Abstract
For human embryonic stem cells (hESCs) to be used clinically, it is imperative that immune responses evoked by hESCs and their derivates after transplantation should be prevented. Human leukocyte antigens (HLA) and ABO blood group antigens are important histocompatibility factors in graft rejection. HLA matching between recipient and unrelated donors, in particular, is important in improving outcomes in hematopoietic cell transplantation (HCT). We have established and successfully maintained 29 hESC lines and analyzed the HLA and ABO genotypes of these lines. HLA-A, -B, -C and -DR (DRB1) genotyping was performed by polymerase chain reaction (PCR) sequence-based typing and ABO genotyping was carried out by PCR restriction fragment length polymorphism methods. To determine what proportion of the Korean population would be covered by these cell lines in organ transplantation, 27 cell lines with HLA-A, -B, and -DR data were evaluated for HCT (cord blood) donors and 28 cell lines with HLA-DR and ABO data were evaluated for solid organ (kidney) transplantation donors, and then compared the data with those from 6,740 donated cord bloods. When 2 HLA mismatches are allowed for HCT, as currently accepted for cord blood transplantation, it was estimated that about 16% and 25% of the possible recipients can find one or more donor cell lines with ≤2 mismatches at A, B, DRB1 allele level and at A, B antigen/DRB1 allele level, respectively. When HLA-DR antigen level matching and ABO compatibility was considered for solid organ (kidney) transplantation, it was estimated that about 29% and 96% of the possible recipients can find one or more ABO-compatible donor cell lines with 0 and 1 DR mismatches, respectively. We provided the first report on the HLA and ABO genotypes of hESC lines, and estimated the degree of HLA and ABO matching in organ transplantation for the Korean population.
Introduction
The development of human embryonic stem cells (hESCs), with their unlimited proliferation and differentiation ability, raises hope in patients that scientists may one day be able to cure currently untreatable illnesses. Reaching that goal will require overcoming certain problems with hESCs. First, the derivation, maintenance, and differentiation of hESCs should be accomplished under xeno-free culture condition using good manufacturing practice (GMP) systems (32,58,66). Second, guided differentiation methods of hESCs into a given functional cell type must be established, and it must be demonstrated that the resulting cells are homogeneous and do not form teratomas or cause cancer (40). Third, the immune response/rejection caused by transplantation of hESCs or their differentiated derivatives should be prevented.
The culture conditions for the derivation and maintenance of hESCs necessary for clinical applications, such as cell therapy, have been established very rapidly, especially given the relatively recent development of the field. The first report of hESC derivation in 1998 (65) described an immunosurgery procedure for the isolation of inner cell mass (ICM) cells, the use of mouse embryonic fibroblast (MEF) cells as feeder cells, and fetal bovine serum (FBS)/leukemia inhibitory factor (LIF) as components of the culture medium. Currently, basic fibroblast growth factor (bFGF) is the most important factor for maintaining pluripotency, and Knockout serum replacement (SR) and various human feeder cells originating from fetal skin, fallopian tube, foreskin, uterine endometrium, and other tissues have replaced FBS and MEFs in hESC culture (1,2,17,27,29,43,54). To avoid contamination of animal-derived components during the derivation procedure, mechanical isolation of ICM has been introduced instead of immunosurgery (60). Researchers have attempted to develop defined culture media to allow culture of hESCs without feeder cells or SR (3,17,35,37,67,68,70—72). The generation of clinical-grade hESC lines under GMP conditions was recently reported, although these cells were prepared using US FDA-approved clinical-grade materials instead of a xeno-free method (10).
It has been expected that stem cells and their derivates could be used for cell therapy as they might replace damaged tissue or organ (44). Unlike other stem cells, hESCs have demonstrated true pluripotency, with the potential to differentiate into all three germ layer lineage cells (4,20,45,52). Thus, differentiated cells from hESCs could be used to replace virtually any damaged tissue or organ. To accomplish their therapeutic goals, transplanted cells must not be rejected by the patient's immune system (56). Previous reports have shown that undifferentiated hESCs expressed low levels of major histocompatibility complex (MHC) class I proteins but do not express MHC class II proteins (14,34). Moreover, it was reported that, during differentiation in vitro and in vivo, the expression levels of MHC class I proteins in hESCs increased but MHC class II proteins were not expressed on differentiated hESCs (14). The presence of MHC antigens on hESCs might induce an immune response; thus, hESCs injected in vivo across histocompatibility barriers would be at risk for rejection (6). Studies have reported that hESCs or their derivatives have low immunostimulatory potential and failed to stimulate proliferation of alloreactive human T cell (34), and have shown that undifferentiated hESCs formed teratomas in immunocompetent mice after humanization by injection of human peripheral blood leukocytes (13). In contrast, it was reported that hESCs transplanted into immunocompetent mice were effectively recognized and rejected by the adaptive murine immune system (62). The immunogenicity of hESCs and their differentiated derivatives, especially in vivo, is still not well understood.
A number of genes for human leukocyte antigens (HLA) are highly polymorphic and encode various MHC class I and II proteins; the three major MHC class I genes are HLA-A, -B, and -C, and the three major MHC class II genes are HLA-DP, -DQ, and -DR. The proteins (antigens) encoded by HLA are expressed uniquely on the surface of cells in the body, and are used to distinguish self from non-self; thus, differences in MHC proteins become a serious impediment for transplantation of hESCs or their differentiated derivatives. In fact, a number of reports have suggested that the risks of severe graft-versus-host disease (GvHD), graft failure, and mortality increase in proportion to HLA disparity in allogeneic hematopoietic cell transplantation (HCT) using bone marrow or umbilical cord blood (7,31,49,50,57). It is very difficult to find suitable, fully HLA-matched donors; nevertheless, it is important to minimize HLA mismatching between donor and patient to maximize graft outcome. Another factor to be considered for transplantation is ABO blood group antigens. ABO compatibility between the donor and recipient has long been considered an absolute requirement for successful solid organ transplantation, although ABO incompatibility is not an obstacle for successful HCT (9).
The ideal solution for avoiding or minimizing immune response/rejection in allogeneic transplantation is to use donor cells that are genetically identical to those of the patient. One way to accomplish this is by derivation of hESCs from oocytes generated by somatic cell nuclear transfer (SCNT); another is to reprogram a patient's somatic cells, creating induced pluripotent stem cells (iPS) (25). There are currently no reports demonstrating successful establishment of hESCs through SCNT, perhaps because of potential ethical concerns associated with this method. Since Takahashi and Yamanaka (63) first reported the induction of pluripotent stem cells from somatic cells in mice, there have been several reports describing the successful production of iPS in mice and humans using defined factors (22,38,41,46—48,69). At present, the procedure for reprogramming to generate iPS cells mostly involves genetic modification using viral transduction, making it difficult to apply the iPS solution in a clinical context, although some researcher have shown that iPS are able to be generated without virus construction (23,59,73). The next option is to establish a hESC bank that has a large number of hESC lines, which collectively match HLA and ABO types in a majority of patients. In the last 10 years, more than 400 hESC lines have been established and studied worldwide (International stem cell registry, University of Massachusetts medical school; http://www.umassmed.edu/iscr/index.aspx) (16,18); therefore, the organization of a large bank of hESC is not impossible. In 2005, using HLA data from 10,000 cardiac organ donors and 6,577 patients awaiting kidney transplants in the UK, Taylor et al. (64) estimated that approximately 150 hESC lines would be needed to establish a therapeutic hESC bank capable of providing an HLA match for most potential patients. Nakajima et al. (42) also estimated that a bank containing 170 hESC lines could offer at least one hESC line with a single mismatch at one HLA loci to 80% of patients in the Japanese population. If parthenogenetic homozygous hESC lines could be established, the number of hESC lines needed to establish such a bank would be reduced from 170 to as few as 55 under the same conditions.
To date, we have established 31 hESC lines under various conditions that comply with the derivation of clinical-grade hESCs, and have successfully maintained 29 such lines in the CHA Stem Cell Institute, Korea. We have analyzed the HLA-A, -B, -C, and -DR genotypes of these 29 hESC lines (CHA-hES 3—26, M1, and R1—R4) by polymerase chain reaction (PCR) sequence-based typing, and performed ABO genotyping using PCR restriction fragment length polymorphism method. We successfully determined the ABO genotypes of all hESC lines and obtained HLA-A, -B, -C, and -DR genotype results from 27 of the 29 lines; the exceptions were two hESC lines that showed chromosomal triploidy/tetraploidy, and HLA-A, -C, and -DR results were obtained in one of these two cell lines. In this study, we compared the HLA and ABO genotypes of the CHA-hESC lines as donors with those of 6,740 donated cord blood samples as recipients to determine what proportion of the Korean population would be covered by these cell lines for cell/tissue transplantation. For a simulation of matching these cell lines as possible organ donors, 27 cell lines with HLA-A, -B, and -DR data were evaluated for HCT (cord blood) and 28 cell lines with HLA-DR and ABO data were evaluated for solid organ (kidney) transplantation. Our results demonstrate that established hESC lines have a potential as HCT donors with ≤2 ABDR (A, B antigen/DRB1 allele level) mismatches and as ABO-compatible solid organ (kidney) donors with 0 DR (antigen level) mismatch for 25% and 29% of the Korean population, respectively.
Materials and Methods
hESC Derivation, Maintenance, and Characterization
Frozen human embryos from pronucleus to blastocyst stage were donated by couples undergoing infertility treatment after obtaining informed consent and approval by the Institutional Review Board (IRB) of CHA Gangnam Medical Center. All processes followed IRB guidelines prior to 2005 and then relevant laws enacted since 2005. After thawing, donated embryos were cultured in sequential media to the blastocyst stage for 1—6 days: P1/Blastocyst medium (Irvine Scientific Inc. Santa Ana, CA) or G1.2/G2.2 medium (Vitrolife, Englewood, CO). hESC lines were established by removing the zona pellucida of blastocysts (except hatched blastocysts) by treatment with 0.3% pronase (Sigma P-5147, St. Louis, MO) in DMEM/F12 or acid Tyrode's solution (Sigma) in DMEM/F12. The trophoectoderm (TE) was then removed by immunosurgery using an anti-human whole serum antibody (Sigma H-8765) and guinea pig complement (Sigma S-1639 or Calbiochem 234395, La Jolla, CA), or by mechanical isolation through manual transfer of ICM outgrowths after seeding intact blastocysts on feeder cells. The STO cell line (ATCC CRL-1503, Manassas, VA) or mouse embryonic fibroblasts (mEFs) were used as mouse feeder cells; human embryonic fibroblasts (hEFs) or human foreskin fibroblasts (hFSs) were used as human feeder cells after mitotic inactivation by mitomycin C (Sigma) treatment. hEFs and hFSs were established from aborted human fetuses and human foreskin tissue after obtaining informed consent. After spreading the ICM clump on feeder cells, ES cell-like colony was cultured and mechanically dissected into small clumps every 5—6 days. ES cell colonies were cultured in DMEM/F12 medium supplemented with 20% Knockout SR, 0.1 mM β-mercaptoethanol, 1% nonessential amino acids, 100 U/ml penicillin, 100 μg/ml streptomycin, and 4 ng/ml basic fibroblast growth factor (bFGF; all products from Gibco/Invitrogen, Grand Island, NY) at 37°C in a humidified 5% CO2 incubator.
hESCs were cyropreserved by transferring hESC clumps stepwise into media containing incremental increases in SR and DMSO (final composition: 90% SR + 10% DMSO), loaded into 0.25 ml straws (IMV, L'Aigle cedex, France; 30—70 clumps/straw), and frozen using a programmed cell freezer (CryoMagic, Mirae Biotech, Korea) (30).
To confirm endogenous alkaline phosphatase activity and hESC-specific surface marker expression in established hESCs, we stained hESC colonies with an FRV-alkaline, sodium nitrite, and naphthol AS-BI alkaline solution mixture (Sigma) for 20 min at room temperature (RT), and performed immunocytochemistry using antibodies against SSEA-4 and TRA 1-60 (Chemicon, Temecula, CA) after fixing with 4% paraformaldehyde (PFA). We also ascertained the Oct-4 gene expression status by reverse transcription-polymerase chain reaction (RT-PCR) or immunocytochemistry using an anti-OCT-4 antibody (Chemicon). The following primer sequences were used for Oct-4: (forward) 5′-gga aag gct tcc ccc tca ggg aaa gg-3′, (reverse) 5′-aag aac atg tgt aag ctg cgg ccc-3′ (460 bp). A fluorescein-labeled goat anti-mouse IgG antibody (Molecular Probes, Eugene, OR) was used as a secondary antibody and nuclei were stained with DAPI (Vector Laboratories, Burlingame, CA).
For the cytogenetic analysis of hESCs, they were incubated for 3 h in hESC culture medium containing 0.1 μg/ml colcemid (KaryoMax colcemid solution; Gibco-BRL). Thereafter, they were treated with a hypotonic solution (1% sodium citrate buffer) for 30 min and then fixed with methanol and acetic acid (3:1, v/v). Cells were spread onto a glass slide and dried, and chromosomes were identified by G banding. For karyotyping of each cell lines, more than 20 metaphase chromosomes were counted by an expert.
Each hESC line was characterized with respect to short tandem repeat (STR) loci by DNA fingerprinting. Genomic DNA was extracted and 15 STR loci (D3S1358, TH01, D21S11, D18S51, Penta E, D5S818, D13S317, D7S820, D16S539, CSF1PO, Penta D, vWA, D8S1179, TPOX, FGA) and the gender marker Amelogenin were co-amplified using a PowerPlex® 16 System (Promega, San Luis Obispo, CA) and an ABI 3100 Genetic Analyzer (Applied Biosystems, Foster City, CA).
The differentiation capacity of each hESC line was confirmed by injecting undifferentiated hESCs into a testicle of an NOD/SCID male mouse as clumps (about 1 × 106 cells/150 μl PBS). After 14—20 weeks, teratomas were excised, fixed in PFA, embedded in paraffin, sectioned, and then analyzed histologically after staining.
HLA and ABO Genotyping
Donated cord blood data, which included HLA genotypes and ABO blood group antigens, was provided by iCORD (Cord Blood Bank of CHA Bio & Diostech Co., Seoul, Korea) and compared with HLA and ABO genotypes determined for CHA-hESC lines.
High resolution (allele level) HLA typing of hESC and cord blood was performed by PCR sequence-based typing using a Secore® Sequencing Kit (Invitrogen, WI), using a DNA Engine Tetrad Thermal Cycler (Bio-Rad, Hercules, CA) and an Applied Biosystems 3130xl Genetic Analyzer (Applied Biosystems, Foster City, CA). HLA-A, -B, -C, and -DR genotypes were determined for hESC, and HLA-A, -B, and -DR genotypes were determined for donated cord blood.
ABO genotyping was performed by PCR restriction fragment length polymorphism using the restriction enzymes, Kpnl and Alul, and a PTC-300 Thermal Cycler (MJ Research, Waltham, MA).
Results
As part of a research effort to develop hESCs for clinical applications, we have established 31 hESC lines under various conditions, 29 of which have been maintained and characterized (Table 1 and refer to http://csci.cha.ac.kr/CHA-hESClines). CHA-hES 3—12 were established after isolation of ICM by immunosurgery and CHA-hES 13—26, M1, and R1—R4 were established by mechanical isolation of ICM with no use of anti-human serum and guinea pig complement, conditions that approach xeno-free (see Discussion). CHA-hES 9, 18, and 20—26 were established on human feeder cells and the remaining CHA-hESC lines were established on mouse feeder cells. Twenty-nine CHA-hESC lines have been cryopreserved at various passages; all 24 hESC lines that have been thawed were found to be viable, and an additional 5 hESC lines that have not yet been thawed are available for the future use (Table. 1).
Derivation Conditions and Characteristics of hESCs Established and Maintained at the CHA Stem Cell Institute, Korea
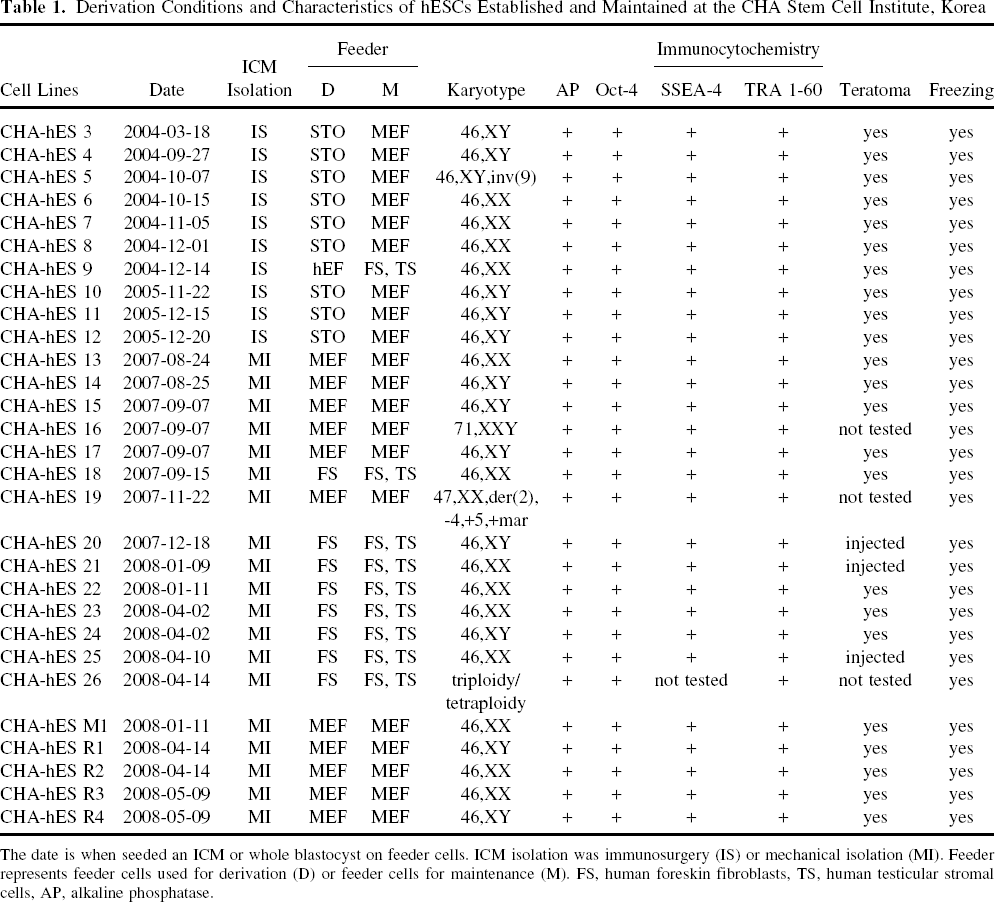
The date is when seeded an ICM or whole blastocyst on feeder cells. ICM isolation was immunosurgery (IS) or mechanical isolation (MI). Feeder represents feeder cells used for derivation (D) or feeder cells for maintenance (M). FS, human foreskin fibroblasts, TS, human testicular stromal cells, AP, alkaline phosphatase.
All hESC lines showed endogenous alkaline phosphatase activity and were positive for SSEA-4, TRA 1-60, and Oct-4. A karyotypic analysis showed that 26 hESC lines were normal [CHA-hES 3, 4, 10, 11, 12, 14, 15, 17, 20, 24, R1, R4 for 46,XY; CHA-hES 6, 7, 8, 9, 13, 18, 21, 22, 23, 25, M1,R2,R3 for 46,XX; CHA-hES 5, 46,XY, inv(9)]; three hESC lines—-CHA-hES 16 (71,XXY), 19 [47,XX, der(2),-4,+5,+mar], and 26 (triploidy/tetraploidy)—-had abnormal karyotypes (Table. 1). DNA fingerprinting results (Table. 2) were in agreement with the gender and triploidy results of karyotypic analyses, and demonstrated that each of the 29 hESC lines were distinct from one other. The differentiation potential of the 26 karyotypically normal hESC lines was determined by injecting hESCs into the testicle of an NOD/SCID male mouse. Twenty-three hESC lines, such as CHA-hES 3—15, 17, 18, 22, 23, 24, M1, and R1—R4, formed teratomas, and their teratomas showed that these hESCs differentiated into various cell types composed of three germ layers (http://csci.cha.ac.kr/CHA-hESClines). Teratoma formation by injection of CHA-hES 20, 21, and 25 is currently under examination to confirm their differentiation ability using various stains. The karyotypically abnormal hESC lines such as CHA-hES 16, 19, and 26 have not been tested for teratoma formation.
DNA Fingerprinting Results From 29 CHA-hESC Lines

Raw data were not shown.
FS, human foreskin fibroblasts were derived from foreskin tissues of two different donors.
Previous reports showing that classical HLA typing by serological methods (serotyping) is inaccurate have led to the development of the more precise gene sequencing method (genotyping). In this study, we performed high-resolution HLA-A, -B, -C, and -DR typing of our 29 hESC lines and obtained complete HLA typing results from all but two (Table. 3). We were unable to obtain data from CHA-hES 16 (all HLA loci) and CHA-hES 26 (HLA-B locus), which failed PCR sequence-based typing (twice in each case). In case of CHA-hES 16, we got the results from PCR reaction and sequencing for HLA-A, -B, -C, and -DR, but failed to get the matching results from DATA base. It was inferred that the computer program recognized input data as an error because CHA-hES 16 had three chromosome 6 and the sequencing results would be three for each HLA gene. For CHA-hES 26, we failed PCR reaction for only HLA-B. Of the 27 CHA-hESC lines, none were homozygous at all of the HLA-A, -B, and -DR loci. Only a few hESC lines were homozygous at a single HLA locus: one (high-resolution, 4 digits) or five (low-resolution, 2 digits) were homozygous for HLA-A; two (low-resolution) were homozygous for HLA-C; and one (high-resolution) was homozygous for HLA-DRB1.
HLA and ABO Genotypes of 29 CHA-hESC Lines
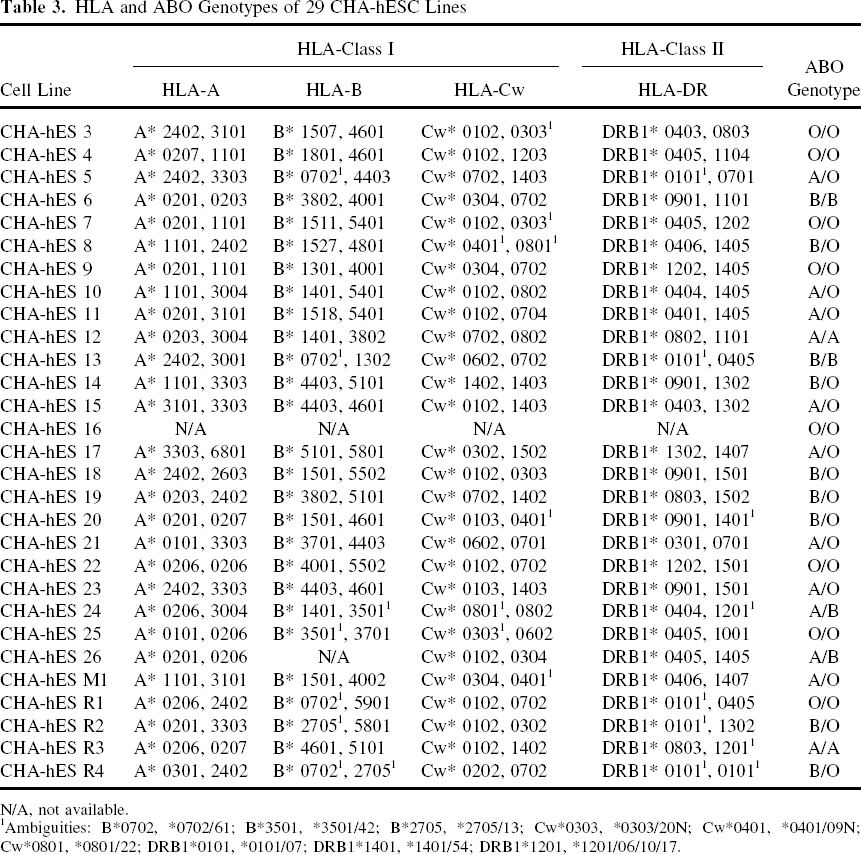
N/A, not available.
Ambiguities: B*0702, *0702/61; B*3501, *3501/42; B*2705, *2705/13; Cw*0303, *0303/20N; Cw*0401, *0401/09N; Cw*0801, *0801/22; DRB1*0101, *0101/07; DRB1*1401, *1401/54; DRB1*1201, *1201/06/10/17.
To determine what proportion of the Korean population would be matched for these hESC lines in terms of organ transplantation, we compared HLA and ABO types of the hESC lines as donors with those of 6,740 donated cord blood samples as recipients. The donated cord blood data were provided by iCORD, one of the Korean private cord blood banks. For a simulation of HCT (cord blood), 27 cell lines with HLA-A, -B, and -DR data were evaluated and for a simulation of solid organ (kidney) transplantation, 28 cell lines with HLA-DR and ABO data were evaluated as possible donors. For HCT, we compared HLA data at A, B, DRB1 allele level (high-resolution, 4 digits) as well as at A, B antigen/DRB1 allele level. The assignment of serological antigens for particular HLA alleles were carried out on the basis of The HLA Dictionary 2008 (21). Table 4 shows that among 6,740 cord blood samples as possible recipients of HCT, 14 (0.21%), 117 (1.74%), and 958 (14.21%) could find 0, 1, and 2 mismatches, respectively, at A, B, and DRB1 allele level. When we considered A, B antigen/DRB1 allele level of matching, these figures were increased to 20 (0.30%), 164 (2.43%), and 1,510 (22.4%), respectively. Thus, a total of 1,089 (16.16%) and 1,694 (25.13%) possible recipients could find one or more donor cell lines with ≤2 mismatches at A, B, DRB1 allele level and at A, B antigen/DRB1 allele level, respectively.
Results of HLA-A, -B, -DRB1 Matching Between 27 CHA-hESC Lines and 6,740 Donated Cord Blood Samples for Simulation of Stem Cell Transplantation
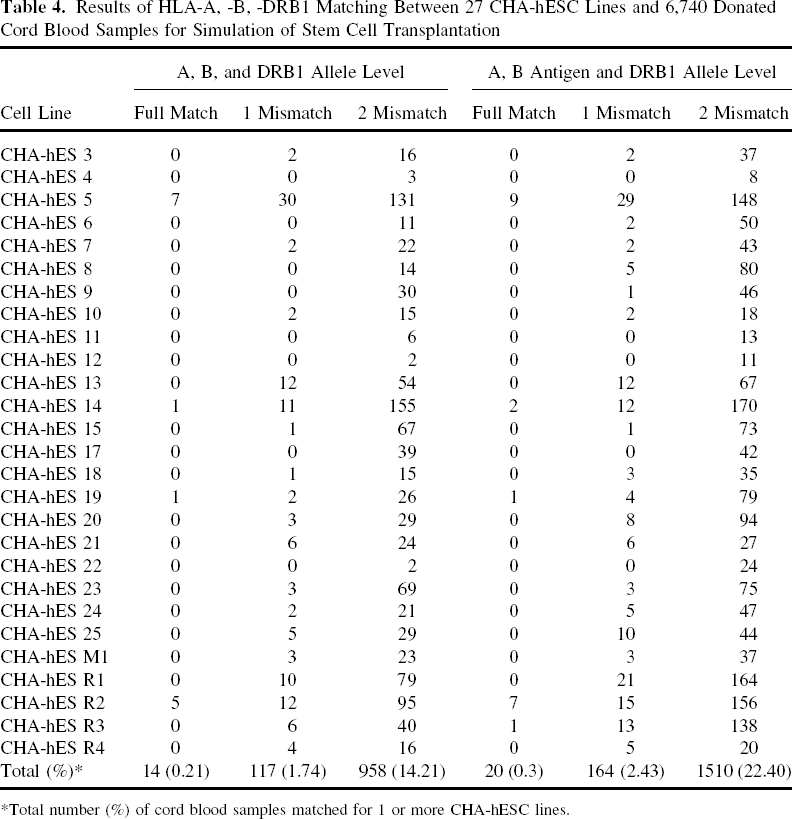
Total number (%) of cord blood samples matched for 1 or more CHA-hESC lines.
A second type of histocompatibility antigens, the ABO blood group antigens, is also important to be considered in matching donor and recipient, especially in solid organ transplantation. Matching of ABO blood type is not essential in HCT, although ABO incompatibility may cause hemolysis after transplantation (61). Table 3 shows the results of our examination of the ABO blood types of 29 CHA-hESC lines. Ten of these CHA-hESC lines were type A (A/A or A/O), nine were type B (B/B or B/O), eight were type O, and two were type AB. In consideration of solid organ (kidney) transplantation (http://www.unos.org), we analyzed HLA-DR (DRB1) antigen level matching and ABO compatibility (O to A, B, O, and AB; A to A and AB; B to B and AB; AB to AB) between 28 CHA-hESC lines as possible donors and 6740 donated cord blood samples as possible recipients (Table. 5). When only HLA match was considered, a total of 3,013 (44.7%) and 6,668 (98.93%) possible recipients could find 0 and 1 DR antigen mismatched donor cell lines, respectively. When ABO compatibility was also considered, these figures were decreased to 1,957 (29.04%) and 6,456 (95.79%), respectively.
Results of HLA-DRB1 Antigen Level and ABO Blood Group Matching Between 28 CHA-hESC Lines and 6,740 Donated Cord Blood Samples for Simulation of Solid Organ Transplantation

Total number (%) of cord blood samples matched for 1 or more CHA-hESC lines.
Discussion
Since the first report of hESC establishment, more than 400 hESC lines have been established worldwide (International stem cell registry, University of Massachusetts medical school; http://www.umassmed.edu/iscr/index.aspx) (16,18). There is also an effort on the part of several stem cell registries to disseminate information regarding various hESC lines; the emergence of large banks of hESC lines would be a logical next step that may not be long in coming. Based on HLA matching using data from donors and patients for organ/cell transplantation, the number of hESC lines needed for a therapeutic hESC bank has been estimated at approximately 150 in the UK and 200 in Japan (42,64). However, no reports have calculated the number of hESC lines needed for a bank using the HLA type of hESC lines. In this study, we performed high-resolution HLA genotyping of HLA-A, -B, -C, and -DR, obtaining complete results from 27 of the 29 CHA-hESC lines typed. An evaluation of the 27 cell lines with HLA-A, -B, and -DR data as possible donors and 6,740 donated cord bloods as possible recipients for HCT (cord blood) showed that about 16% and 25% of the possible recipients could find one or more donor cell lines with ≤2 mismatches at A, B, DRB1 allele level and at A, B antigen/DRB1 allele level, respectively. We also analyzed the ABO genotypes of all 29 CHA-hESC lines and found that 34.5% (10/29) were type A, 31.0% (9/29) were type B, 21.6% (8/29) were type O, and 6.9% (2/29) were type AB. An evaluation of the 28 cell lines with HLA-DR and ABO data as possible donors and 6,740 donated cord bloods as possible recipients for solid organ (kidney) transplantation revealed that about 29% and 96% of the possible recipients could find ABO compatible donor cell lines with 0 and 1 HLA-DR antigen mismatches, respectively.
Realizing this potential and applying hESC-based cell therapy clinically requires that a number of technical issues be surmounted. First, the derivation of clinical-grade hESC lines must not involve exposure to animal materials during the derivation/propagation process so as to avoid the risk of infection or an immune reaction to animal substances. Also, all processes from embryo production to hESC establishment and differentiation should be performed according to a GMP system in consideration of clinical use (33,58,66). As the hESC derivation method and culture condition have been developed into xeno-free system, we have successfully established and maintained 29 hESC lines in GMP-grade culture room under various conditions (Tables 1 and 2). The modifications made to achieve these near-clinical-grade hESCs include changing the hESC derivation/propagation method from immunosurgery to mechanical isolation of ICMs, and switching from mouse feeder cells to human feeder cells. We have also recently begun using human serum instead of FBS for feeder cell culture, but have not yet had an opportunity to apply this modification to the establishment of new hESC lines. Xeno-free SR from Invitrogen has recently come into the market as a next-generation substitute for SR, but its ability to support growth and maintain the characteristics of hESCs has not yet been verified. Crook et al. (10) reported the generation of six clinical-grade hESC lines using current GMP-compliant materials and procedures. Although the authors of this study did not use material that originated from humans, they developed a model for applying GMP to the production of clinical-grade hESCs that complies with regulations governing the use of human cells and cellular-based products intended for tissue repair and replacement in the US. If these hESC lines prove suitable for therapeutic use, the first step toward clinical application of hESCs would be largely solved.
A second important aspect of hESC technology that must be resolved is the immunogenicity of hESCs, an issue that has not been well studied. At the core of the immune system are the MHC genes, which encode proteins expressed on the surface of cells; in humans, the genetic subset that encodes cell surface antigens are the HLA genes. MHC class I molecules are found on most cells of the body, where they present peptides from cytosolic proteins that are recognized by cytotoxic T cells. MHC class II molecules are found only on a few cell types, including macrophages, dendritic cells, and B cells, and present peptides from exogenous proteins that are recognized by helper T cells. Previous reports have shown that undifferentiated hESCs weakly express MHC class I antigens on the cell surface, but do not express MHC class II (14,34). When hESCs differentiate into embryoid bodies (EBs) in vitro, their expression of MHC class I is increased, but MHC class II proteins remain undetected (14). Treatment of undifferentiated hESCs with interferon-γ has also been shown to induce the expression of MHC class I proteins, which are also increased in EBs by treatment with interferon-α, -β, or -gamma (14). Drukker et al. (13) reported that hESCs or their differentiated derivatives transplanted into human peripheral blood mononuclear cell-reconstituted mice were not rejected and developed normally into teratomas. Cabrera et al. (8) suggested that the low expression of MHC class I on hESCs was caused by downregulation of antigen-processing machinery (APM) components, similar to mechanisms in tumor cells, as shown by the low or absent expression of APM components in the HS293 hESC line. Based on these data, it has been suggested that hESCs and their derivatives have an immune-privileged status. A recent report has reached an opposite conclusion, however, reporting that hESCs transplanted into immunocompetent mice were effectively recognized and rejected by the adaptive murine immune system (62). Using molecular imaging techniques, this article showed that hESC survival after transplantation was significantly limited in immunocompetent mice compared to immunodeficient mice, and repeated transplantation of hESCs into immunocompetent mice resulted in accelerated hESC death.
If the immune character of hESCs is capable of eliciting an immune response upon transplantation, the immune reaction caused by transplanted hESCs must be solved before clinical application of hESCs. The strategies for preventing hESC immune recognition are (i) establishment of large banks of immunophenotyped hESC lines to match MHC alleles between hESC lines and patients; (ii) generation of isogenic hES cell lines by somatic cell nuclear transfer using the patient's own somatic cells or parthenogenesis of oocytes; (iii) induction of tolerance by hematopoietic chimerism using hematopoietic cells differentiated from hESCs; (iv) creation of universal cells by genetic modification to reduce the expression of MHC molecules; (v) transplantation of hESCs into immune-privileged sites, including the eye, brain, or testis; and (vi) use of immunosuppressive medications (5,11,12,51).
Matching ABO blood group antigens is another factor that contributes to the success of transplantation (61). Recently, Mölne et al. (39) reported that hESCs and differentiated hepatocyte- and cardiomyocyte-like derivatives expressed ABO blood group antigens. Although ABO matching may seem less important than HLA type matching in hematopoietic stem cell transplantation, in contrast to its prominent significance in solid organ transplantation (19,26), it is still worth considering. Our results indicate that the possibility of finding 0 HLA-DR mismatches between our 28 CHA-hESC lines as possible donors and 6.740 donated cord bloods as possible recipients for solid organ transplantation decreased from 45% to 29% when ABO compatibility was taken into account.
Studies of HLA matching in unrelated donor HCT using high-resolution matching have shown that matching all five HLA loci [HLA-A, -B, -C (class I) and -DRB1, -DQB1 (Class II)] lowers the risks of clinically severe GvHD, graft failure, and mortality, and further showed that these risks are amplified with increasing degree of locus mismatch (49). Although a perfect match is clearly ideal, in reality it is very difficult to find HLA full-matching unrelated donors. The current standard for the selection of cord blood units includes HLA-A, -B, and -DR typing, and the vast majority of cord blood transplantations have been performed using units with match grades of 6/6, 5/6, or 4/6 at HLA-A, B antigen level, and HLA-DR allele level (15,24,28,55). A similar situation exists with respect to using hESCs and their derivates for donor cell source for HCT: while it would be desirable to find HLA full-matching hESCs for each patient, in practice it would require quite a large number of hESC lines. If two HLA-type mismatches are assumed to be acceptable for transplantation of hESCs, as is the case for cord blood transplantation programs, it was estimated that our 27 CHA-hESC lines can cover about 16% and 25% of the Korean population with A, B, DR allele level and A, B antigen/DR allele level matches, respectively. None of our 27 CHA-hESC lines was homozygous at all of the HLA-A, -B, and -DR loci, and very few were homozygous at an individual locus. Ultimately, establishing hESC lines with homozygous HLA genotypes would be very helpful as this would reduce the number of hESC lines needed for a therapeutic hESC bank. In this context, hESC lines established from human parthenogenetic blastocysts have been shown to possess homozygous HLA genotypes (36,53). Although this latter strategy raises practical problems that would need to be resolved, this approach might prove to be a practical way to create a full complement of HLA genotypes using a minimal number of hESC lines.
When we analyzed HLA-DR antigen level matching and ABO compatibility between 28 CHA-hESC lines as possible donors and 6,740 donated cord bloods as possible recipients for solid organ (kidney) transplantation (Table. 5), it was estimated that 28 CHA-hEC lines can cover almost one third (29%) of the Korean population with 0 DR mismatch and vast majority (96%) of the population with 1 DR mismatch.
In summary, we provide the first report on the HLA and ABO blood group genotypes of 28 CHA-hESC lines, and estimated the degree of HLA and ABO matching among these hESC lines as possible donors and 6,740 donated cord blood samples as possible recipients for organ transplantation. As expected, only a small percentage (<0.5%) of possible recipients was fully matched with donor cell lines at the HLA-A, -B, and -DR loci. However, if two HLA-type mismatches were considered the minimum requirement for transplantation as currently accepted for umbilical cord blood transplantation, it was estimated that our 27 CHA-hESC lines can provide a coverage for 16% and 25% of the Korean population with A, B, DR allele level and A, B antigen/DR allele level matches, respectively. On the basis of these results, we estimate that approximately 190 (for A, B, DR allele level match) or 120 (for A, B antigen/DR allele level match) hESC lines would be needed to establish a therapeutic hESC bank for HCT to provide broad coverage for the Korean population. In addition, a stem cell bank derived from therapeutic cloning or reprogramming technologies would be required to provide patient-specific stem cells and accommodate minor populations.
Footnotes
Acknowledgments
This research was partly supported by a grant (SC-1140) from Stem Cell Research Center of the 21st Century Frontier Research Program funded by the Ministry of Education, Science and Technology, Republic of Korea and by a grant for Stem Cell Research funded by CHA University.
