Abstract
BACKGROUND:
LINC00885 is a novel oncogenic long noncoding RNA (LncRNA) which is upregulated in various types of cancer, but its function in triple-negative breast cancer (TNBC) remains unknown.
OBJECTIVE:
This study aimed to determine the role of LINC00885 on TNBC development.
METHODS:
Clinical interrelation and survival analysis were determined using online database. The CCK-8 and Transwell assays were used to detect the proliferation and migration behaviors in TNBC cell lines. The interaction among genes was detected by RNA pull down assay.
RESULTS:
LncRNA LINC00885 was highly expressed in TNBC compared to normal breast like. Low levels of LINC00885 was related to good prognosis in TNBC patients compared to TNBC patients with high LINC00885. LINC00885-downregulation inhibited, whereas LINC00885-overexpression promoted the proliferation and migration capability of TNBC cell lines. In TNBC cell lines, noncatalytic region of tyrosine kinase 1 (NCK1) expression was positively associated with LINC00885 expression, and shRNA-mediated the depletion of NCK1 significantly abolished LINC00885 upregulation-mediated pro-tumor effects. Combined with online databases, miR-654-3p was screened as the direct target gene of LINC00885, which could directly bind to 3’-untranslated regions (3‘-UTR) of NCK1, resulting in the decreased expression of NCK1 in TNBC cell lines. LINC00885 overexpression-mediated the upregulation of NCK1 was abrogated by miR-654-3p mimics. MiR-654-3p mimics significantly rescued the tumor promotive role caused by LINC00885-overexpression. However, exogenous NCK1 notably eliminated the anti-tumor effects caused by miR-654-3p mimics in LINC00885-overexpressed cells.
CONCLUSIONS:
LINC00885 is expressed at a high level in TNBC. LINC00885 promoted proliferation and migration by regulating the miR-654-3p/NCK1 axis in TNBC cell lines. Possibly, LINC00885 can be served as a potential therapeutic target for TNBC.
Keywords
Introduction
Breast cancer (BC) is the most widespread malignancy and the second leading cause of cancer-related death among the female population worldwide, accounting for
TNBC is more aggressive than other BC subtypes because it is associated with increased angiogenesis and epithelial-mesenchymal transition (EMT) [8, 9, 10]. At the molecular level, EMT alters the adhesion, polarity and differentiation of epithelial cells and renders them more prone to migration and invasion [10, 11, 12]. The noncatalytic region of tyrosine kinase 1 (NCK1) is an adaptor protein acting as a linker of the proximal and downstream signaling events that are related to the occurrence and development of a variety of tumors [13, 14]. NCK1 upregulation promotes the development of ovarian carcinoma (OC) [15, 16], colorectal cancer (CoC) [17] and cervical squamous cell carcinoma [18]. NCK1 specifically localizes in invadopodia but not in podosomes and can be used to recognize cell surface degradative structures of metastatic mammary carcinoma cells [19]. NCK is implicated in podosome formation and ECM degradation [20, 21]. Our previous study also indicates that NCK1 elevation promotes the migration of TNBC cell lines and NCK1 may serve as a diagnostic and prognostic marker of metastatic TNBC [22]. Therefore, the ECM degradation and the subsequent EMT process may be involved in NCK1-mediated cell migration.
Long non-coding RNAs (lncRNAs), a cluster of transcripts
In the current study, the expression of LINC00885 in patients with TNBC and TNBC cell lines was analyzed. Combining database analysis and RNA pull-down assay, the downstream target genes of LINC00885 were screened and verified. The effects of LINC00885 and its downstream signaling axis on cell growth and migration were also investigated. LINC00885 may serve as a prognostic and predictive marker and as a novel therapeutic target.
Materials and methods
Data collection and analysis
Breast Cancer Gene-Expression Miner v4.4 (
Subjects
Primer sequences of RT-PCR
Primer sequences of RT-PCR
A total of 40 pairs of TNBC and normal tissue samples were acquired from The First People’s Hospital of Pingdingshan, China. The present study was approved by the Ethics Committee of Pingdingshan University. Matched pericarcinomatous tissue was defined as tissue
A total of four TNBC cell lines (MDA-MB-468, BT549, Hs578t and MDA-MB-231) and one normal breast epithelial cell line (MCF-10A), were purchased from the Cell Bank of Type Culture Collection of Chinese Academy of Sciences. MDA-MB-468, Hs578t, MDA-MB-231 and MCF-10A cells were cultured in complete Dulbecco’s modified Eagle’s medium (cat. no. 11965092; Lot. no. 8121295; Gibco; Thermo Fisher Scientific, Inc.) and BT549 cells were cultured in RPMI 1640 medium (cat. no. 11875119; Lot. no. 8117293; Gibco; Thermo Fisher Scientific, Inc.) in a humidified incubator at 37
Reverse transcription-quantitative (RT-q) PCR
Total RNA was extracted from cultured cells and tumour tissue using TRIzol
Cell Counting Kit-8 (CCK-8) assay
Cell viability was assessed using the Cell Counting Kit-8 assay (CCK-8; cat. no. CA1210; Lot. no. 20220715; Solarbio), according to the manufacturer’s instructions. After transfection for 48 h, cells were plated into 96-well plates at the density of 5
Western blot analysis
Total protein from tumor cells was quantified using a bicinchoninic acid assay kit and 20
Dual-luciferase reporter gene experiment
The NCK1 and LINC00885 wild type (WT) and mutant (MUT) sequences of the 3’-untranslated regions binding to miR-654-3p were subcloned in the luciferase reporter plasmid pGL3.1 to synthesize NCK1-WT/-MUT and LINC00885-WT/-MUT plasmids, respectively. Plasmids were co-transfected with miR-654-3p mimic or negative mimic for 48 h. Subsequently, a dual-luciferase reporter assay system (cat. no. E1910; Lot. no. 0000449129; Promega Corporation, US) was used to assess the relative luciferase activity, according to the manufacturer’s protocol. Firefly luciferase activity was normalized to Renilla luciferase activity.
RNA pulldown
Relative LINC00885 expression in TNBC and its clinical significance. (A) The differential expression of LINC00885 in different BC Subtypes including basal-like (
Briefly, 1
Cells were inoculated in 6-well plates. After transfection and incubation, cells were digested with 0.25% trypsin and washed with phosphate buffer (PBS). After homogenized on ice, protein concentration was quantified by the BCA Protein Assay Kit (cat. no. 23227, lot. no. OB184645, Thermo Fisher Scientific, Inc.). LDH activity was measured by colorimetric assay using specific test kit (cat. no. BC0685, lot. no. 20220926, Solarbio, CN). The absorbance of LDH was measured at the wavelength of 450 nm. The activity of LDH was calculated according to the manufacturer’s introduction. The activity of LDH in control group was normalized to 1.0.
Statistical analysis
All assays were performed
Results
LINC00885 is upregulated in TNBC tumor tissues and cells
Firstly, the expression pattern of LINC00885 in different subtypes of BC was analyzed according to online databases. Increased LINC00885 expression was observed in patients with basal-like (
Effect of LINC00885 on the migration of triple-negative breast cancer cells. The plasmid overexpressing LINC00885, LINC00885 siRNA and the corresponding control vector/NC-siRNA were injected into BT549 and MDA-MB-231 cells. MDA-MB-231 (A) and BT549 cell (B) viabilities are determined using the Cell Counting Kit-8 assay. Transwell assay for pcDNA3.1-LINC00885 and siRNA-LINC00885 (C) MDA-MB-231 and (D) BT549 cells. Relative quantification of migrated cells is shown in the right panel (scale bar, 25 
Given the upregulation of LINC00885 in TNBC, the effects of LINC00885 on the malignant behaviors of TNBC cell lines were investigated. Firstly, LINC00885-overexpressed and LINC00885-knocked down MDA-MB-231 and BT549 cells were constructed (Fig. S1). A significant increase in cell growth was observed at 48 and 72 h in LINC00885-overexpressed MDA-MB-231 and BT-549 cells, whereas cell growth decreases in LINC00885-knocked down MDA-MB-231 and BT-549 cells, compared with that in the control group (Fig. 2A and B). These results demonstrated that the overexpression of LINC00885 could promote cell viability of TNBC cell lines, whereas LINC00885 knockdown could negatively impact the growth rate of TNBC cell lines (Fig. 2A and B). Consistently, Transwell migration assays also indicated that LINC00885 knockdown suppressed the migration of MDA-MB-231 and BT549 cells, whereas this was increased following the LINC00885 overexpression (Fig. 2C and D). Additionally, the levels of lactate dehydrogenase (LDH) that were detected using colorimetric analysis increased in LINC00885-overexpressed MDA-MB-231 cells and moderately decreased in LINC00885-doenregulated MDA-MB-231 cells (Fig. S2A). These results indicated that LINC00885 facilitated the viability and migration of TNBC cell lines in vitro and might also be related to glycolysis.
Association between LINC00885 and NCK1 in TNBC. (A) The expression of LINC00885 and NCK1 are determined by RT-qPCR. The association between LINC00885 and NCK1 expression in fresh TNBC specimens as analyzed using SPSS 22.0 software (IBM Corp.). (B) MDA-MB-231 and BT549 cells were transfected with pcDNA3.1-LINC00885, siRNA-LINC00885 and the corresponding negative controls pcDNA3.1 or NC-siRNA. NCK1 mRNA was detected by RT-qPCR in MDA-MB-231 and BT549 cells. 
NCK1 is a protein that has previously been reported to be closely associated with cancer progression [17, 18]. In the present study, NCK1 expression was positively correlated with LINC00885 expression in 40 patients with TNBC (Fig. 3A). In addition, mRNA expression of NCK1 was increased by 2.5-fold along with the increased LINC00885 expression, while NCK1 expression was effectively knocked down by 75% in MDA-MB-231 and BT549 cells (Fig. 3B). Moreover, NCK1 protein level increased following LINC00885 overexpression, while it decreased after LINC00885 knockdown in MDA-MB-231 and BT549 cells (Fig. 3C). These results indicated that NCK1 expression was positively associated with LINC00885 expression.
Effects of NCK1 on the migration of LINC00885-mediated triple-negative breast cancer cells. MDA-MB-231 and BT549 cells were transfected with pcDNA3.1, LINC00885-overexpression plasmid, pcDNA3.1-LINC00885 
Relationship between LINC00885 and miRNA. (A) Two databases predicting miRNAs that bound to LINC00885. (B) The binding sites between LINC00885 and miR-654-3p as analyzed using the ENCORI database (
Subsequently, the functional role of NCK1 inLINC00885-mediated alteration of cell proliferation and migration was investigated. Overexpression of LINC00885 significantly increased the viability of MDA-MB-231 and BT549 cells from 48 to 72 h (Fig. 4A). However, shRNA-mediated NCK1 knockdown decreased the cell viability of LINC00885 overexpression plasmid-transfected MDA-MB-231 and BT549 cell lines, compared with that of MDA-MB-231 and BT549 cells transfected with negative control shRNA and pcDNA3.1-LINC00885 plasmid (Fig. 4A). Consistently, LINC00885 upregulation promoted the migration of TNBC cell lines, which were not affected by the co-transfection of pcDNA3.1-LINC00885 and negative control shRNA (Fig. 4B and C). Once NCK1 was downregulated, cell migration and migrated cell number were restrained in LINC00885-overexpressing TNBC cell lines (Fig. 4B and C). In addition to that, LINC00885-overexpression-mediated the increase of LDH was also restored by the knockdown of NCK1 in MDA-MB-231 cells (Fig. S2B). Therefore, LINC00885 upregulation promoted cell viability, migration and LDH level in TNBC cell lines, that could be rescued by NCK1 knockdown.
Correlation among LINC00885, miR-654-3p and NCK1. (A) Study on the relationship between NCK1 and miR-654-3p in breast cancer using the ENCORI database (
Effect of LINC00885/miR-654-3p/NCK1 pathway on the migration of triple-negative breast cancer cells. MDA-MB-231 and BT549 cells were transfected with pcDNA3.1-LINC00885, pcDNA3.1-LINC00885 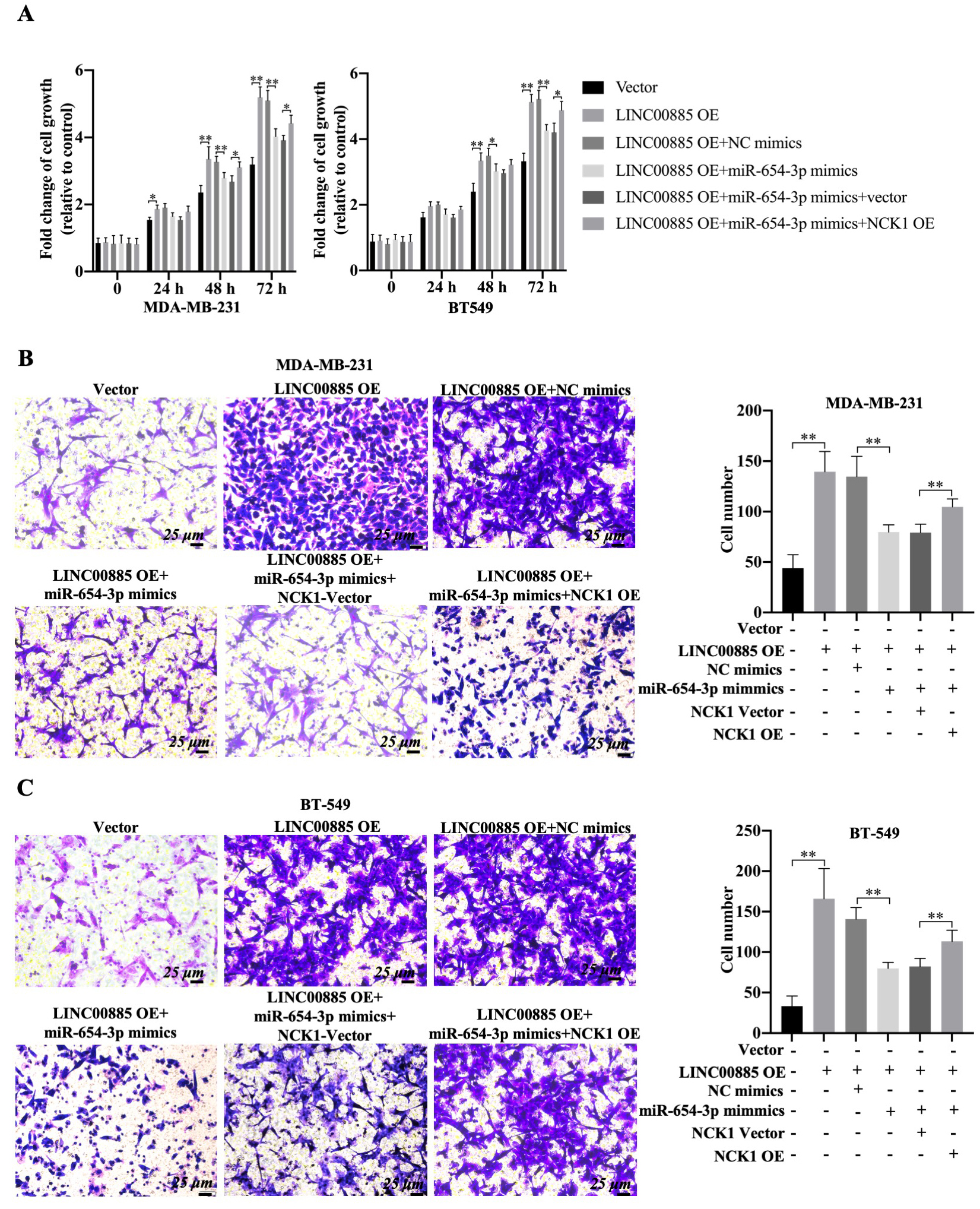
LncRNAs can function as ceRNAs to inhibit the degradation of mRNAs by sponging miRNAs in the cytoplasm of cells [35]. Thus, the present study investigated if LINC00885 could regulate NCK1 expression through the ceRNA mechanism by directly binding to miRNAs in TNBC cell lines. Using LncRNASNP and ENCORI databases, 17 miRNAs that could bind to LINC00885 were predicted and screened (Fig. S3A). Meanwhile, miRNAs binding to NCK1 were predicted using TargetScan, miRDB, miRWalk and ENCORI, and 14 candidate miRNAs were identified (Fig. S3B). The overlapping gene between the 17 miRNAs and 14 candidate miRNAs was miR-654-3p. Thus, miR-654-3p was screened as the only one candidate that could bind to LINC00885 and NCK1 (Fig. 5A). Moreover, the dual-luciferase reporter assay demonstrated that the luciferase activity in the cells transfected with the luciferase reporter plasmid LINC00885-WT was inhibited by miR-654-3p mimic (Fig. 5B). However, relative luciferase activity in cells transfected with LINC00885-MUT luciferase reporter plasmid was not affected by miR-654-3p mimic (Fig. 5B). These results, combined with the outcomes of the RNA pull-down assay, confirmed the direct binding of LINC00885 to miR-654-3p in MDA-MB-231 cells (Fig. 5C). In addition , miR-654-3p emerged as a potential negative correlation factor of LINC00885 in 40 pairs of TNBC and normal tissue samples (Fig. 5D). Moreover, LINC00885 overexpression also inhibited miR-654-3p expression, while LINC00885 knockdown increased the expression of miR-654-3p in MAD-MB-231 cells (Fig. 5E). These results indicated that miR-654-3p was the potential binding target of LINC00885 in TNBC cell lines.
MiR-654-3p is negatively correlated with NCK1 in TNBC cell lines
Based on the ENCORI database, miR-654-3p appeared to be not associated with NCK1 in patients with BC (BRCA,
Schematic diagram of the mechanism. LINC00885 upregulated NCK1 expression by sponging miR-654-3p in TNBC cell lines, thereby promoting cancer progression.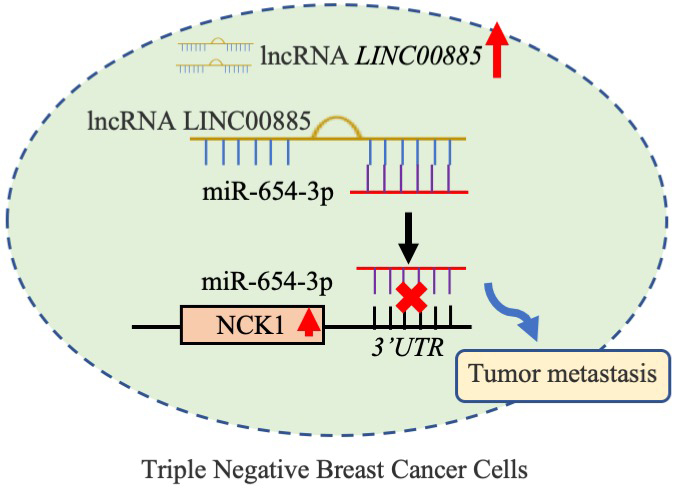
The functional roles of NCK1 and miR-654-3p in LINC00885-mediated alteration of cell migration were investigated. Using CCK-8 assay, overexpression of LINC00885 significantly increased the viability of BT549 cells at 48 and 72 h and promoted cell proliferation of MDA-MB-231 cells from 24–72 h (Fig. 7A). However, miR-654-3p mimic decreased the proliferation of LINC00885-overexpressing TNBC cell lines compared with that of TNBC cell lines transfected only with pcDNA3.1-LINC00885 (Fig. 7A). On the other hand, the proliferation inhibitory effect of miR-654-3p mimic was partially restored in TNBC that overexpressed both LINC00885 and NCK1 (Fig. 7A). In addition, the combined transfection of pcDNA3.1-LINC00885 and miR-NC did not affect the pro-migration role and the upregulation of LDH by LINC00885 overexpression (Fig. 7B, 7C and Fig. S2C). Once the miR-654-3p mimic was transfected, the cell migration and LDH level decreased in LINC00885-overexpressing TNBC cell lines (Fig. 7B, 7C and Fig. S2C). Upon NCK1 overexpression, the inhibitory effect of miR-654-3p mimics could be slightly restored. Therefore, LINC00885 overexpression promoted the proliferation and migration of MDA-MB-231 and BT549 cells. Possibly, the tumor-promoting properties of LINC00885 could be regulated by the miR-654-3p/NCK1 signaling axis.
Discussion
TNBC is a subtype of BC with a considerable migration characteristics and a high degree of malignancy and heterogeneity [36, 37]. TNBC is one of the most fatal cancers in females because of its poor prognosis and limited treatment options; moreover, blockade of cell migration is the primary approach for ameliorating TNBC [38, 39]. The present findings indicated that LINC00885 was highly expressed in TNBC cell lines and LINC00885 increased the tumor behaviors of TNBC cell lines through promoting NCK1 expression; therefore, LINC00885 could be a positive regulator of cell migration and aerobic glycolysis in TNBC cell lines.
LINC00885 is an oncogenic factor in cervical carcinoma (CeC) [40, 41] and early-stage BC [42]. Increased levels of LINC00885 in CeC tissues may be considered prognostic biomarkers [41]. These results suggested that LINC00885 had a high level of upregulation and was related to overall survival in patients with TNBC. LINC00885 promoted tumorigenesis and invasion of CeC cells [40, 43]. LINC00885 silencing in BC cells significative decreased cell migration and motility by downregulating the expression of proliferation-related transcripts such as epiregulin, MYC proto-oncogene and cyclin D1 [42]. The present functional analysis revealed that overexpression of LINC00885 could promote the migration of TNBC cell lines. These results showed that LINC00885 was involved in the migration of TNBC cell lines. Possibly, the abnormally increased expression of LINC00885 in TNBC might be related to the increased cell migration. Previous study also suggested that LINC01605 remarkably restrained cell proliferation and migration, as well as aerobic glycolysis by inhibiting LDH in TNBC cell lines [44]. Herein, LINC00885 also positively regulated the level of LDH in TNBC cell lines. Thus, the promotive role of LINC00885 on cell migration might also be related to the process of aerobic glycolysis.
The ceRNA acts as a miRNA ‘sponge’ and competes with miRNA-targeted mRNAs to influence miRNA-mediated gene regulation [45]. LINC00885 promoted tumorigenesis of colorectal cancer (CoC) in in vivo model by upregulating MACC1 expression as ceRNA for microRNA-432-5p [46]. LINC00885 promoted CoC progression by sponging miR-3150b-3p and upregulating bromodomain adjacent to zinc finger domain 2A [41, 42, 43, 46]. Thus, LINC00885 could regulate cell proliferation and migration via the ceRNA mechanism, thereby influencing the development of TNBC. MiR-654-3p inhibited non-small cell lung cancer tumorigenesis [47]. Previous study demonstrated that miR-654-3p might exert a suppressor role in hepatocellular carcinoma through inhibiting tumor cell proliferation, migration, and invasion [48]. The in vitro restoration of miR-654-3p decreased cell proliferation and migration and induced reprogramming of metastasis-related genes in Papillary thyroid carcinoma cells [49]. However, in the present study, miR-654-3p inhibition could reverse LINC00885-mediated the increase of proliferation and migration in TNBC cell lines. Therefore, LINC00885 induced cell viability and migration in TNBC cell lines by sponging miR-654-3p. In addition, the present results demonstrated that NCK1 expression was positively correlated with LINC00885 expression and miR-654-3p was negatively associated with NCK1. LINC00885 regulated downstream NCK1 expression by targeting mir-654-3p through the ceRNA mechanism. A previous study indicated that NCK1 regulated EGFR-induced cell migration and tumor migration with an on-off mechanism [50]. The migration-promoting effect of NCK1 was demonstrated in several types of cancer including BC [51]. In the present study, the miR-654-3p-mediated improvement of the tumor process could be counteracted by the upregulation of NCK1. Collectively, LINC00885 upregulated NCK1 expression by sponging miR-654-3p in TNBC cell lines, thereby promoting cancer progression. The LINC00885/miR-654-3p/NCK1 pathway may play an important role in TNBC cell migration (Fig. 8).
Although the present study identified a novel lncRNA associated with an increased malignant potential in TNBC, which may become a novel biomarker and potential therapeutic target for TNBC, there are still some drawbacks to be solved in future studies. Firstly, the present findings could not be further confirmed due to the lack of in vivo tumor migration models. Secondly, a single lncRNAs/miRNAs signaling axis is not sufficient to prompt a clinical application and change the status quo of clinical treatment. Simultaneously, further studies are needed to determine how NCK1 affects tumor cell migration.
Conclusion
In summary, the current study uncovered the promotive role of LINC00885 on cell migration in TNBC, elucidating a novel ceRNA network for LINC00885/miR-654-3p/NCK1 in TNBC cell lines. LINC00885 might become a new biomarker for the prediction of cell migration and the treatment of TNBC.
Author contributions
Conception: S.J.Y.
Interpretation or analysis of data: H.P.N., L.Z. and Q.J.X.
Preparation of the manuscript: H.P.N. and S.J.R.
Revision for important intellectual content: S.J.Y., L.Z. and Q.J.X.
Supervision: S.J.Y and H.P.N.
All authors had full access to the data in the study and take responsibility for the integrity of the data and the accuracy of the data analysis.
Funding
This work was supported by Henan Provincial Health Commission Project (Wjlx2021091 and Wjlx2021455).
Availability of data and materials
All data generated or analyzed during this study are included in this published article and its additional files.
Ethics approval and Consent to participate
All experiments were approved by the Ethics Committee of Pingdingshan University. The data analyzed in this study all were downloaded from online database.
Patients consent for publication
Not applicable.
Supplementary data
The supplementary files are available to download from
List of abbreviations
TNBC, triple-negative breast cancer; ER, estrogen receptor; PR, progesterone receptor; HER2, human epidermal growth factor 2; OC, ovarian carcinoma; EMT, epithelial mesenchymal transition; ECM, extracellular matrix; NCK1, Noncatalytic region of tyrosine kinase 1; TCGA, The Cancer Genome Atlas; GEPIA, Gene Expression Profiling Interactive Analysis; BRCA, breast carcinoma; CSCC, cervical squamous cell carcinoma; ceRNA, competing endogenous RNAs; CC, cervical cancer;
Footnotes
Acknowledgments
Not applicable.
Conflict of interest
The authors declare that they have no competing interests.
