retroviral infections, cells were first transduced with Bmi-1 or p16INK4a shRNA and then with hTERT.
The four stably transduced cell lines with an expanded lifespan were designated as iHAM3 542 (CSII-CMV-16E6E7;E6E7), iHAM3 542-344 (CSII-CMV-16E6E7 and MSCVpuroACChTERT;E6E7/T), iHAM4 136-344 (LXSN-hbmi-1 and MSCVpuroACChTERT;bmi/T), and iHAM4 344-96 (MSCVpuroACChTERT and LXSN-16E6E7;T/E6E7) (Table 1).
Primer Sequences Specific to Gene mRNA and Annealing Temperature and Cycles Use in the Gene Expression of Undifferentiated HAM and iHAM Cells

Cell Culture and Proliferation
The HAM cells and iHAM cells were cultured in DMEM with high glucose (Sigma-Aldrich) supplemented with 10% fetal bovine serum (FBS) and 1% antibiotic–antifungal solution (GIBCO BRL, Grand Island, NY). The population doubling level (PDL) at each passage was calculated as [log (cell number at passage) – log (cell number seeded)] / log2. The population doubling time was calculated from the slope of the linear regression curve of cell number versus time of treatment over a 72-h period. Based on the formula, the cell population doubling time (TD) =
Immunofluorescence
The cells were fixed in −20°C acetone for 10 min and then air dried for the subsequent immunocytochemical reactions. After blocking for nonspecific reaction with BLOCK ACE (Dainippon Pharmaceutical, Osaka, Japan) for 45 min, cells were incubated in the following primary antibodies diluted 1:200 in PBS containing 1% bovine serum albumin (BSA) and 0.1% Triton X-100 for 24 h at 4°C: octamer binding transcription factor 3/4 (Oct3/4), Nanog, Krüppel-like factor 4 (KLF4), stage-specific embryonic antigen 4 (SSEA4), α-smooth muscle (α-SM) actin, nestin, c-myc (all from Santa Cruz Biotechnology, Inc., CA, USA), sex-determining region Y box 2 (Sox2), anti-human Lin28 (R&D System, Inc., Minneapolis, MN, USA), vimentin (Dako, Glostrup, Denmark), and cytokeratin 18 (CK18; Progen, Heidelberg, Germany). After washing, the cells were incubated with biotinylated second antibody (Nichirei, Tokyo, Japan) for 1 h at room temperature, followed by incubation with fluorescein isothiocyanate (FITC) or Texas red-conjugated streptavidin (Vector Laboratories, Inc., Burlingame, CA) for 30 min at room temperature. The nuclear staining was performed with Hoechst 33342 (Dojindo Laboratories, Japan).
All samples were visualized using fluorescent microscopy (Leica DM/RBE, Wetzlar, Germany), and the figures were recorded with a DP-70 and analyzed using its control software (Olympus, Japan).
Flow Cytometry Analysis for Cell Surface Markers
HAM cells and iHAM cells were trypsinized and resuspended in 10% FBS/DMEM. After 4% paraformaldehyde (PFA) fixation, cells were resuspended in 0.5% BSA/0.01M PBS solution and incubated for 1 h at room temperature with the following FITC-conjugated primary antibodies (Beckman Coulter): CD105, CD34, CD14, CD44, human leukocyte antigen (HLA)-DR (mouse), and phycoerythrin (PE)-conjugated antibodies (BD Bioscience Pharmingen, Vienna, Austria); CD73, CD45, and phycoerythrin-Cyanin 5.1 (PC5)-conjugated CD90 (Beckman Coulter). Mouse IgG1-isotyped antibodies conjugated to FITC or PE or PC5 (Beckman Coulter, France) served as the negative control. Samples were analyzed on a BD FACSCalibur™ system (BD Biosciences). 3 × 104 events were acquired in a histogram figure with Cell Quest software (BD Biosciences). Data were further analyzed with WinMDI software ver.2.9 (The Scripps Research Institute).
RT-PCR
For undifferentiated cells: Total RNA was extracted from 5 × 106 fresh HAM cells (fHAM cells), p2 HAM cells (second generation of fHAM cells), and iHAM cells using ISOGEN RNA extraction buffer (Nippon Gene, Tokyo, Japan). cDNAs were synthesized from 1 mg of total RNA using an Omniscript RT Kit (QIAGEN, Tokyo, Japan). cDNA was subjected to polymerase chain reaction (PCR) using a Taq PCR Core Kit (QIAGEN, Tokyo, Japan) (25 ml). Total RNA was extracted using an RNase Mini Kit (QIAGEN, Tokyo, Japan). DNase I digestion of RNA was performed prior to the reverse transcriptase reaction.
For differentiation cultured iHAM cells: After differentiation, RT-PCR was performed on purified RNA using the One-Step RT-PCR kit (QIAGEN, Inc., Valencia, CA) as per the manufacturer's recommendations. After the reaction, the PCR products were subjected to 2% agarose gel electrophoresis with ethidium bromide. Digital images were captured by LAS-3000 (Fujifilm, Tokyo, Japan) and Multi-Gauge v3.0 software (Fujifilm, Tokyo, Japan). The primer sequences and optimal annealing temperatures and cycles are listed in Table 1.
Sphere Formation
The cells of fHAM cells and iHAM cells were cultured on ultra low attachment dishes in 24 wells for 2 weeks. The cell concentration was a density of 1.0 × 106 cells per well.
Tumorigenicity
Cells of each cell line were examined for tumorigenicity by injection of 1.0 × 105 cells per site into the rear leg muscles (
Differentiation–Induction Experiments
Adipogenic Differentiation
Cells were seeded into two-well Lab-Tek chamber slides (Nalge Nunc International, USA) at 5 × 103 cells/cm2 in MSCGM™ (Mesenchymal Stem Cell Growth Medium; Lonza, Walkersville, MD, USA). The cells were exposed to Adipogenic Induction Medium (AIM, Lonza, Walkersville, MD, USA) followed by 1–3 days of culture in Adipogenic Maintenance Medium (AMM, Lonza) for 3 weeks with the medium changed every third day. Cells were fixed with 4% PFA and stained with Oil red O (Nakarai Tesque, Kyoto, Japan), which was utilized to visualize fat drops (18).
Chondrogenic Differentiation
Cells were resuspended into 15 ml polypropylene culture tubes at a concentration of 5.0 × 105 cells/ml and treated with hMSC differentiation BulletKit-Chondrogenic (Lonza) containing chondrogenic basal medium, insulin transferrin selenous acid, BSA and linoleic acid (ITS+) supplement, dexamethasone, ascorbate, sodium pyruvate, proline, penicillin/streptomycin, l-glutamine, and 10 ng/ml transforming growth factor β3 (TGF-β3; R&D Systems, Inc., Minneapolis, MN), prepared according to the manufacturer's protocol (Lonza). Cells were then centrifuged at 150 ×
Osteogenic Differentiation
Cells plated in the two-well chamber slide were treated by Differentiation BulletKit-Osteogenic (Lonza). Cells received full medium changes every 3–4 days for 3 weeks. To identify osteogenic differentiation, cells were fixed with 4% PFA, and then mineralization was observed by staining with Alizarin red S (Fisher Scientific, Pittsburg, PA) and alkaline phosphatase activity was observed using Vector Red Alkaline Phosphatase Substrate Kit I (Vector Laboratories, Burlingame, CA) (5).
Results
Establishment of HAM Cells with an Extended Lifespan
To extend the lifespan of HAM cells, four different types of cells were established by combination of bmi-1, E6E7, and/or hTERT genes. Cells transduced with E6/E7 were designated iHAM3 542(E6E7); cells transduced with E6/E7 and hTERT genes were named iHAM3 542-344(E6E7/T); cells transduced with bmi-1 and hTERT were designated iHAM4 136-344(bmi/T) and iHAM4 344-96(T/E6E7). In these cell lines, the rates of transfected genes were different, even if the same genes were induced, because of the use of different promotors.
iHAM cells were spindle shaped, like mesenchymal stem cells. These cells were expandable over 200 population doublings (PDs) by passage over 50 generations (Fig. 1). The cell growth curve of iHAM cells exhibited an S-curve pattern with a shorter lag period, and they stopped dividing after reaching confluence. The doubling times of each iHAM cell population were observed. iHAM E6E7 was 35.45 ± 3.19 h, iHAM3 E6E7/T was 25.28 ± 4.67 h, iHAM4 bmi/T was 34.69 ± 5.58 h, and iHAM4 T/E6E7 was 29.47 ± 4.09 h. The doubling times of iHAM3 E6E7/T and iHAM4 T/E6E7 were slightly shorter than those of iHAM3 E6E7 and iHAM4 bmi/T. The names of cell lines are summarized together with the names of transfected genes in Table 2.

Proliferation of immortalized human amniotic mesenchymal cells (iHAM cells). All cell lines of established iHAM cells were cultured for at least 50 generations (>350 days) and the population doubling level (PDL) was over 150 population doublings (PDs). The immortalized cell lines with combinations of genes transferred were (a) iHAM3 542 transduced with human papillomavirus E6/E7, (b) iHAM3 542-344 transduced with E6/E7 and human telomerase reverse transcriptase (hTERT), (c) iHAM4 136-344 transduced with hTERT and E6/E7, (d) iHAM4 344-96 transduced with B lymphoma Mo-MLV insertion region 1 homolog (bmi-1) and hTERT. No differentiation was seen between cell lines.
Immortalized Human Amniotic Cell Lines

Tumorigenicity
When iHAM cells were transplanted into the testis, liver, and muscle of nude mice at a concentration of 1 × 105 cells/part, respectively, no tumors were seen during the monitoring period of more than 6 months.
Surface Markers Analysis of HAM Cells
Surface markers of four cell lines, iHAM3 542, iHAM3 542-344, iHAM4 344-96, and iHAM4 136-344 were evaluated by flow cytometric analysis (Fig. 2 and Table 3). All of the iHAM cells were positive for CD105 (endoglin), CD90 (Thy-1), CD73, and CD44 [phagocytic glycoporotein-1 (Pgp-1)/ly-24] and negative for CD34, CD45 (leukocyte common antigen), CD14 (a marker for macrophage and dendritic cells), and HLA-DR. All types of iHAM cells displayed similar expression patterns of surface antigen, and the rates of positive cells indicated high values of 90% or more in each cell type. Compared with immediately isolated fresh HAM; fHAM cells and passage 2 HAM cells; p2 HAM cells, p2 HAM cells expressed higher values than fHAM cells for all stem cell markers. Interestingly, p2 HAM displayed CD45 15.3% and HLA-DR 16.7%, although these were not observed in fHAM and iHAM (Fig. 2).

Surface markers of iHAM cells lines by FACS analysis. Top: freshly isolated HAM cells (fHAM cells); bottom: one of the iHAM cell lines (iHAM3 542). Mesenchymal stem cell markers CD105 and CD90 were expressed positively in the fHAM cells, but CD73 and CD44 reacted little. iHAM cells expressed all mesenchymal stem cell markers, CD105, CD90, CD73, CD44 with at least 90% positivity. Few hematopoietic stem cell markers were expressed in either fHAM cells or iHAM cells. HLA, human leukocyte antigen.
Analysis of Pluripotency of iHAM Cells
In order to assess the stem cell characteristics of iHAM cells, we investigated immunofluorescence analyses, RT-PCR, and sphere formation ability of cells at passages 45–50.
The expressions of stem cell markers on the fHAM cells and iHAM cells were detected by immunofluorescence. Oct3/4 and Sox2 were strong, while Klf4 and Nanog showed little reaction. Comparing fHAM cells and iHAM cells to Oct3/4, fHAM cells were scattered positive cells stained in the cytoplasm, with few cells positive in the nucleus, whereas in the iHAM cells, all nuclei reacted strongly. The same phenomenon was seen with Sox2. The expressions of Klf4 and Lin28 were limited to the nucleus, like Oct3/4. Nanog was observed weakly in the cytoplasm, and slight reactions of SSEA-4 and nestin were seen. In contrast, α-SM actin, which is a differentiation marker for mesoderm, was expressed on several cells. No cytokeratin (CK)-positive cells were observed (Fig. 3).
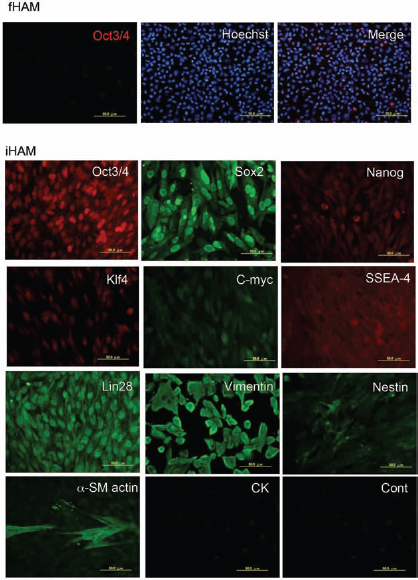
Immunohistochemical analysis of fHAM cells and iHAM cells. The expressions of stem cell markers on the fHAM cells and iHAM cells were detected by immunofluorescence. Most of the upper layer was fHAM cells, including some octamer binding transcription protein 3/4 (Oct3/4)-positive cells. From the third layer, they were iHAM cells. The nuclei of most cells were stained by Oct3/4, sex-determining region Y box 2 (Sox2), Lin28, and Krüppel-like factor 4 (Klf4). Some nuclei were stained strongly by Nanog, nestin, and stage-specific embryonic antigen-4 (SSEA-1). Vimentin and anti-α-smooth muscle (α-SM) actin were clearly expressed in the cytoplasm. No cytokeratin (CK)-positive cells were observed. Scale bar: 50.0 mm.
These results were consistent with the results of RT-PCR in the four cell lines. iHAM3 542 and iHAM4 136-344 expressed strongly for the stem cell markers; Oct-4, Sox2, Nanog, Klf4, c-myc, breast cancer resistance protein (BCRP), and nestin (neural progenitor cells marker) mRNAs were expressed strongly through over 45 passages. However, the expression of mRNA of Sox2 was low in iHAM3 542-344 (Fig. 4, d lane). Nanog showed low expression in iHAM344-96 (Fig. 4).

(A) Expression of representative stem cell marker genes by RT-PCR. Samples are as follows: lane -RT, reagent control; lane 1, freshly isolated HAM cells (fHAM cells); lane 2, HAM cells cultured in Dulbecco's modified Eagle's medium (DMEM) with 10% fetal bovine serum (FBS) at passage 2 (HAM cells P2); lanes 3–6, iHAM cells lines (iHAM3 542, iHAM3 542-344, iHAM4 136-344, and iHAM4 344-96). All iHAM cells lines showed stronger reactions than fHAM cells and HAM cells P2 for Oct-4, Klf4, c-myc, breast cancer resistance gene (BCRP), and nestin (neural progenitor cell marker). However, expression was weaker than fHAM cells and HAM cells P2 for Sox2 and Nanog. β-Actin mRNA was used as a positive control. (B) Formation of spheres. iHAM cells made spheres easily after they were cultured on the ultralow attachment dish. There were various sizes of spheres. Notice that they indicated alkaline phosphatase (ALP) activity (scale bar: 500 mm) and Oct3/4 reaction (scale bar: 200 mm).
When iHAM cells were cultured in an ultralow attachment dish, they made spheres easily even if their concentration was low. There were various sizes of spheres, showing strong alkaline phosphatase (ALP) activity and Oct3/4 reaction (Fig. 4B).
Adipogenic, Chondrogenic, and Osteogenic Differentiation
The multipotency of iHAM cells was assessed by histochemical staining and RT-PCR assays.
Adipogenic differentiation of iHAM cells was estimated by Oil Red O staining for neutral lipids (Fig. 5A). Various sizes of lipids droplets were seen in the cytoplasm of iHAM cells after 3 weeks of adipogenic induction (Fig. 5A, a-2, b–d). However, uninduced cells also showed a few small lipid droplets in cytoplasm (Fig. 5A, a-1).

Differentiation potentials of immortalized HAM cells (iHAM cells). (A) Adipogenic differentiation. (1) After culture for 3 weeks, cells were stained with Oil Red O. a-1, no induction. However, a few small particles were seen in the cytoplasm. From a-2 to d, induced iHAM cells, they were induced for 3 weeks by Lonza's induction medium. Almost all cells contained various sizes of particles in the cytoplasm. (2) Expression of peroxisome proliferating-activated receptor γ2 (PPARγ2) mRNA following the induction of adipogenic differentiation. Scale bar: 50 mm. (B) Chondrogenic differentiation. Histochemical evidence for the chondrogenic differentiation by staining with toluidine blue. a-1, no pellets; from a-2 to d, the pellets were surrounded with two or three layers of fibroblast-like cells like a perichondrium. The cellular matrix did not show clear metachromasy. Scale bar: 100 mm. Expression of collagen type II (Col2A1) and Sox9 mRNA following the induction of chondrogenic differentiation. The appearance of collagen type II mRNA varied according to the difference of the cell lines. Scale bar: 100 mm. a, iHAM3 542 (a-1 without induction, a-2 induction); b, iHAM3 542-344; c, iHAM4136-344; d, iHAM4 344-96.
RT-PCR analyses indicated very strong expression of peroxisome proliferator-activated receptor γ2 (PPARγ2) mRNA, the adipogenic marker, in the induced cells (Fig. 5A, 2).
Chondrogenic differentiation was observed by pellet culture and immunostaining. In the uninduced cells, it was not possible to make pellets. Induced cells made pellets in which two or three layers of fibroblast-like cells surrounded the pellets like a perichondrium. The cellular matrix was not metachronous, with toluidine blue staining for proteoglycan suggesting that there was insufficient proteoglycan to produce a matrix.
On RT-PCR analysis, Sox9 was strongly expressed in induced cells. Both iHAM3 cell lines without induction also expressed it. The reaction of collagen type II (Col2A1) mRNA was seen in all lanes, although the reaction of iHAM4 344-96 was strongest among them (Fig. 5B).
We observed the osteogenic differentiation of iHAMs cell lines by mineralization with alizarin red staining and ALP activity (Fig. 6) after 3 weeks of induction. All cell lines showed presence of reactive materials on the cells after induction (Fig. 6, 1-i). ALP activity was seen on some cells in all cell lines after induction (Fig. 6, 1-ii).

Osteogenic differentiation. Histochemical evidence for osteogenic differentiation by staining with Alizarin Red S and alkaline phosphatase activity. Scale bar: 50.0 mm. Expression of alkaline phosphatase (ALP) and osteopontin mRNA following the induction of osteogenic differentiation. Scale bar: 50.0 mm. a, iHAM3 542 (a-1 without induce, a-2 induce); b, iHAM3 542-344; c, iHAM4 136-344; d, iHAM4 344-96.
On RT-PCR analysis, mRNA of ALP and osteopontin (OPN) was detected in all cell lines after differentiation stimulation (Fig. 6, 2). Especially, the iHAM4 136-344 cell line demonstrated strong ALP and osteopontin mRNA (Fig. 6, 2 lane d).
Flow Cytometric Analysis of Cell Surface Antigen Expression of iHAM Cells

fHAM, freshly isolated HAM cells; P2HAM, HAM cells cultured in DMEM with 10% FBS at passage 2. iHAM cell lines: iHAM3 542, iHAM3 542-344, iHAM4 136-344 and iHAM4 344-96.
Discussion
In this study, we found that combinations of bmi-1 or E6/E7 with hTERT genes or transduction with E6/E7 alone could extend the lifespan of HAM cells near or over 200 PDs although normal somatic cell populations proliferated no more than 50 PDs (18). This result suggested that these immortalized cell lines were successfully established by down-regulation of p16 INK4a and p53 expression and activation of telomerase (13, 19). It is reported that extension of the lifespan of human umbilical cord blood-derived mesenchymal stem cells (UCBMSCs) by transfecting hTERT alone did not require inhibition of the p16INK4a/RB pathway (20). Indeed, in our study, the lifespan of HAM cells transduced with hTERT alone could not be extended over 10 passages (data not shown). This indicates that telomerase activity was not sufficient to immortalize HAM cells and agrees with inhibition of the p16INK4a/RB pathway being necessary for immortalization of human placenta-derived mesenchymal cells (hPDMCs) (29). Recently, Wolbank et al. (26) succeeded in the establishment of two cell lines of immortalized human amnion-derived mesenchymal stem cells by ectopic expression of the catalytic subunit of hTERT. They were largely unaltered in terms of surface marker profile and morphology, but one of them showed increased immunogenicity. The expressions of the cell surface markers were very different between the cell lines. It is important to investigate the mechanism of immortalization in detail. We had established cell lines by oncogenes or viral genes in addition to hTERT. Although the oncogene or viral is not suitable for regenerative medicine, it was advantageous for the study of somatic stem cells because the nature of stem cells was stable between the cell lines.
In the experiments reported here, iHAM cells did not form teratomas or tumors in nude mice. Unlike hESCs and Hela cells, iHAM cells do not express telomerase and are not tumorigenic upon transplantation.
In this study, we showed that cells after genes transfection with an extended lifespan were able to retain their differentiation potential. It has been reported that HAM cells can differentiate into osteogenic, chondrogenic, and adipogenic as mesodermal lineages and also into three germ layers—the ectoderm (neural), mesoderm (skeletal muscle, cardiomyocytic, and endothelial), and endoderm (pancreatic) (2, 8, 16, 25, 27). Our results showed that the differentiation potential of iHAM cells was maintained even after long-term culture. Freshly isolated HAM cells were rarely differentiated into chondrocytes. However, when freshly isolated HAM were differentiated, the cells were well differentiated into chondrocytes. However, iHAM cells showed slightly lower differentiation ability for chondrogenic differentiation; even if the mRNA of collagen type II was expressed, it was too weak to produce proteoglycan. This seems to be due to prolonged culture (over 45 passages) as reported previously (22, 30). Another possibility was that some iHAM cells spontaneously and randomly differentiated into a variety of cell types under attachment culture in DMEM with 10% FBS. For example, we observed a few iHAM cells spontaneously differentiate to α-smooth muscle actin-positive cells without direct induction, as well as some pluripotent markers of differentiated cells that also might have changed in this process. When we recultured iHAM cells for several passages in special ES culture media, such as mesenchymal stem cell growth medium BulletKit (MSCGM; Cambrex, Walkersville, MD) (18, 20) or CHANG MEDIUM™ C (Irvine Scientific), the cells reverted to high proportions of CD90, CD73, and CD105 positivity (>95%). This result agreed with the criteria recently proposed by Parolini et al. (15), in which the specific pattern of surface antigen expression at passages 2–4 for hHAM cells should be positive (>95%) for CD73, CD90, and CD105 and negative (<2%) for CD14, CD34, CD45, and HLA-DR (1). The surface marker expression of human chorionic mesenchymal stromal cells (hCMSCs) (15) was unaffected by the exogenously expressed bmi-1, E6, E7, and/or TERT genes. When iHAM cells were compared to freshly isolated HAM cells (fHAM cells) and cultured HAM cells at passage 2 (P2HAM cells), fHAM cells and P2HAM cells displayed lower proportions of positivity for CD73, CD90, CD105, and CD44. However, our results suggested that the proportion of mesenchymal stem cell marker increased gradually from the primary culture, as repeated cultures were performed. In addition to surface markers, iHAM cells expressed molecular markers of pluripotent stem cells including OCT-4, SOX-2, Nanog, SSEA4, KlF4, c-myc, and Lin28, indicating that iHAM cells with an extended lifespan after long-term culture showed “stemness” characteristics.
All iHAM cells showed stronger expressions of mRNA of stem cell markers, such as Oct-4, Klf4, c-myc, BCRP, and nestin (neural progenitor cells marker) compared to fHAM cells and P2 HAM cells at the RNA level. However, Nanog and Sox2 were weaker than fHAM cells in iHAM 3 542-344 and iHAM4 344-96 cells. Recently, the mRNAs of isoforms of OCT-4 were found to encode proteins that have identical POU DNA binding domains and C-terminal domains but differ in their N-terminal domains. OCT-4B is mainly localized to the cytoplasm (11). In this experiment, we did not distinguish between the isoforms OCT-4A and OCT-4B because we were focused on expression in cells.
For the ability of pluripotency, with stimulation for chondrogenesis, all cell lines expressed mRNA of collagen type II, with iHAM4 344-96 cells expressing particularly strongly. However, metachromasy staining with toluidine blue solution was not clear. It seems that these cell lines were induced to chondrogenesis, but not completely. These cell lines possessed the ability of chondrogenic differentiation because they had Sox9, which is as essential gene for making chondrocytes (1). In the process of chondrogenesis, aglycan appears following type II collagen.
Osteogenesis, too, was not complete. Some cells with ALP activity remained 3 weeks after osteogenic stimulation. Two or 3 weeks after the induction for osteogenesis, transient overexpression of ALP activity was seen and then decreased with osteopontin or another protein being produced instead.
These cells possessed the ability of differentiation to cartilage and bone. iHAM3 542-344 and iHAM4 344-96 transduced the same genes, hTERT and E6E7, but their differentiation potentials were different. This is consistent with Parolini et al. who reported that the surface marker expression of hCMSCs (15) was unaffected by the exogenously expressed bmi-1, E6, E7, and/or TERT genes. It is necessary to investigate in detail the relationship between the expressions of stem cell markers and differentiation abilities with immortalized genes.
In conclusion, combination of hTERT with human papillomavirus E6 and E7 or bmi-1 genes successfully produced iHAM cells with potential for adipogenic, osteogenic, and chondrogenic differentiation. The properties, such as (1) iHAM cells being of fetal origin, (2) less ethical controversy, (3) maintained multipotency with extended lifespan, (4) not tumorigenic, and (5) short population doubling time allowing production of large numbers, suggested that iHAM cells are a good material for investigations of somatic stem cells.
Footnotes
Acknowledgments
