Abstract
Rooting and root development in Acer rubrum have important effects on overall growth. A. rubrum does not take root easily in natural conditions. In this study, the mechanisms of the miR396b-GRF1 module underlying rooting regulation in A. rubrum were studied. The subcellular localization and transcriptional activation of miR396b and its target gene growth regulating factor 1 (GRF1) were investigated. These experiments showed that GRF1 was localized in the nucleus and had transcriptional activation activity. Functional validation experiments in transgenic plants demonstrated that overexpression of Ar-miR396b inhibited adventitious root growth, whereas overexpression of ArGRF1 increased adventitious root growth. These results help clarify the molecular regulatory mechanisms underlying adventitious root growth in A. rubrum and provide some new insights into the rooting rate in this species.
Introduction
Acer rubrum L., a large deciduous tree in the family Sapindaceae, 1 is a high-quality ornamental tree with colorful leaves. As color-leafed trees have become increasing popular in landscaping applications, the adaptability of introduced A. rubrum has become a hot research topic. 2 Because sexually propagated A. rubrum seedlings frequently lose parental traits, breeding programs have to propagate this tree asexually. 3 At present, A. rubrum is primarily propagated using tender cuttings, which are extremely difficult to root. 4 Low rooting rates lead to decrease in branch-survival rates, which pose a challenge to the rapid propagation of A. rubrum and its large-scale application in landscape greening. 5 Although rooting and root development have important effects on the overall growth of A. rubrum specimens, there is a lack of research on the related molecular mechanisms. 6 It is therefore important to explore molecular mechanisms underlying root formation in A. rubrum to improve the cutting rooting rate and reduce propagation cost.
Auxin indoleacetic acid (IAA) was the first plant hormone used to stimulate the rooting of cuttings. Then, indolebutyric acid (IBA), a second auxin analog, was found to promote rooting more effectively than IAA. At present, auxin has been shown to play a central role in adventitious root formation in model plants, and in actual production, synthetic auxin analogs are often used to induce adventitious root development in difficult-to-root woody plants.
Rooting and root development are regulated by miR396 and GRF. miR396 is highly conserved in many species, and miR396 is crucial in rooting. 7 Different miR396 members may play different roles in regulating root development. 8 In Arabidopsis thaliana, overexpression of miR396a and miR396b under the control of a strong 35S promoter has been shown to have different effects on root length. However, in wild-type Arabidopsis, 35S::MIR396a was found to exhibit a longer root phenotype, while 35S::MIR396b showed no significant change in root length. 8 In Larix leptolepis, transgenic plants overexpressing pre-miR396 had abnormal root tip tissue and a low rooting rate. 9 In Tribulus sativa, miR396 negatively affected root growth, the size of the root apical meristem (RAM), and the proliferation of apical cells, with primary root length and dry weight significantly reduced, RAM length considerably reduced by 37.5%, and the percentage of replicating cells in roots significantly reduced by 7.89% in plants overexpressing miR396. 10 Some GRF genes and protein are also expressed in roots. For example, 7 of the 9 At-GRF genes regulate the root meristem, whereas 2 of the 12 OsGRF genes are expressed in the root. 11 In L. leptolepis, downregulating the expression of LaGRFs reduced the rooting rate, suggesting that LaGRFs might positively regulate the rooting of L. leptolepis. 12 In Populus, PTGRF1/2D was mainly expressed in the pericyclic and apical root positions, indicating that these genes participated in root formation. 13
Studies in apples found that MdGRF11-overexpressing strains cultured under the same conditions had longer taproots than wild-type strains, suggesting that overexpression of MdGRF11 enhanced the growth of young roots. 14
miR396 mainly targets a class of important growth regulating factors (GRFs). Homeostasis is established between miR396 and its target genes GRF3 and GRF31 via mutual feedback regulation, and GRF3/GRF1 and miR396 have a negative regulatory relationship. 7 Overexpression of At-miR396a reduces the transcription levels of target genes such as GRF and b HLH74 (BASIC HELIX-LOOP-HELIX74), resulting in a shorter root phenotype. 8 MiR396-OE transgenic A. Arabidopsis thaliana was found to inhibit root growth, possibly because miR396 downregulated GRF expression, thereby disrupting the balance of cell-cell interactions. 15
miR396 and its target GRFs play a crucial role in the growth and development of plant roots. Although the molecular regulatory mechanism associated with the miR396-GRF module has been reported in some plants, especially in model plants such as A. thaliana and Populus L., it has not been reported in A. rubrum. In this study, we aim to comprehensively study the regulatory roles of miR396b-GRF1 in adventitious root growth and provide new insights into the improvement of the rooting rate in A. rubrum.
Materials and Methods
Plant materials
In this study, we used the A. rubrum hybrid “Autumn Fantasy.” Our preliminary analyses showed that treatment with 300 mg/L IBA for 1 hour was optimal and significantly improved the rooting of A. rubrum. 16 Cuttings (8-10 cm long) were divided into two groups. Cuttings in the first group (CK group) were soaked in deionized water for 1 hour. before cutting; cuttings in the second group (treatment group) were treated with 300 mg/L IBA for 1 hour. before cutting. After 10 days of culture in the greenhouse, phloem samples were taken within 3 cm of the base of each cutting for follow-up tests. Phloem samples were stored in an ultra-low temperature freezer at −80°C for later use.
Phylogenetic and conserved motif analysis of ArGRF1
The amino acid sequences of ArGRF1 were compared using the National Center for Biotechnology Information (NCBI) database. Protein sequences with greater than 50% homology with ArGRF1 were aligned using MEGA 7, 17 and a phylogenetic tree was then constructed using the neighbor-joining (NJ) method. The conserved motifs of the ArGRF1 protein were detected using MEME online tool (http://meme-suite.org/tools/meme). The parameters used for the MEME analysis were as follows: number of repetitions = any, maximum number of memes = 10, minimum width = 6, and maximum width = 200.
Differential expression analysis of the GRF family in A. rubrum
Gene expression data for GRF family members were screened from the transcriptome data of A. rubrum. Genes were considered differentially expressed when the |log2FoldChange| was >1 and the P-value was <.05. A matrix file was uploaded to R software to generate a heatmap of the GRF family in A. rubrum.
Quantitative reverse-transcription PCR (qRT-PCR) validation
Ar-miR396b and its target gene ArGRF1 were validated using qRT-PCR. The polysaccharide polyphenol RNA rapid extraction kit (Aibosen, Beijing, China) was used to extract total RNA from the phloem samples in the CK and IBA 300 mg/L groups. Reverse transcription kits, such as the TransScript miRNA First-strand cDNA Synthesis Super Mix Kit (TransGen, Beijing, China) and the Transcript One -step gDNA Removal and cDNA Synthesis Super Mix Kit (TransGen, Beijing, China), were then used to reverse transcribe RNA into cDNA qRT-PCR was performed with 2×SYBR GREEN (Aibosen, Beijing, China). The conditions for PCR amplification were as follows: 95°C for 2 minutes, followed by 40 cycles of 94°C for 15 seconds, 60°C for 15 seconds, and 72°C for 20 seconds. Each reaction was performed 6 times in parallel. The reference gene used for the Ar-miR396b analysis was U6. 18 The reference gene used for the ArGRF1 analysis was Actin 6. 19 Table 1 presents the qRT-PCR primers used in this study.
Primers used for qRT-PCR.

Abbreviations: F, forward primer; R, reverse primer.
Subcellular localization and transcriptional activation
In this study, we established a transient transformation system in tobacco leaves, and transient expression in tobacco leaves was judged based on the subcellular localization of transient green fluorescent protein (GFP) expression. 20 We constructed a recombinant expression vector that contains pBWA(V)HS—Glosgfp and the coding sequence (CDS) of the ArGRF1 gene, and we transferred this vector into Agrobacterium competent cells (GV3101) using the freeze-thaw method. Positive strains were cultured and screened. The selected positive strains were mixed with the infection solution to make a suspension, and the OD value was adjusted to 0.8-1.2. Robust, 5-6-week-old tobacco plants with good growth and no flowers were selected. A small hole was scraped under the epidermis of the tobacco leaves with a needle. Then, the infected solution was extracted with a needleless syringe and injected into the epidermis of the tobacco leaf. The injected tobacco plants were cultured in darkness for 16 hours, and then grown in the light for 1 to 2 days. Finally, the epidermis was removed and prepared for observation.
The transcriptional activation technique was used on ArGRF1 to verify its transcriptional activation activity. Firstly, a recombinant expression vector composed of pGBKT7 and ArGRF1 gene CDS was constructed. After successful construction, it was transformed into AH109 yeast receptor cell, an experimental strain of the GAL4 system developed by Clontech, which was coated on SD-Trp solid medium after transformation and cultured inverted at 29°C for 2 to 3 days. Monoclonal strains were selected for PCR verification. The positive results were inoculated in SD-Trp liquid medium and cultured until the liquid medium was cloudy. An appropriate amount of the liquid medium was taken and diluted with ddH2O to OD600 = 0.1. then absorbed 1 µL, sampled on SD-Trp monodeficiency medium, and cultured at 29℃ until the colony grew. Appropriate amount of solution was absorbed and coated into double-shortage medium containing SD-His-Trp-3AT (3AT concentrations were 25 mM), and incubated at constant temperature. Among them, HIS is the reporter gene that needs to be detected using the HIS defect screening media. 3-AT is a competitive histidine inhibitor that inhibits leakage expression and slight autoactivation of HIS3 to avoid false positive test results, Appropriate colonies were then selected for observation and photographic record.
Gene cloning and transgenic functional verification
DNA was extracted from the A. rubrum phloem samples using the CTAB plant genome DNA rapid extraction kit (Aidlab, Beijing, China), following the manufacturer’s instructions. Pre-miR396b was cloned from the DNA extracted from the phloem samples. Total RNA was extracted from A. rubrum by grinding samples in liquid nitrogen. RNA was extracted using the polysaccharide polyphenol RNA rapid extraction kit (Aibosen, Beijing, China). The total RNA extracted from the phloem was reverse transcribed, and first-strand cDNA was synthesized using TransScript One-step gDNA Removal and cDNA Synthesis Super Mix (TransGen, Beijing, China). ArGRF1 was cloned from the cDNA. The overexpression vectors PCAMBIA1304-Ar-miR396b and PCAMBIA1304-ArGRF1 were constructed using the Nimble Cloning technique. The constructed overexpression vectors were transformed into Agrobacterium GV3101.
Wild-type A. thaliana (Columbia-0) was transformed using the inflorescence infection method. An appropriate number of seeds from the T2 generation and from wild-type A. thaliana were dried and stored at 4°C. After storage, seeds were placed in the medium and vernalized at 4°C for 3 days. After 5 hours of light induction, seeds were wrapped in 3 layers of tinfoil and cultured at 19°C in darkness for 4 days until seedling hypocotyl length reached 6 mm on average. All the root organs of the A. thaliana seedlings were removed with a sterilized blade on an ultra-clean workbench; only the hypoembryonic axis (6 mm in length) remained of the lower part of the seedling. The treated A. thaliana seedlings were placed on 1/2 MS medium with tweezers and cultured under long-day conditions to induce adventitious root formation. 21 For each strain, we performed 10 biological replicates and 2 technical replicates. Daily changes in adventitious root length and quantity were photographed and recorded; specific quantitative characters were measured using ImageJ. 22 Statistical analysis was performed with GraphPad8.
Ar-miR396b and ArGRF1 were verified using qRT-PCR. Total RNA was extracted from the adventitious roots of wild-type A. thaliana, transgenic Ar-miR396b, and transgenic ArGRF1 plants using Trizaol RNA Extraction Kits (Aidlab, Beijing, China). TransScript miRNA first-strand cDNA Synthesis Super Mix (TransGen, Beijing, China) was used to reverse transcribe cDNA from the total RNA extracted from transgenic Ar-miR396b plants. Transcript one-step gDNA Removal and cDNA Synthesis Super Mix (TransGen, Beijing, China) was used to reverse transcribe cDNA from the total RNA extracted from transgenic ArGRF1 plants.
qRT-PCRs were conducted using 2×SYBR GREEN (Aibosen, Beijing, China). The PCR cycling conditions were as follows: 95°C for 2 minutes; followed by 40 cycles of 94°C for 15 seconds, 60°C for 15 seconds, and 72°C for 20 seconds. Each reaction was performed 6 times in parallel. The reference gene for Ar-miR396b was U6, 18 and the reference gene for ArGRF1 was 18S rRNA. 23 The qRT-PCR primers used are listed in Table 1.
Results
Phylogenetic analysis of ArGRF1
A phylogenetic tree of A. rubrum and 16 other plants was constructed using MEGA 5.0. As shown in Figure 1A, ArGRF1 is closely related to PvGRF1 of Pistacia vera, and they are clustered into one class. ArGRF1 mainly consists of 2 conservative domains, QLQ, and WRC. The QLQ domain contains Motif5, Motif7, Motif9, and Motif16. The WRC domain contains Motif3, Motif6, and Motif8 (Figure 1B).

Phylogenetic and conserved motif analysis. (A) Phylogenetic analysis. Bootstrap estimates based on 1000 replicates are indicated by numbers on the branches. The gene marked in red is ArGRF1 (A). (B) Conserved motifs of the ArGRF1 protein. Colored boxes, numbered 1 to 20 on the right, denote each motif. The sequence of motifs is arranged from 1 to 20 according to the size of the genome.
Differential expression analysis of the GRF family in A. rubrum
Twenty differentially expressed GRFs were analyzed, of which 6 were down-regulated including ArGRF2, ArGRF8, and ArGRF20, 14 were upregulated, including ArGRF1, ArGRF12, and ArGRF16. The gene marked in red is ArGRF1. as shown in the figure, ArGRF1 was upregulated after IBA treatment (Figure 2).

GRF family hierarchical clustering heat map of A. rubrum. CK indicates: cuttings (8-10 cm long) of the A. rubrum hybrid “Autumn Fantasy” that were soaked in deionized water for 1 hour before cutting. IBA300 indicates: cuttings that were treated with 300 mg/L IBA for 1 hour before cutting. The gene marked in red is ArGRF1.
qRT-PCR validation of Ar-miR396b and ArGRF1
qRT-PCR analysis showed that, after treatment with 300 mg/L IBA, the expression patterns of Ar-miR396b and ArGRF1 exhibited an inverse relationship. Compared with CK, the relative expression of Ar-miR396 decreased, while that of ArGRF1 increased (Figure 3).
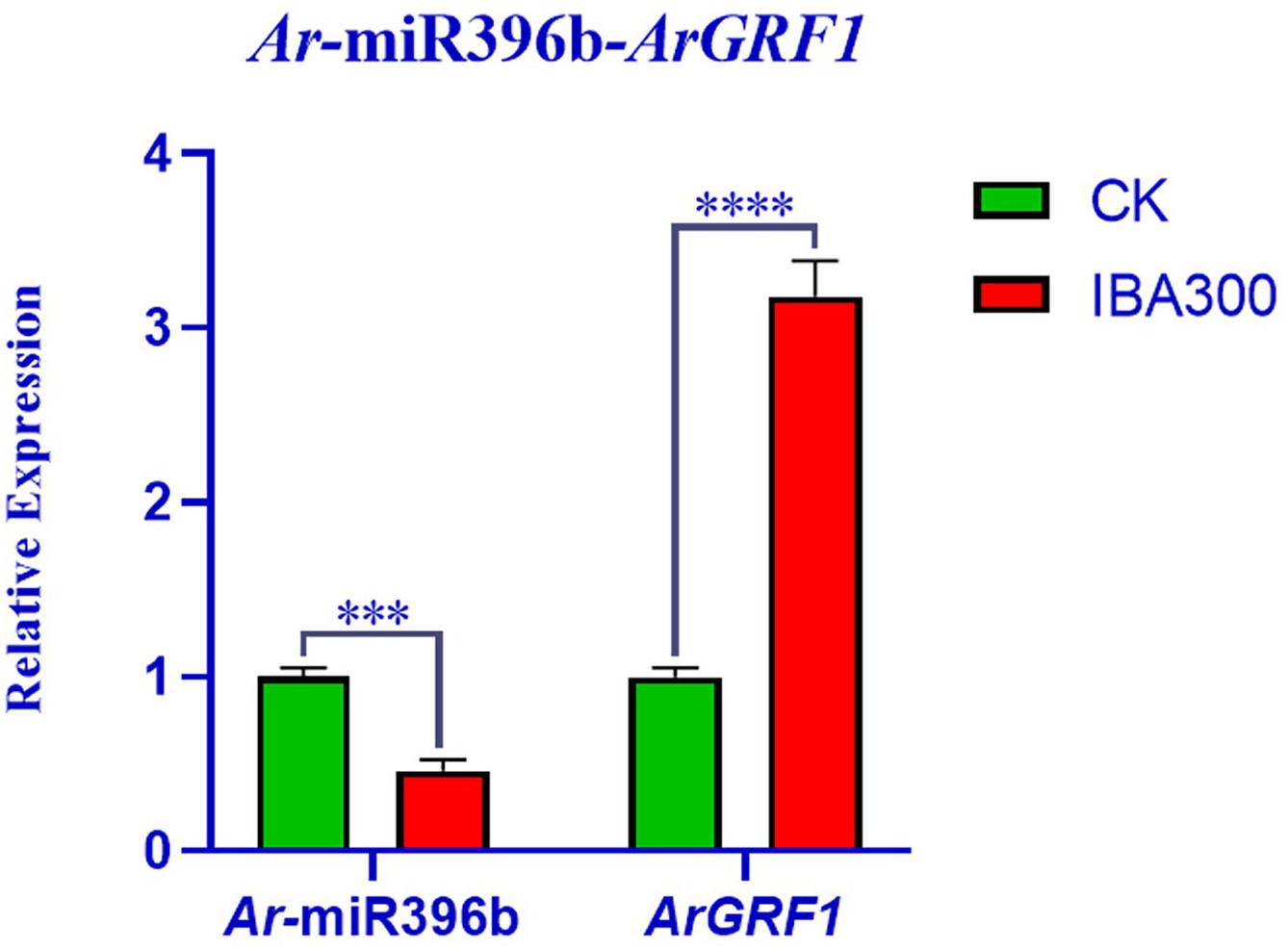
Expression profiles of Ar-miR396b and ArGRF1 after treatment with 300 mg/L IBA. The expression of the control (CK) was adjusted to 1.0. Bars represent the standard error of 3 repetitions.
Subcellular localization and transcriptional activation
Fluorescent protein labeling was used to investigate whether ArGRF1 was a nuclear localization protein like other transcription factors. The findings revealed that empty GFP was dispersed throughout the cell, whereas GFP-ArGRF1 fusion protein was only expressed in the nucleus, where green fluorescence was evident, which was consistent with predictions (Figure 4).

Subcellular localization of ArGRF1 protein in that leaf epidermal cells of tobacco. From left to right are green fluorescent protein fields (GFP), nuclear marker field, bright fields, and merged pictures. The first row is the control group, and the second row is the experimental group. Excitation light, 488 nm; emitted light, 510 nm. Scale bar = 20 μm.
The transcriptional activation results are as follows. In SD/-Trp monodeficient medium, strains in both control BD and experimental BD-ArGRF1groups could be grown, which demonstrated that BD plasmid was transferred into yeast strains. However, in In SD/-Trp-His+3AT (25 mM) double-deficient medium, only strains in the experimental group that carried pGBKT7-ArGRF1 recombinant vector plasmid could grow, while strains in the control group could not grow, which demonstrated that ArGRF1 could activate the transcription of His reporter gene, and that this transcription is only affected by ArGRF1 (Figure 5).

Results of the transcriptional activation of the ArGRF1 in yeast. BD was the negative control; and pGBKT7-53 was the positive control; SD/-Trp and SD/-Trp-His refer to nutrient deficiency medium.
Overexpression of Ar-miR396b and ArGRF1 in A. thaliana
T3 generation A. thaliana was cultured on 1/2 MS medium for 10 days. After 10 days of growth, adventitious root number and length were recorded, and the adventitious root phenotypes were photographed. ArGRF1 and Ar-miR396b respectively promoted and inhibited A. thaliana adventitious root growth (Figure 6A). OE-miR396b had fewer adventitious roots during 10 days of culture, while OE-ArGRF1 had more adventitious roots (Figure 6B). Statistical analysis of the numbers of adventitious roots of A. thaliana on day 10 showed that OE-miR396b had fewer roots than the control and OE-ArGRF1 had more roots than the control, but these differences were not significant (Figure 6C). The adventitious roots of OE-miR396b were shorter and those of OE-ArGRF1 were longer during the 10 days of culture (Figure 6D). Statistical analysis of A. thaliana adventitious root lengths on day 10 showed that the adventitious roots of OE-miR396b were significantly shorter than those of the control, while the adventitious roots of OE-ArGRF1 were considerably longer than those of the control (Figure 6E). The expression levels of OE-Ar-miR396b and OE-ArGRF1 in the T3 generation transgenic strains were detected using qRT-PCR. The analysis of the data indicated that the expression levels of Ar-miR396b and ArGRF1 were significantly higher in transgenic A. thaliana than in Col-0 (Figure 6F and G).

Overexpression of Ar-miR396b and ArGRF1 respectively inhibited and promoted adventitious root growth in Arabidopsis thaliana. (A) Comparison of the adventitious root phenotypes of Col-0, OE-Ar-miR396b, and OE-ArGRF1 plants after 10 days of culture. Bar = 1 cm. (B) Comparison of the average numbers of adventitious roots among Col-0, OE-Ar-miR396b, and OE-ArGRF1 during 10 days of culture. (C) Comparison of the numbers of adventitious roots among Col-0, OE-Ar-miR396b, and OE-ArGRF1 on day 10 of culture. (D) Comparison of adventitious root lengths among Col-0, OE-Ar-miR396b, and OE-ArGRF1 during 10 days of culture. (E) Comparison of adventitious root lengths among Col-0, OE-Ar-miR396b, and OE-ArGRF1 on day 10 of culture (**P < .01, n = 3). (F and G) The expression levels of OE-Ar-miR396b and OE-Ar-GRF1 in transgenic A. thaliana. The reference gene for Ar-miR396b was U6. The reference gene for ArGRF1 was 18S rRNA. Error bars indicate SD.
Discussion
miRNAs targets are mostly transcription factors, which are of great significance in the regulation of plant growth and development. Different transcription factor families regulate a series of complex metabolic networks that can help plants grow under normal and adverse environmental conditions. To date, transcription factor regulating root growth have been identified primarily in the myeloblastosis oncogene (MYB), WRKY, NAC (NAM, ATAF1, 2 and CUC2), GRF, and auxin response factors (ARF) families 24 ; the GRF family has mostly been implicated in root growth, flower development, and seed development.25 -27 miRNAs interact with transcription factors and other target genes to influence regulatory networks related to root development in the root transcriptomes and make key functions in the final translation of mRNA. 28 Most GRF genes are the target genes of miR396, and the miR396-GRF pathway is considered one of the main pathways to control organ size. 27 In A. thaliana, 2 genes from the miR396 family, miR396a and miR396b, were predicted and shown to regulate GRF. 29 The repression of MtGRF expression by miR396 in Medicago truncatula was accompanied by root-growth inhibition and a decrease in root apical meristem size due to alterations in cell cycle activity. RNA interference-mediated silencing of MtGRF2 and MtGRF4 also caused short-root phenotypes, implying that the effects of miR396 on root growth were mainly due to changes in MtGRF mRNA levels. 30
In this study, the phylogenetic relationships and conserved motifs of ArGRF1 were analyzed. The phylogenetic results showed that ArGRF1 was closely related to GRF1 from P. vera. Two conserved domains, QLQ and WRC, were found to be mainly present in ArGRF1, similar to PeGRF1 in Phyllostachys edulis. These domains, initially discovered in rice and A. thaliana GRFs, are remarkably consistent across all plant species. 27 The combination of WRC and QLQ domains is unique to GRFs. The QLQ domain is evolutionarily ancient compared to the WRC domain. The QLQ domain was named after the conserved glutamine-leucine-glutamine motif and is distinguished by the typical arrangement of this motif (QX3LX2Q) with aromatic, hydrophobic, and acidic amino acid residues. 31 The QLQ domain is found at the N-terminus of the SWI2/SNF2 protein, which has been identified as a mediator of protein-protein interactions. 32 This domain has thus been postulated to be involved in mediating protein interactions. The WRC domain, named after the conserved tryptophan–arginine–cysteine motif, contains 2 distinctive features: a putative nuclear localization signal and a zinc-finger motif (C3H). 33 The WRC domain may play a role in DNA binding. 25
In A. rubrum, different ArGRF family members showed different expression patterns under IBA treatment, suggesting that ArGRF family members are induced by IBA and play an important function in the hormone signal transduction pathway. ArGRF1 was upregulated after IBA treatment, and it is proposed that ArGRF1 plays a critical role in the growth of the adventitious roots in A. rubrum.
qRT-PCR validation of Ar-miR396b and ArGRF1 showed that the relative expression level of Ar-miR396 decreased and that of ArGRF1 increased after IBA treatment compared with the CK, indicating that Ar-miR396b and ArGRF1 may have complementary expression patterns. The miR396b-GRF1 network may play a significant role in A. rubrum rooting in response to IBA.
Localization of proteins within plant cells is key to a better understanding of protein function, gene regulation, and protein-protein interactions. 34 Previous studies showed that all HvGRF family members in barley were localized in the nucleus. 35 Here, we confirmed that ArGRF1 was primarily located in the nucleus, as is typical of transcription factors, using instantaneous expression to visualize subcellular localization patterns in a tobacco leaf. Previous studies have shown that OsGRF1 in rice has transcriptional activation activity and is a transcriptional activator. Using the GAL4 system in O. sativa, OsGRF1 was shown to fuse with the GAL4 DNA binding domain, thereby activating the HIS reporter gene and allowing yeast cells to grow on histidine-deficient medium. 11 A transcriptional activation experiment was used to verify the transcriptional activation activity of ArGRF1. The results indicated that ArGRF1 can activate the transcription of HIS reporter genes. ArGRF1 had transcriptional activation activity and thus might play an important role as a transcription factor in A. rubrum adventitious root growth.
Ar-miR396b and ArGRF1 overexpression vectors were further constructed and transfected into A. thaliana. Transgenic A. thaliana plants were successfully obtained. qRT-PCR analysis showed that, compared with the wild type, Ar-miR396b and ArGRF1 were significantly upregulated in T3 generation transgenic A. thaliana. This indicated that Ar-miR396b and ArGRF1 had been successfully transferred into A. thaliana and overexpressed.
OE-Ar-miR396b plants exhibited noticeable phenotypic changes compared to WT plants, including reduced adventitious root length. Consistent with this, M. truncatula root growth was inhibited by overexpression of the miR396b precursor; transgenic plants overexpressing the mtr-miR396b precursor (miR396-OE) had significantly reduced primary root lengths and dry weights. 10 To date, GRFs have been shown to promote root growth in transgenic A. thaliana overexpressing GRFs from Brassica rapa, as well as in B. oleracea, rice, and M. truncatula.10,36 -38 Compared to control plants, transgenic B. oleracea rGRF3#10 had greater primary root elongation rates and longer roots. 37 RNAi-mediated inactivation of MtGRF2, MtGRF4, and MtGRF6 in M. truncatula significantly reduced root length and weight, suggesting that MtGRF2, MtGRF4, and MtGRF6 may promote root growth. 10 In O. sativa, OsGRF6 positively regulated root length, and the roots of the OsGRF6 mutant were much shorter than those of the wild type. 38 A. thaliana overexpressing B. rapa GRF genes (BrGRF3, BrGRF5, BrGRF7, BrGRF8, and BrGRF9) had longer primary roots than wild-type plants of the same age. 36 Together, these previous studies show that GRFs can act as positive regulators of plant root growth.
Here, ArGRF1 significantly promoted the growth of adventitious roots in A. thaliana, and transgenic plants overexpressing ArGRF1 (OE-ArGRF1) had significantly longer adventitious roots than the WT. In M. truncatula, a GRF gene containing miR396 binding site has been shown to be cleaved in roots. 39 The expression of GRF1 was regulated by the miR396b, 40 and miR396b-GRF1 regulatory modules are functionally conserved in plants.40,41 The cleavage of the GRF1 gene by miR396 mainly occurs between nucleotides 10 and 11 in the miR396 pairing region, and this process is seen in root tissues.7,42 Root development in A. thaliana is negatively affected by overexpression of miR396a and miR396b. 43 In transgenic A. thaliana overexpressing GRF1, the abundance of pri-miR396b and mature miR396 decreased significantly, while in GRF1/GRF1/GRF3 triple knockout mutants, the abundances of primary transcripts and mature miR396 were significantly increased25,43 The data indicate that there is a reciprocal regulatory relationship between miR396 and its target gene GRF1, which establishes an internal environmental balance at the post-transcriptional level. 43 Therefore, we speculated that ArGRF1 promoted the growth of adventitious roots and that Ar-miR396b inhibited the expression of ArGRF1 by directed cleavage, which in turn inhibited rooting. Ar-miR396b negatively regulated the mRNA abundance of its target gene ArGRF1, and, in turn, ArGRF1 negatively regulated the expression of Ar-miR396b. Therefore, ArGRF1 negatively regulates Ar-miR396b expression in A. thaliana overexpressing ArGRF1, thereby increasing adventitious root length. In A. thaliana overexpressing Ar-miR396b, Ar-miR396b significantly downregulated ArGRF1, inhibiting rooting and reducing adventitious root length.
Through a comprehensive analysis of the transcriptome, sRNA, and degradome data, GRF1 was identified as the target of miR396b. After IBA treatment, the expression of miR396b was downregulated and the expression of GRF1 was upregulated. In addition, according to the representative type of T3 generation transgenic A. thaliana, it could be observed that miR396b inhibited the growth and development of adventitious roots, whereas GRF1 promoted the growth and development of adventitious roots. Furthermore, these theoretical analyses were consistent with the experimental results and could be applied to other species. In future studies, the identification of protein interaction will be conducted to further verify the function of ArGRF1. Thus, these results are of great scientific significance, as they clarify the molecular mechanism of Ar-miR396b and ArGRF1 in regulating rooting in A. rubrum, as well as provide a basis for the improvement of A. rubrum rooting ability through molecular breeding to increase its utility in landscape applications.
Conclusion
Through a series of bioinformatics analyses and experimental verification, this study demonstrated that ArGRF1 was a transcription factor, and it also confirmed the negative regulatory relationship between Ar-miR396b and ArGRF1. Furthermore, the results of this study revealed the functions of Ar-miR396b and ArGRF1 and provided a basis for further study of molecular mechanisms on rooting in A. rubrum.
Footnotes
Acknowledgements
The authors hope to express their appreciation to the reviewers for this manuscript.
Author Contributions
Kezhong Zhang, Wei Ge, and Hewen Zhao designed the study; Manyu Zhang and Huiyu Zhu collected and prepared the materials; Manyu Zhang and Huiju Li conducted the experiments and data analysis and wrote the manuscript; Wei Ge revised the manuscript. All authors read and approved the final draft.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This project was supported by The Project of Cultivation for young top-notch Talents of Beijing Municipal Institutions (grant No. BPHR202203100) and Beijing Innovation Consortium of Agriculture Research System (grant No. BAIC09-2022).
Declaration of Conflicting Interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Consent for Publication
All authors agree to publish this paper.
Data Availability
Not applicable.
