Abstract
BACKGROUND:
Vasculogenic mimicry (VM) is characterized by formation of three-dimensional (3D) channels-like structures by tumor cells, supplying the nutrients needed for tumor growth. VM is stimulated by hypoxic tumor microenvironment, and it has been associated with increased metastasis and clinical poor outcome in cancer patients. cAMP responsive element (CRE)-binding protein 5 (CREB5) is a hypoxia-activated transcription factor involved in tumorigenesis. However, CREB5 functions in VM and if its regulated by microRNAs remains unknown in breast cancer.
OBJECTIVE:
We aim to study the functional relationships between VM, CREB5 and microRNA-204-5p (miR-204) in breast cancer cells.
METHODS:
CREB5 expression was evaluated by mining the public databases, and using RT-qPCR and Western blot assays. CREB5 expression was silenced using short-hairpin RNAs in MDA-MB-231 and MCF-7 breast cancer cells. VM formation was analyzed using matrigel-based cultures in hypoxic conditions. MiR-204 expression was restored in cancer cells by transfection of RNA mimics. Luciferase reporter assays were performed to evaluate the binding of miR-204 to 3
RESULTS:
Our data showed that CREB5 mRNA expression was upregulated in a set of breast cancer cell lines and clinical tumors, and it was positively associated with poor prognosis in lymph nodes positive and grade 3 basal breast cancer patients. Silencing of CREB5 impaired the hypoxia-induced formation of 3D channels-like structures representative of the early stages of VM in MDA-MB-231 cells. In contrast, VM formation was not observed in MCF-7 cells. Interestingly, we found that CREB5 expression was negatively regulated by miR-204 mimics in breast cancer cells. Functional analysis confirmed that miR-204 binds to CREB5 3
CONCLUSIONS:
Our findings suggested that CREB5 could be a potential biomarker of disease progression in basal subtype of breast cancer, and that perturbations of the miR-204/CREB5 axis plays an important role in VM development in breast cancer cells.
Introduction
Vasculogenic mimicry (VM) was first described in malignant melanoma cells and involves the formation of three-dimensional (3D) channel-like networks by tumor cells [1]. These cellular networks allow the supply of blood and oxygen in which tumors can feed themselves, operating independently or simultaneously with classical endothelial vessels [2]. Remarkably, VM has been described in multiple solid tumors such as glioblastoma [3], ovarian [4], prostate [5], lung [6], hepatocellular [7] and breast cancers [8, 9]. Unfortunately, the presence of VM has been associated with aggressive tumor phenotypes, increased metastasis, poor clinical prognosis, and shorter overall survival in cancer patients. Thereby, the protein factors and modulators participating in VM are considered as potential biomarkers and therapeutic targets [10]. The tumor microenvironment is a key factor in the development of VM. Hypoxia is the most typical feature of the inner microenvironment in tumors; it is also the most important factor inducing the VM. Hypoxia through the action of the hypoxia inducible factor (HIF-1
Recent studies indicated that CREB5 expression could be regulated at posttranscriptional level by non-coding RNAs including long non-coding RNAs (lncRNAs), circular RNAs (circRNAs) and microRNAs in cancer cells [19, 20, 21]. MicroRNAs (miRNAs) are non-coding single-stranded small RNAs of 21-25 nucleotides in length that function as negative regulators of gene expression [22]. The post-transcriptional regulatory functions of miRNAs are mediated by degradation of the target mRNAs and inhibition of mRNA translation, through complementarity between the miRNA seed region and the 3
Materials and methods
Cell lines
Human MDA-MB-231 and MCF-7 breast cancer cell lines were obtained from the American Type Culture Collection, and routinely grown in Dulbecco’s modification of Eagle’s minimal medium (DMEM-F12) supplemented with 10% fetal bovine serum and penicillin-streptomycin (50 unit/ml) at 37
Analysis of CREB5 expression in public databases
The public datasets for CREB5 gene expression from breast tumors samples was obtained from Cancer Genome Atlas (TCGA) using Xena repository data (
Kaplan-Meir analysis
Kaplan-Meier curves for CREB5 expression in breast cancer patients were determined as previously described [28]. Briefly, Kaplan-Meir on line tool (
MicroRNA-204 mimics transfection
MicroRNA-204 precursor and miR-204 scramble control were both transfected at 30 nM in MDA-MB-231 and MCF-7 cells using siPORT amine transfection agent (Ambion). Briefly, pre-miR-204 was diluted in 25
CREB5 gene silencing
Two oligonucleotides pairs (21–23 nt length) corresponding to specific short hairpin RNAs (shRNA) targeting the CREB5 gene were designed. To minimize the possibility of shRNA off targeting effects, a nucleotide BLAST search was carried out. Each oligonucleotide pair was cloned into the pSilencer 5.1 U6 retro plasmid (ThermoFisher) and sequences were confirmed by automatic sequencing. The resulting recombinant plasmids were named sh-CREB5.2 and sh-CREB5.3. The efficacy of the constructs in CREB5 gene silencing was tested by transient transfection of MDA-MB-231 and MCF-7 cells (2
Vasculogenic mimicry assays
Vasculogenic mimicry experiments were performed through 3-dimensional cell cultures on matrigel. Firstly, MDA-MB-231 cells (1
Western blots
Proteins were separated on 10% polyacrylamide gels and transferred to 0.2
Immunofluorescence analysis
Briefly, cells were fixed in 4% formaldehyde in PBS 1X for 30 min at room temperature. Coverslips were incubated with 0.1% Triton X-100 for 3 min. Following washing with PBS 1X, cells were blocked for 40 min at room temperature with 0.2% BSA in PBS 1X, and incubated with anti-CREB5 antibodies overnight at 4
Luciferase reporter assay
The 3
Statistical analysis
Experiments were performed three times by triplicate and results were represented as mean
Results
Expression of CREB5 is upregulated in breast tumors
To evaluate the CREB5 expression in breast cancer patients, we first examined its mRNA levels in a cohort of 1,391 breast tumors and matched adjacent normal samples from TGCA datasets. Our results showed that CREB5 mRNA was significantly (
CREB5 is overexpressed in breast tumors and cancer cell lines. (A) CREB5 expression in 1391 breast tumors and matched adjacent normal samples from TCGA datasets (***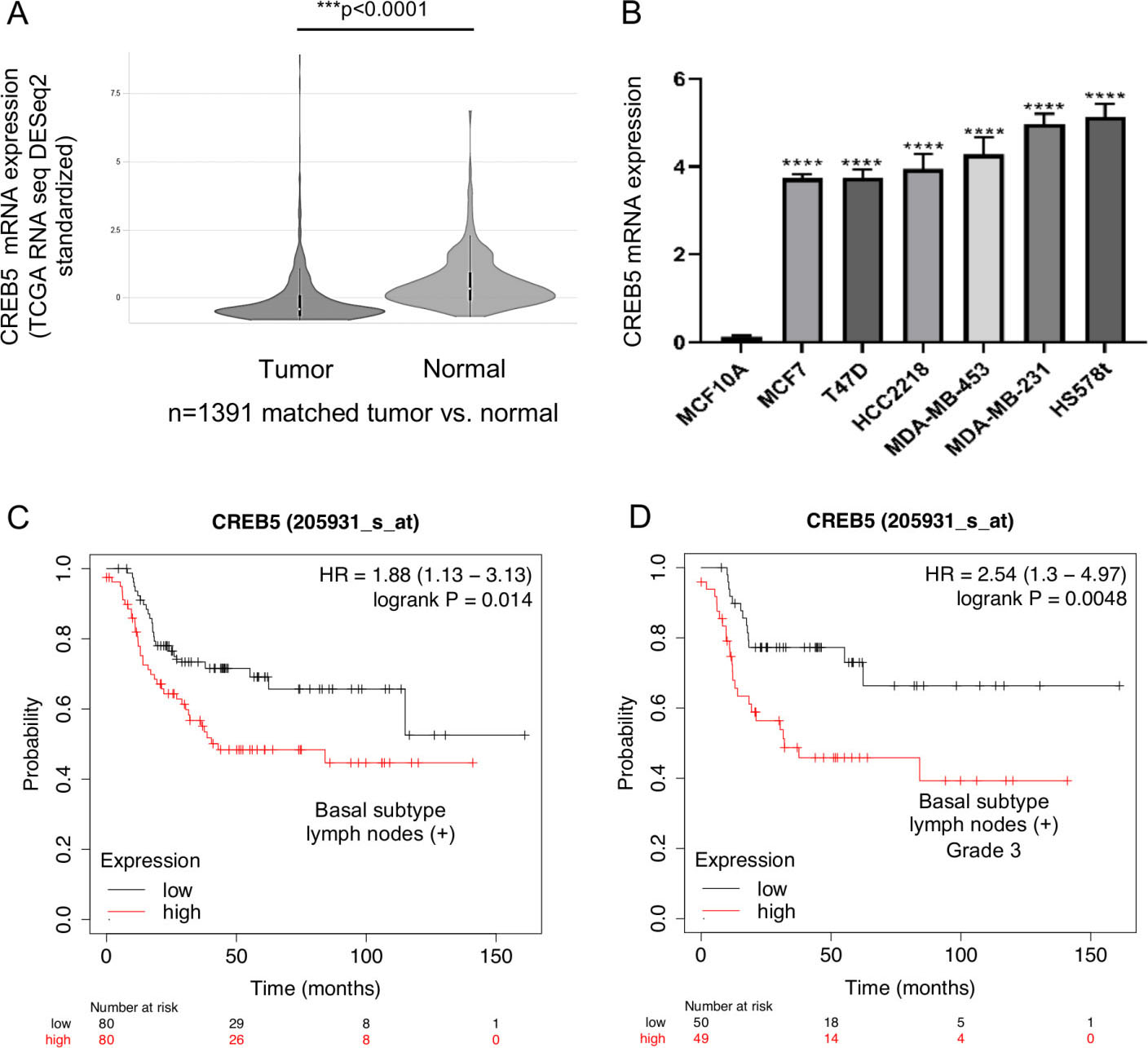
CREB5 silencing in MDA-MB-231 and MCF-7 breast cancer cell lines. (A) Western blot assays for CREB5 knockdown in MDA-MB-231 and MCF-7 cells transfected with the sh-CREB5.2 or sh-CREB5.3 constructs using CREB5 antibodies (1:1000). GAPDH was used as an internal loading control. (B) Densitometric analysis of immunodetected bands in panel A. Data represents the mean of three independent assays 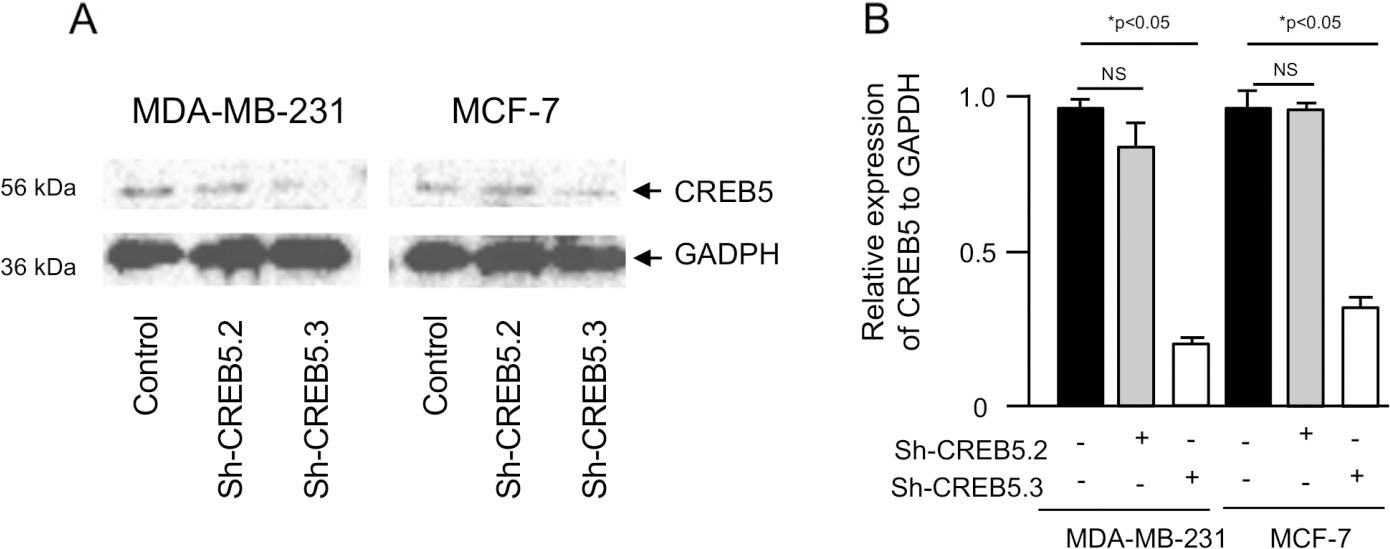
Silencing of CREB5 expression inhibits vasculogenic mimicry. (A) Images of vasculogenic mimicry of MDA-MB-231 cells transfected with sh-CREB5.3 (right panel), scramble control (middle panel) and non-transfected (left panel). Cells were submitted to hypoxia for 48 h, and then incubated in matrigel during 0–9-h. (B) Graphical representation of quantification of number of capillary tubes and (B) nodes as depicted in panel A. (D) Images of vasculogenic mimicry of MCF-7 cells after 48 h hypoxia and subsequent incubation in matrigel for 9 h. Experiments were performed three times in triplicate and the data were expressed as mean 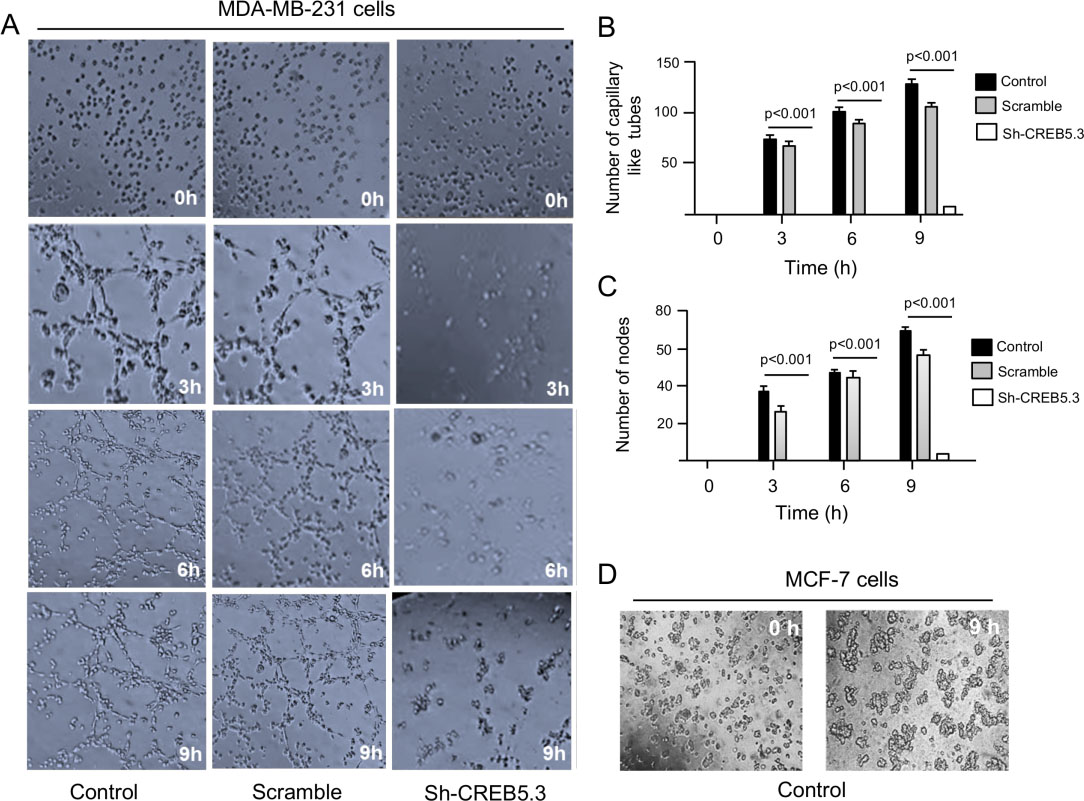
Then, we wondered whether high expression of CREB5 had clinical implications in the prognosis of breast cancer patients. Therefore, we performed an overall survival analysis using the Start Kaplan-Meier tool as implemented for breast cancer, which utilizes genome-wide transcriptome data and clinical overall survival information from breast cancer patients with a median follow-up of 150 months. Data showed that high expression of CREB5 transcript was associated with poor overall survival of breast cancer patients with basal subtype and lymph node-positive tumors (HR
MicroRNA-204 targets the CREB5 gene. (A) Western blot of MDA-MB-231 and MCF-7 cells, non-transfected and transfected with pre-miR-204, using the CREB5 polyclonal antibodies (1:1000). GAPDH specific antibodies were used as control. (B) Densitometric analysis of bands in panel A. Data were normalized using GAPDH expression. Images are representative of three independent experiments. (C) Immunofluorescence assays using anti-CREB5 antibodies for CREB5 protein detection and DAPI for nuclear DNA staining (blue channel) in MDA-MB-231 (Alexa-555, red channel) and MCF-7 (Alexa-488, green channel) cells transfected with miR-204 and scramble as control. (D–E) Densitometric analysis of immunofluorescence signals in panel C. (F) Schematic representation of p-miR-Report construct containing the miR-204 binding sites of the 3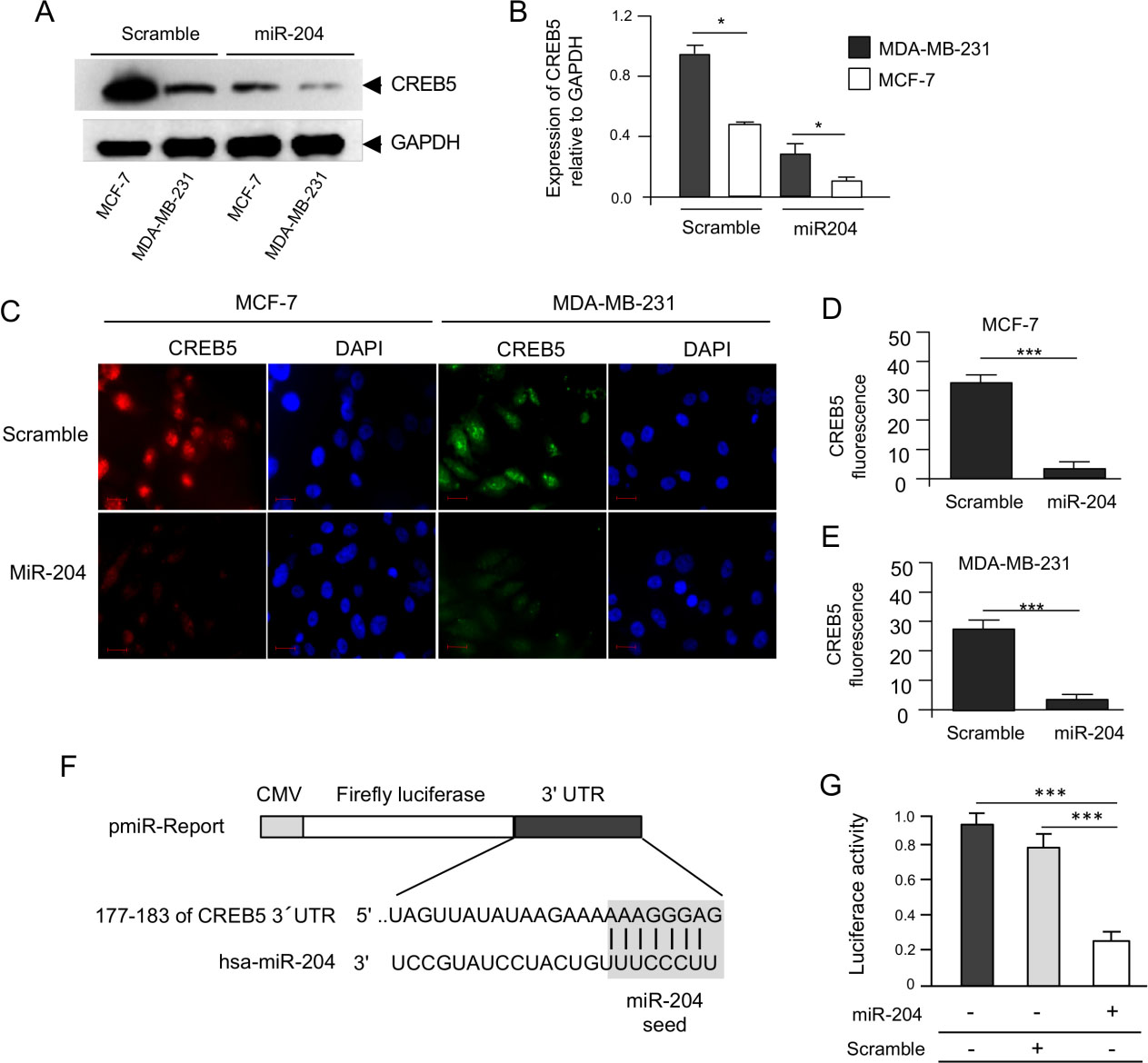
CREB5 is a hypoxia-activated gene but its role in VM is unknown. To determine if selective inhibition of CREB5 affects 3D channels-like formation representative of VM, its expression was knocked-down by RNA interference. Two specific short hairpin RNAs (designated as shCREB5.2 and shCREB5.3) targeting the CREB5 mRNA were designed and cloned into the pSilencer vector. Both constructs were individually transfected in triple negative MDA-MB-231 and luminal MCF-7 breast cancer cell lines, and subsequently CREB5 protein expression was analyzed by Western blot assays using specific antibodies 48 h post transfection. The results showed that the shCREB5.3 sequence efficiently down-regulated CREB5 expression in both cell lines (Fig. 2A), whereas no significant effect was observed with the shCREB5.2 interfering sequence. GADPH protein expression used as a loading control, showed no significant changes between both treatments. Densitometric analysis of the immunodetected bands showed that silencing induced by the shCREB5.3 construct was effective, as this sequence suppressed CREB5 expression about 85% in MDA-MB-231 cells and by 62% in MCF-7 cell line (Fig. 2B). Then, the effects of CREB5 silencing using shCREB5.3 were evaluated by hypoxia-induced 3D channels-like network formation assays representative of the early stages of VM in MDA-MB-231 cells. Results showed that, after 48 h hypoxia, VM was effectively induced in control cells showing an increased number of branch points and capillary tubes during the course of time (0–9 h) reaching the highest levels at 9 h (Fig. 3A, left panel). Interestingly, CREB5 silencing using the sh-CREB5.3 resulted in a dramatic inhibition of VM under hypoxia at 3–9 h (Fig. 3A, right panel). Likewise, a reduction in the number of branch points and capillary tubes compared to untreated control cells was found since the initial time points until 9 h of channels-like formation assays (Fig. 3A, right panel). No significant inhibition of VM was observed in scramble-transfected control cells, as expected (Fig. 3A, middle panel). Intriguingly, VM was not detected in poorly invasive MCF-7 breast cancer cells, suggesting that this process could be a specific feature of highly metastatic tumors such as MDA-MB-231 cancer cells (Fig. 3D).
MicroRNA-204 targets the transcription factor CREB5
In previous studies, we have shown that miR-204 inhibits the hypoxia-induced channels-like formation [4]; therefore, it was reliable to propose that it may regulates VM at least in part by targeting CREB5 as the bioinformatics analysis showed that it contains potential 3
Discussion
In spite of recent advances in chemotherapeutic treatment of locally advanced and metastatic disease, breast cancer still remains as a lethal disease. The tumor microenvironment is a key factor in the development of aggressive disease, as it activates diverse cell signaling pathways impacting the cancer hallmarks including angiogenesis, therapy resistance, invasion, metastasis and VM. Notably, VM has been associated with more aggressive tumors, poor prognosis, and shorter overall survival in breast cancer patients. Thereby, there is an urgent need for potential biomarkers of disease progression and novel therapeutic agents targeting VM [10]. Hypoxia activates the hypoxia inducible factor (HIF-1
cAMP response element-binding protein (CREB) is family of proteins that promote tumorigenesis in different types of cancer. CREB proteins have the ability to activate the expression of diverse target genes which are involved in important processes in tumor development including metabolism, angiogenesis, cell cycle, cell survival, proliferation, and DNA repair [29]. Most of our limited knowledge about the functions of CREB5 in tumors comes from colorectal cancer studies. Previous studies have shown that CREB5 is overexpressed in colorectal cancer, and showed that it may plays a key role in cell proliferation and metastasis by regulation of c-Jun, CSF1R, MMP9, PDGFRB, FIGF and IL6 [30]. Also, it was showed that CREB5 expression was upregulated in colorectal cancer and positively correlated with advanced WHO stages and TNM stages and shorter survival in patients. CREB5 silencing reduced the invasiveness and metastatic capacity of colorectal cancer cells
As CREB5 seems to be a novel oncogene, here we focused in the study of its functions in vasculogenic mimicry in breast cancer. Silencing of CREB5 abolished the hypoxia-induced formation of 3D channels-like structures representative of the early stages of vasculogenic mimicry in breast cancer cells. These findings are novel as no previous role for CREB proteins have been reported, and added complexity to the molecular mechanisms operating at transcriptional level through the action of transcription factors in VM. To obtain clues on the mechanisms of CREB5 regulation we evaluated if its expression could be modulated by miR-204 in breast cancer cells, as they have been reported as major regulators of genes involved in angiogenesis and vasculogenic mimicry. CREB5 contains 4 predicted binding sites for miR-204 suggesting a posttranscriptional regulation by miR-204, a non-coding RNA that has been previously reported to be downregulated in several cancer cell lines and solid tumors, including breast cancer. As a negative regulator, miR-204 may inhibits the angiogenic by directly targeting key factors involved in, such as ANGPT1 and TGFBRII [23]. Interestingly, here we found that CREB5 expression was negatively regulated after restoration of miR-204 levels in breast cancer cells, suggesting that posttranscriptional regulation of CREB5 could represent an important control point of gene expression.
All together these finding raises several questions which remain to be solved in future studies: i) What is the transcriptional program activated by CRE5 transcriptional factor involved in vasculogenic mimicry activation? ii) Are there additional microRNAs potentially regulating CREB5 expression and 3D channel-like formation?, and iii) To what extend do CREB5-miR-204 interactions may occur in other types of human cancers?. In conclusion, we showed that tumor suppressor miR-204 exhibit a potential role in the regulation of vasculogenic mimicry through the miR-204/CREB5 axis in breast cancer.
Footnotes
Acknowledgments
We acknowledge to Universidad Autónoma de la Ciudad de México for support. This research was partially funded by Consejo Nacional de Ciencia y Tecnologia, grant number A1-S-13656.
Conflict of interest
The authors declare that they have no competing interests.
Author contributions
Conception: C.L.C. L.A.M and C.P.P.
Interpretation or analysis of data: E.C.S., C.P.F., A.F.P., Y.M.S.V., M.B.S.C., O.N.H.C., R.G.A.B.
Preparation of the manuscript: C.L.C and M.E.A.S. Revision for important intellectual content: Supervision: C.L.C., L.A.M., and M.E.A.S.
All authors have read and agreed to the published version of the manuscript.
