Abstract
BACKGROUND AND OBJECTIVE:
We aimed to further study the role of Myelin Transcription Factor 1(MyT1) in tumor and other diseases and epigenetic regulation, and better understand the regulatory mechanism of MyT1.
METHODS:
Using bioinformatics analysis, the structure and function of MyT1sequence were predicted and analyzed using bioinformatics analysis, and providing a theoretical basis for further experimental verification and understanding the regulatory mechanism of MyT1. The first, second and third-level structures of MyT1 were predicted and analyzed by bioinformatics analysis tools.
RESULTS:
MyT1 is found to be an unstable hydrophilic protein, rather than a secretory protein, with no signal peptide or trans-membrane domain; total amino acids located on the surface of the cell membrane. It contains seven zinc finger domains structurally. At sub-cellular level, MyT1 is localized in the nucleus. The phosphorylation site mainly exists in serine, and its secondary structure is mainly composed of random coils and alpha helices; the three-dimensional structure is analyzed by modeling.
CONCLUSIONS:
In this study, the structure and function of MyT1 protein were predicted, thereby providing a basis for subsequent expression analysis and functional research; it laid the foundation for further investigation of the molecular mechanism involved in the development of diseases.
Introduction
A large number of transcription factors exist in an organism, acting as trans-acting factors. They occur in different signal transduction pathways, and can specifically regulate gene expression with cis-acting factors; thereby influence various biochemical and physiological activities, thereby regulating various biochemical and physiological activities. Myelin transcription factor (Myt)/Neural zinc finger (NZF) proteins constitute a unique family of transcription factors that play an important role in differentiation of a mass of cell types. Myt/NZF family proteins are mainly expressed in the developing nervous system and have characteristic CCHHC-type zinc finger motifs as DNA-binding domains [1].
Myelin proteins account for 30% of the total myelin sheath mass, and the protein (PLP) and myelin basic protein (MBP) are the main myelin proteins in central nervous system, constituting 80% of the total myelin content in the central nervous system [2]. MyT1 is a DNA binding protein that binds to the promoter of PLP
In the normal physiological state, MyT1 acts as a transcription factor mainly expressing in the development of the central nervous system cells [4, 5], and mediates the formation of myelin and the proliferation and differentiation of oligodendrocyte [6, 7, 8]. MyT1 also exists in some subgroups of neural precursor and mature neurons [9]. MyT1’s DNA binding domain has a special CCHHC (Cys-X4-Cys-His-X7-His-X5-Cys) structure [10], which can identify a target sequence with “AAGTT” as the core, specifically bind to various neural development-related
With the rapid development of bioinformatics, the use of bioinformatics-related software for rapid and effective analysis has become the mainstream research methodology in the field of
Materials and methods
The first, second and third-level structures of MyT1 were predicted and analyzed by bioinformatics analysis tools. The flowchart of the methods is shown in Fig. 1, and the software name and address are shown in Table 1.
Software name and address
Software name and address
Flowchart of the methods.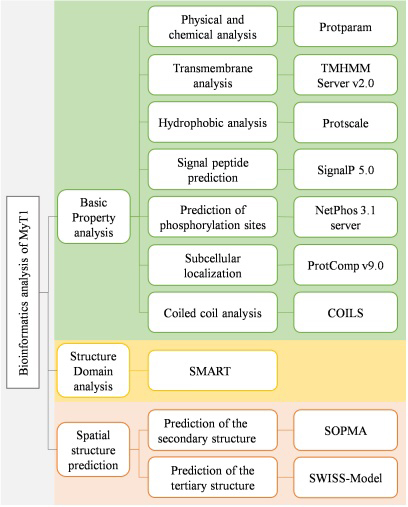
Physical and chemical analysis
According to the physical and chemical properties of amino acids, we can analyze the unknown proteins in electrophoresis experiments, as well as the physical and chemical properties of known proteins. We used ExPASy ProtParam to analyze the physicochemical properties of MyT1 [20]. ExPASY can analyze the basic properties of proteins.
Transmembrane analysis
The protein contains transmembrane region, which suggests that it may act as a membrane receptor or a membrane protein localized on the membrane. TMHMM is a program based on Markov model to predict the transmembrane helix, which can be used to predict the transmembrane region and the inner and outer membrane region. We used TMHMM Server v.2.0 to predict transmembrane domain [21]. The parameter settings: Output format choose Extensive, with graphics.
Hydrophobic/hydrophilic analysis
Hydrophobicity of amino acids reflects protein folding. Hydrophobic regions appear in potential transmembrane regions and play an important role in maintaining the tertiary structure of proteins (such as maintaining the structure of biofilm). The hydrophilicity and hydrophobicity of proteins can provide reference for the identification of protein transmembrane regions. ProtScale was used for hydrophobic/hydrophilic analysis [20].
The parameter settings: Amino acid scale choose HPhotb. /Kyte and Doolittle; Window size choose 9; Relative weight of the window edges compared to the window center in 100%; There is no need to normalize the scale from 0 to 1.
Signal peptide prediction
Signal peptide is the N-terminal amino acid sequence of secretory protein polypeptide chain, which is used to guide protein transmembrane translocation. It is generally composed of 20
The parameter settings: Organism group choose Eukarya; Output format choose Long output.
Phosphorylation sites prediction
Protein phosphorylation refers to the process of transferring phosphate groups from one compound to another by enzymatic reaction. It is a universal regulation mode existing in organisms and plays an extremely important role in the process of cell signal transmission. We used NetPhos 3.1 Server [23] to predict protein phosphorylation sites. Netphos used neural network to predict phosphorylation sites.
The parameter settings: Residues to predict choose all three; Output format choose classical; Choose Generate graphics.
Subcellular localization
Protein surface is directly exposed to organelle environment, which is determined by sequence folding process, which depends on amino acid composition. Therefore, it is possible to predict subcellular localization by amino acid composition. ProtComp v9.0 was used to check sub-cellular localization.
Coiled coil analysis
Coiled coil is a structural model existing in many natural proteins. Many proteins with coiled coil structure have important biological functions, such as transcription factors in the regulation of gene expression. We used COILS [24] for curl helix analysis. COILS compared the sequence with the known parallel double stranded coil database to obtain the similarity score, and then calculated the probability of the sequence forming the coiled coil.
The parameter settings: Window width choose all; matrix chose MTIDK; 2.5 fold weighting of positions a, d chose no.
Structural domain analysis
Domain is a kind of structural level between secondary and tertiary structure. It is the basic structural unit of protein tertiary structure, which cannot be changed and is the core of gene. We used SMART [25] for structure domain analysis. SMART is a tool for protein family comparison based on Hidden Markov chain algorithm. It provides protein sequences and queries their domains and transmembrane domains in domain database.
The parameter settings: we choose PFAM domains.
Spatial structure prediction
Predict protein secondary structure
Secondary structure refers to the regular local structural elements such as alpha-helical and beta-turn. Different amino acid residues have different tendency to form different secondary structural elements. Most of the algorithms for predicting protein secondary structure are based on the known three-dimensional structure and secondary structure of protein, and the prediction methods are constructed by artificial neural network and genetic algorithm. Sopma is a self optimizing prediction method with band comparison, which integrates several independent secondary structure prediction methods into “consistent prediction results”. We used SOPMA [26] to predict protein secondary structure.
The parameter settings: Number of conformational states choose 4 (Helix, Sheet, Turn, Coil); Similarity threshold choose 8; Window width choose 17.
Homology modeling of protein tertiary structure
According to the data in the database established by the analysis of the structure and function of natural protein, the spatial structure and biological function of a certain amino acid sequence can be predicted. Protein structure prediction can be used to determine the three-dimensional shape of protein according to its amino acid sequence. SWISS-MODEL is an automated protein comparison modeling server, which is used to predict protein structure models We used SWISS-MODEL [27] for homology modeling of protein tertiary structure. SWISS-MODEL is an automatic online software for predicting protein tertiary structure by homologous modeling method. The credibility range of GMQE (global model quality estimation) is 0–1, and the higher the value, the better the quality.
Results
The basic property analysis of MyT1
Physical and chemical analysis
ExPASy ProtParam was used, the result shows that MyT1 encodes 1157 amino acids, the most abundant amino acid is glutamic acid, the relative molecular weight is 126447.96, the isoelectric point (pI) is 4.91, negative, the molecular formula C
Prediction of the amino acid composition of MyT1
Prediction of the amino acid composition of MyT1
Amino acid species: Amino acid can be divided into four groups according to the properties of R-group. Number: The number of the specific amino acids in MyT1 protein. Ratio: The percentage of occurrences of the specific amino acids in MyT1 protein.
TMHMM Server v.2.0 was used, as shown in Fig. 2, the MyT1 has no transmembrane domain and the total amino acids are located on the surface of cell membrane.
Prediction of transmembrane of MyT1.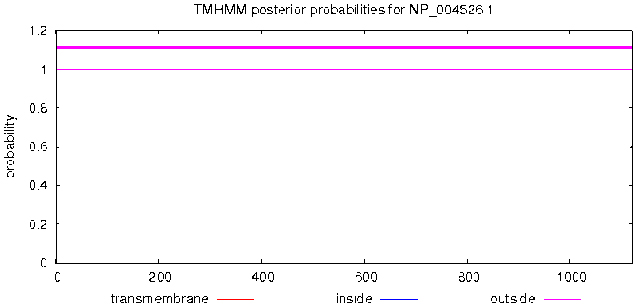
ProtScale was used, as shown in Fig. 3 and Table 3, MyT1 was found to be hydrophilic, the sum number of hydrophilic amino acid residues is
The hydrophobic and hydrophilic amino acid residue positions situations of MyT1
The hydrophobic and hydrophilic amino acid residue positions situations of MyT1
Prediction of hydrophobic/hydrophilic of MyT1.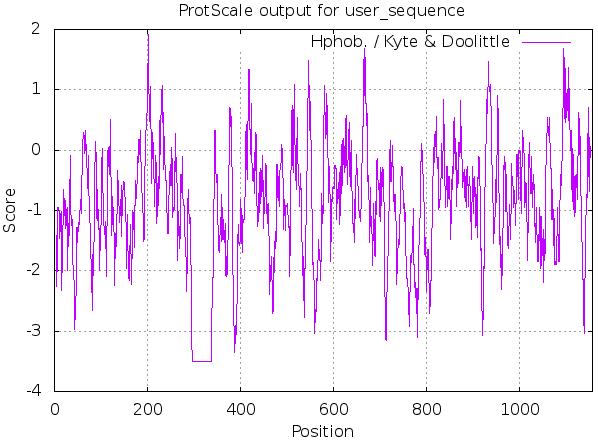
SignalP-5.0 server was used, it is shown in Fig. 4, there is no signal peptide in MyT1.
Signal peptide prediction of MyT1.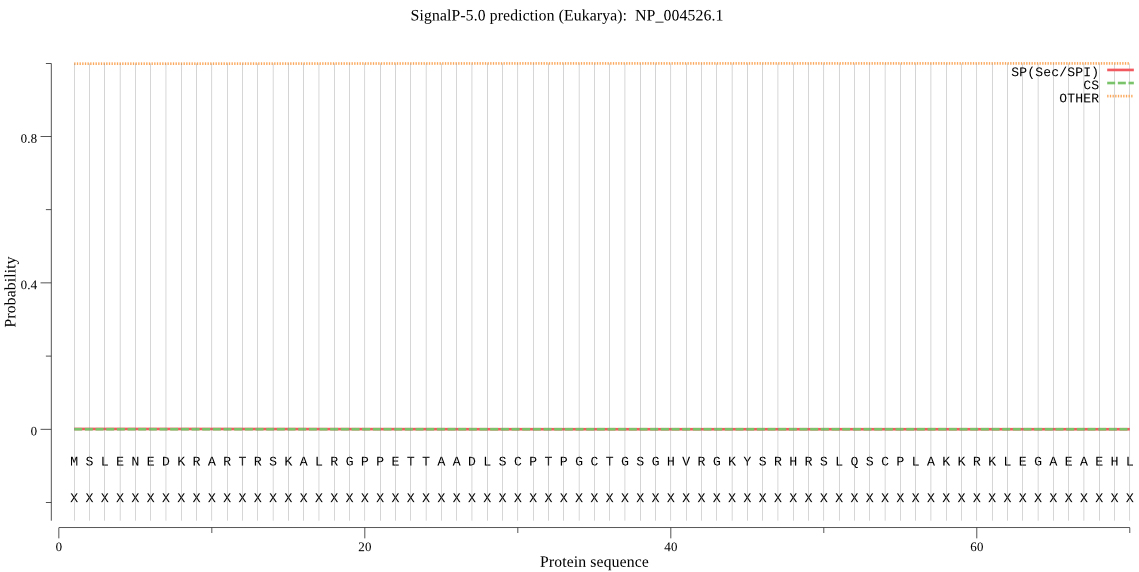
Prediction of phosphorylation sites of MyT1.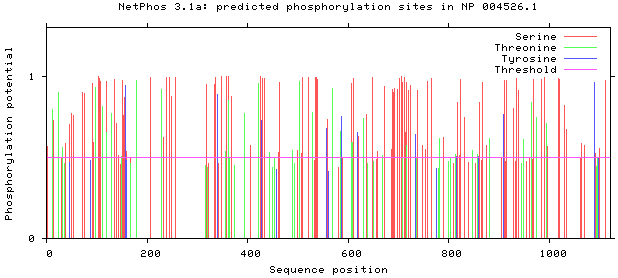
NetPhos 3.1 server was used, as shown in Figs 5–8. There are 34 threonine, 108 serine and 11 tyrosine sites in the potential phosphorylation sites of MyT1.
Prediction of serine phosphorylation sites.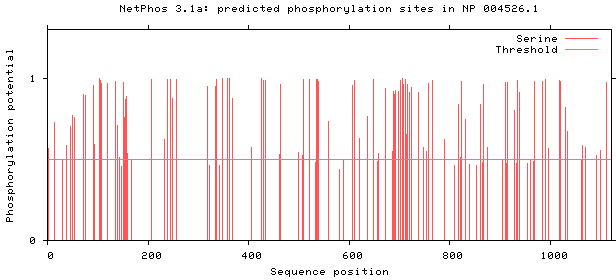
The subcellular localization of MyT1
Prediction of tyrosine phosphorylation sites.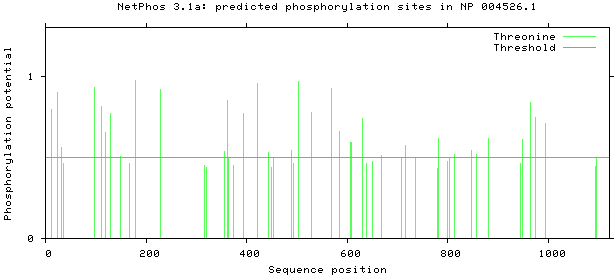
Prediction of threonine phosphorylation sites.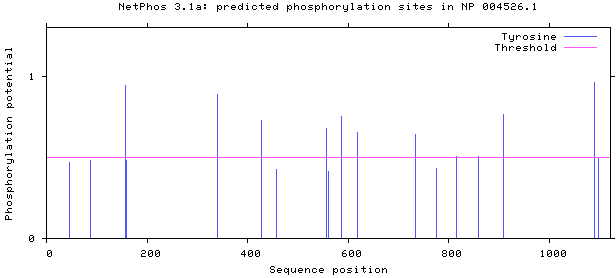
ProtComp v9.0 was used, as shown in Table 4. MyT1 is located in nuclear at subcellular level.
Coiled coil analysis
COILS was used, as shown in Fig. 10. The coiled coil is at sites 291–344, 1033–1091.
Structural domain analysis
SMART was used, as shown in Fig. 9 and Table 5. MyT1 has seven zinc finger domains.
Position of MyT1 Conservative structural domain
Position of MyT1 Conservative structural domain
Prediction of conservative structural domain of MyT1.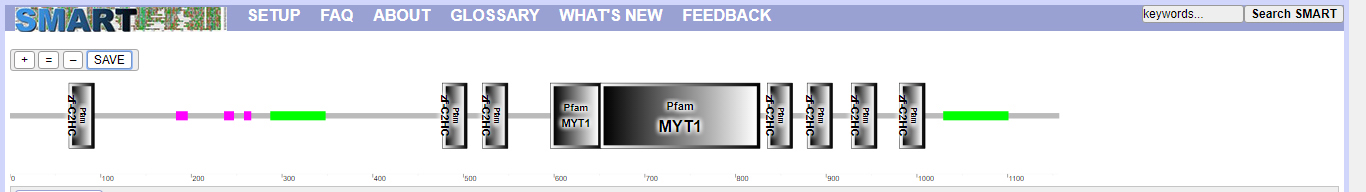
Prediction of MyT1 Coiled coil.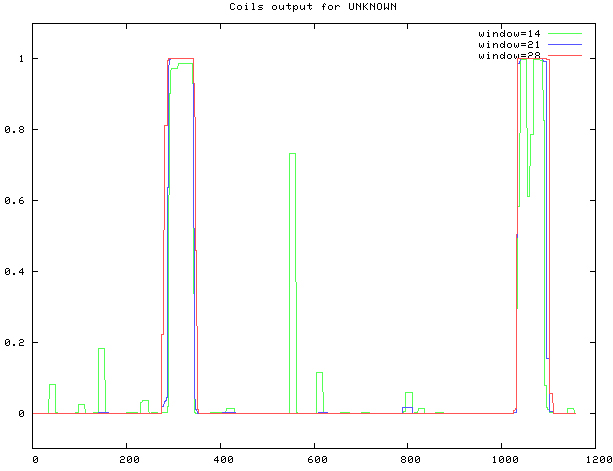
Predict protein secondary structure
SOPMA was used, as shown in Fig. 11. The secondary structure consisted of 36.99% alpha-helical, 7.95% extended strand, 2.25% beta-turn and 52.81% random coil.
Prediction of the secondary structure of MyT1.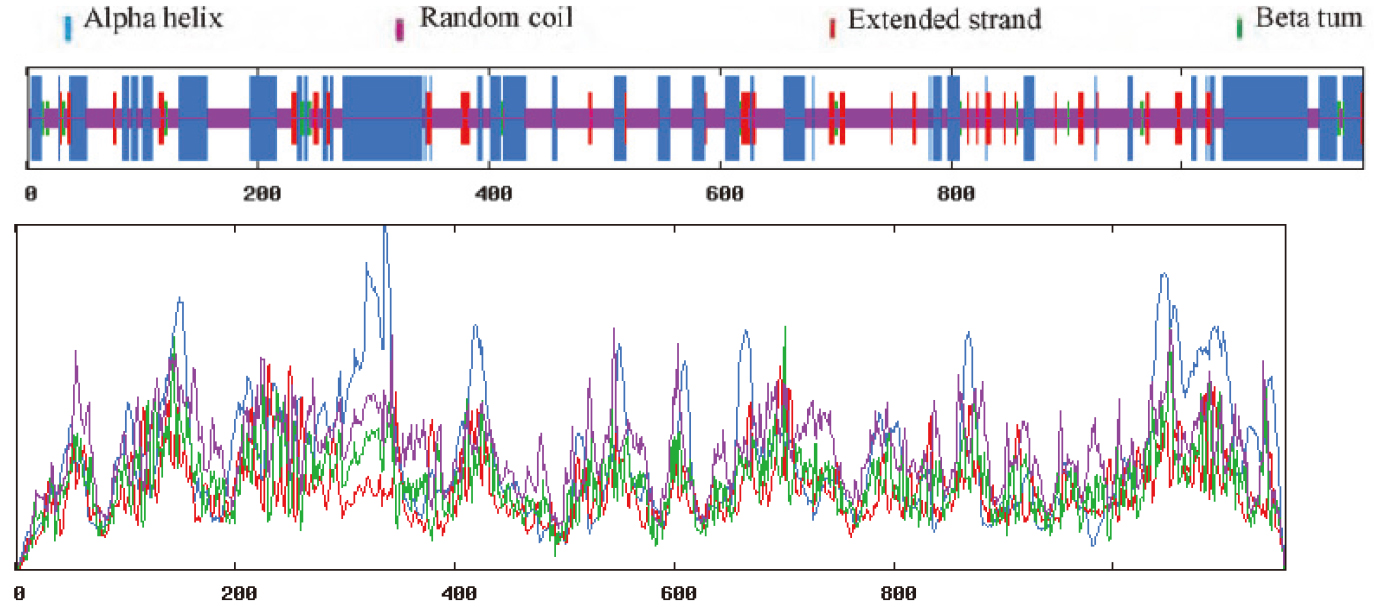
SWISS MODEL was used, as shown in Fig. 12. The score of GMQE is 0.04.
The tertiary structure prediction of MyT1.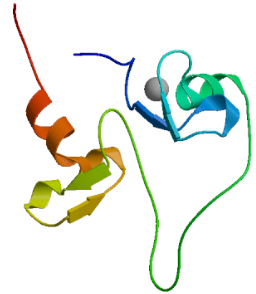
The present study showed that MyT1encodesed 1157 amino acids, the most abundant amino acid is glutamic acid, the relative molecular weight was 126447.96, the isoelectric point (pI) was 4.91, negative, the molecular formula C
MyT1 is an important regulatory
We studied the structure and function of MyT1 based on bioinformatics, the results show that t MyT1 is found to be an unstable hydrophilic protein, rather than a secretory protein, with no signal peptide or trans-membrane domain. The total amino acids are located on the surface of the cell membrane. It contains seven zinc finger domains structurally. At sub-cellular level, MyT1 is localized in the nucleus. The phosphorylation site mainly exists in serine, and its secondary structure is mainly composed of random coils and alpha helices; the three-dimensional structure is analyzed by modeling.
In conclusion, MyT1 is an unstable hydrophilic protein, which may be induced by environmental conditions. Furthermore, we can judge that transcription regulation may be induced by environmental conditions. There is no trans-membrane domain, demonstrating that it is not a secretory protein, which is consistent with its biological function as a transcription factor. Zinc finger domains regulate
Footnotes
Acknowledgments
This work was supported by a project grant from the Tianjin Key Technology R&D Program (No. 17ZXRGGX00020) and the National Natural Science Foundation of China (Grand No. 81801240).
Conflict of interest
None to report.
