Abstract
BACKGROUND:
Numerous studies reveal the clinical significance of tumor microenvironment (TME) in multiple cancers. The association between TME in oral squamous cell carcinoma (OSCC) and clinical outcomes remains unsolved.
OBJECTIVE:
This study aims to exhibit the TME of OSCC and identified the prognostic marker.
METHODS:
Gene expression profile and clinical data OSCC patients were from the TCGA database. The validated stage data was from the Gene Expression Omnibus (GSE65858). Immune/stromal scores of each patient were calculated by ESTIMATE algorithm. Biological functional prediction was conducted. Prognostic genes identified by survival analysis. Nomogram and Receiver operating characteristic curves were employed to test the predicting power. TIMER database was applied to evaluated the immune infiltrates.
RESULTS:
Lower immune scores were observed in male patients (
CONCLUSION
: The current study analyzed the TME and presented immune related prognostic biomarkers for OSCC.
Introduction
Oral squamous cell carcinoma (OSCC) is one of the most common types of cancers in the world. The risk factors including tobacco and alcohol consumption or betel nut chewing contribute to the high incidence of OSCC [1]. Despite the advances in OSCC patients’ treatment, such as surgery, chemotherapy, radiotherapy or biologic therapy, the overall survival rate for OSCC remains poor in the recent decades [2]. In addition, OSCC patients have a poor quality of life due to the dysfunction in chewing and swallowing [3].
Tumor microenvironment (TME) refers to the cellular environment in which tumors or cancer stem cells exist. Apart from the tumor cells, the TME includes surrounding blood vessels, other non-malignant cells, the extracellular matrix (ECM) and signaling molecules. TME is a complex integrated system, which can be divided into immune microenvironment dominated by immune cells and non-immune remission dominated by fibroblasts [4]. It has been reported that TME is associated with tumor survival and recurrence [5], such as glioblastoma [6], colon cancer [7] and prostate cancer [8] by estimation of stromal and immune cells in malignant tumor tissues using expression data (ESTIMATE) algorithm [9], which was applied for calculating the immune and stromal cells in tissues. However, up to now, no studies have been conducted on the association between tumor microenvironment and prognosis in oral squamous cell carcinoma.
With the development of data sharing, transcriptome profiles and clinical data of cancer patients can be easily obtained from some public database such as The Cancer Genome Atlas (TCGA) database. TCGA were launched in 2005 using genome analysis technologies to understand the genetics of cancers, which helping to improve the diagnosis, treatment and prevention of cancers [10]. Here, we conducted the current study to identify prognostic genes which were associated with OSCC. A total of 331 OSCC patients with gene expression profiles and clinical data were collected to evaluate the TME by ESTIMATE, then patients were divided into two groups based on the median of immune scores as well as stromal scores to access the association of survival and differentially expressed genes (DEGs). Kyoto Encyclopedia of Genes and Genomes (KEGG) and Gene Ontology (GO) pathway enrichment analysis were applied to investigate the biological functions of identified genes. Survival analysis was used to explore the prognostic genes and prognostic genes were validated in Gene Expression Omnibus (GEO) database. In addition, receiver operating characteristic curve (ROC) and nomogram were applied to access the prognostic value of the identified key gene and clinical characteristics. Furthermore, the TIMER database was applied to evaluate the association between the key gene HGF (hepatocyte growth factor) and immune infiltration, the whole work flow was exhibited in Supplementary Fig. 1.
We present the following article in accordance with the MDAR checklist.
Methods and materials
Data collection
Level 3 gene transcriptome profiles for 331 OSCC patients were obtained from The Cancer Genome Atlas (TCGA) database (
Screening for differentially expressed genes (DEGs)
In groups divided into by immune scores and stromal scores, the transcriptome data were transformed by log2 (x
(A) Immune and stromal scores were associated with the clinical characteristics of OSCC patients. Scatter plots revealed the distribution of immune scores in sex (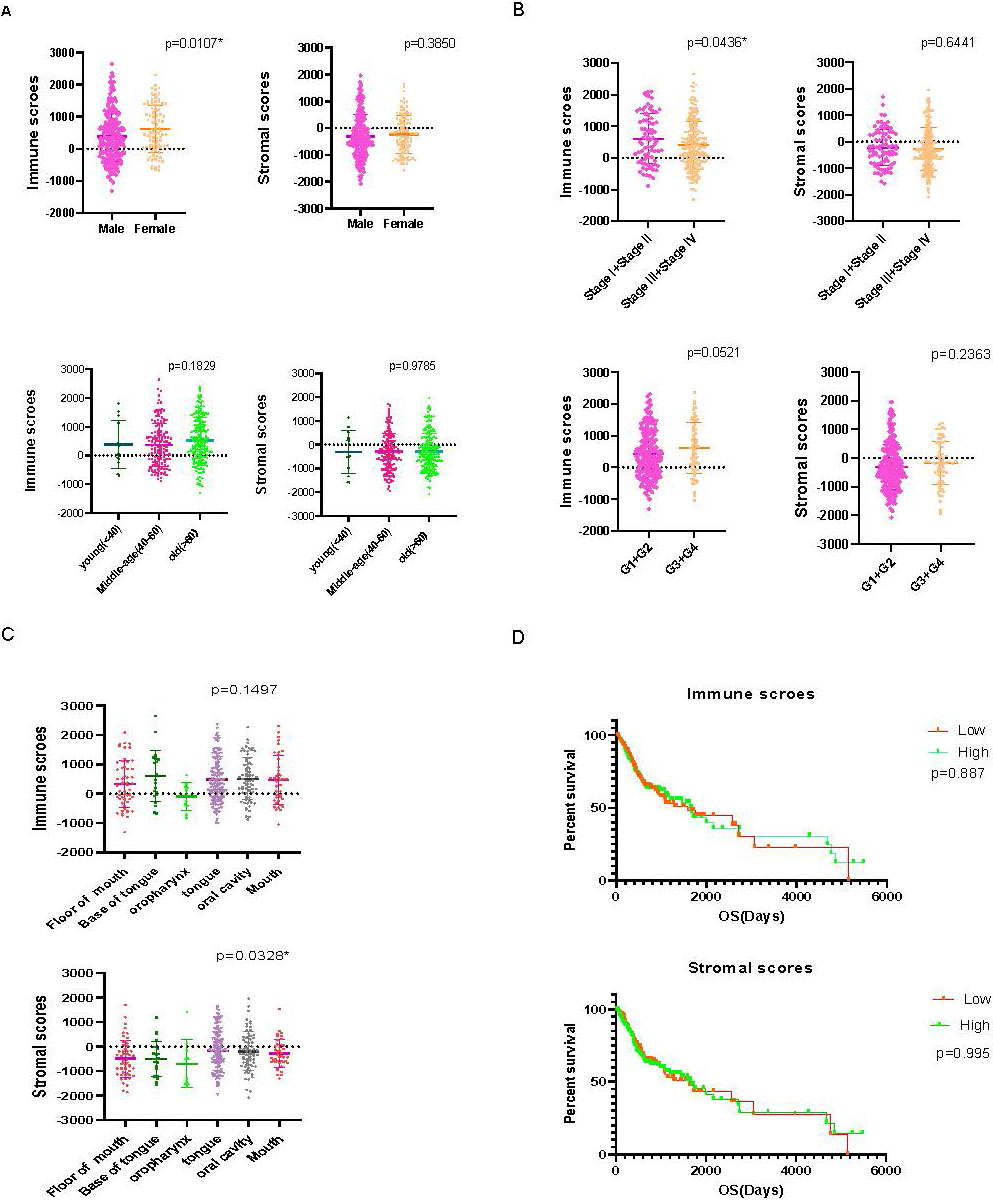
(A) Differentially expressed genes between the high immune scores-group and the low immune scores-group and (B) differentially expressed genes between the high stromal scores- group and the low stromal scores-group were exhibited in volcano plot. (C) Venn diagram showed the intersection of differentially expressed genes which were upregulated, (D) and which were downregulated.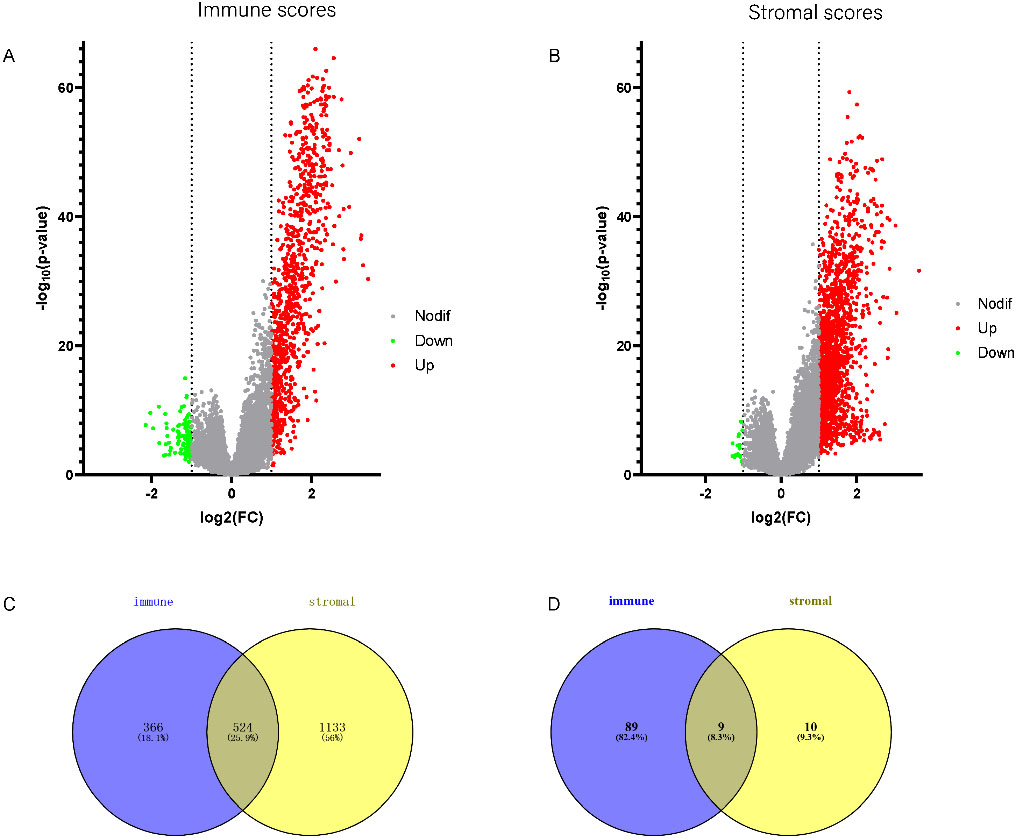
KEGG and GO analysis. (A) Pathway enrichment in KEGG analysis, (B) Results of CC, (C) MF and (D) BP in GO analysis revealed the relationship between differentially expressed genes and functional pathways.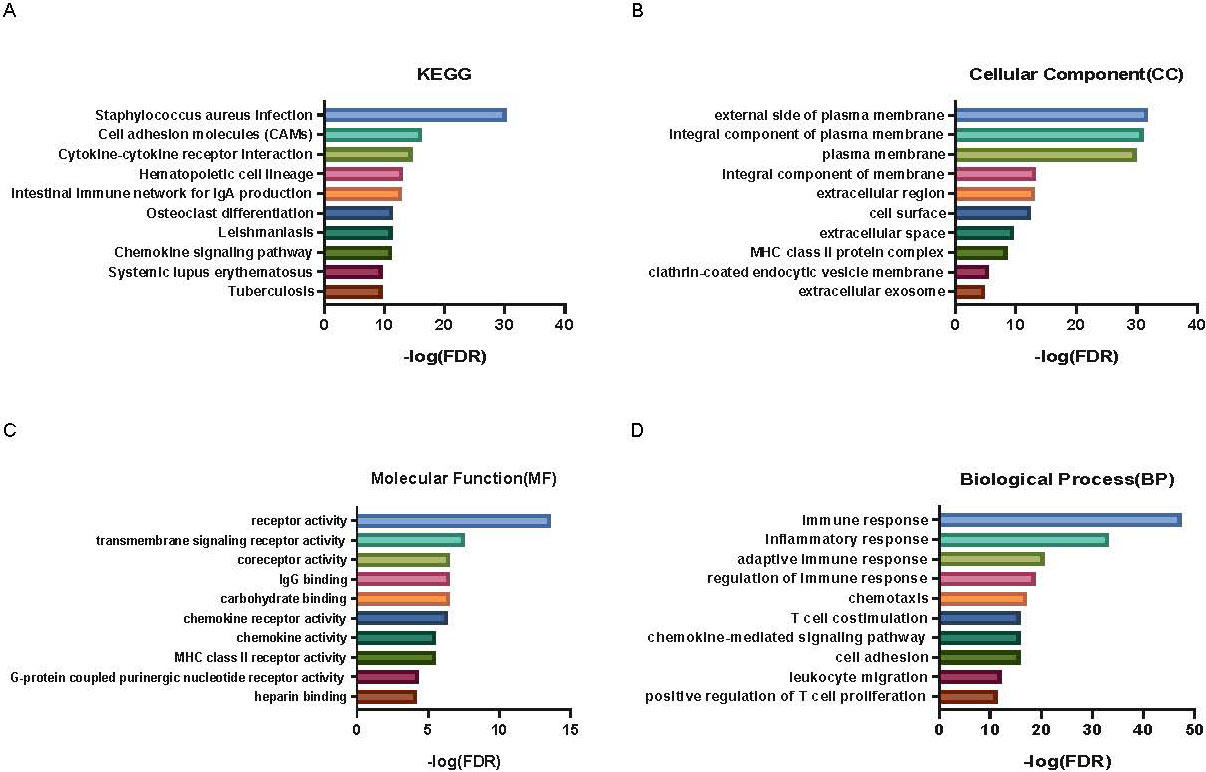
Online tool DAVID (
Nomogram and ROC curves
Cox proportional hazards regression models were used for univariate analyses and to build a nomogram. The predictors analyzed included age, gender, TMN stage, tumor grade and the expression value of HGF. The predicted 1- and 5-year survival probabilities were calculated for patients with the nomogram. Receiver operating characteristic (ROC) curves were drawn to compare single gene (HGF) model with gene (HGF) combined with clinical characteristics model, and the areas under the curve (AUCs) at the 1- and 5-year survival probabilities were calculated to assess the discriminative power of the model. The risk scores (RS) were calculated for ROC curves by considering the expression value of HGF, age, gender, TMN stage and tumor grade with their corresponding coefficient in the formular of risk score
Timer database analysis
The deconvolution algorithm provided by the TIMER (tumor immune estimation resource) database (
Statistical analysis
The R software (version 4.0.5) was used to execute all the statistical analyses and
Results
Immune scores and stromal scores are associated with OSCC subtypes
We divided stage into early stage (stage I and II) and advanced stage (stage III and IV),the immune scores represented the distribution of immune cells in the immune microenvironment, we found that the lower immune scores were observed in male patients (
(A) 72 genes predicting the significant poor overall survival selected by univariate cox proportional hazards regression analyses (TCGA cohort). (B) Survival analysis of hub genes. The K-M curves revealed that the top 9 hub genes and HGF were significantly associated with survival outcomes (TCGA cohort). (C) HGF was proved to be associated with poor prognosis in GEO cohort.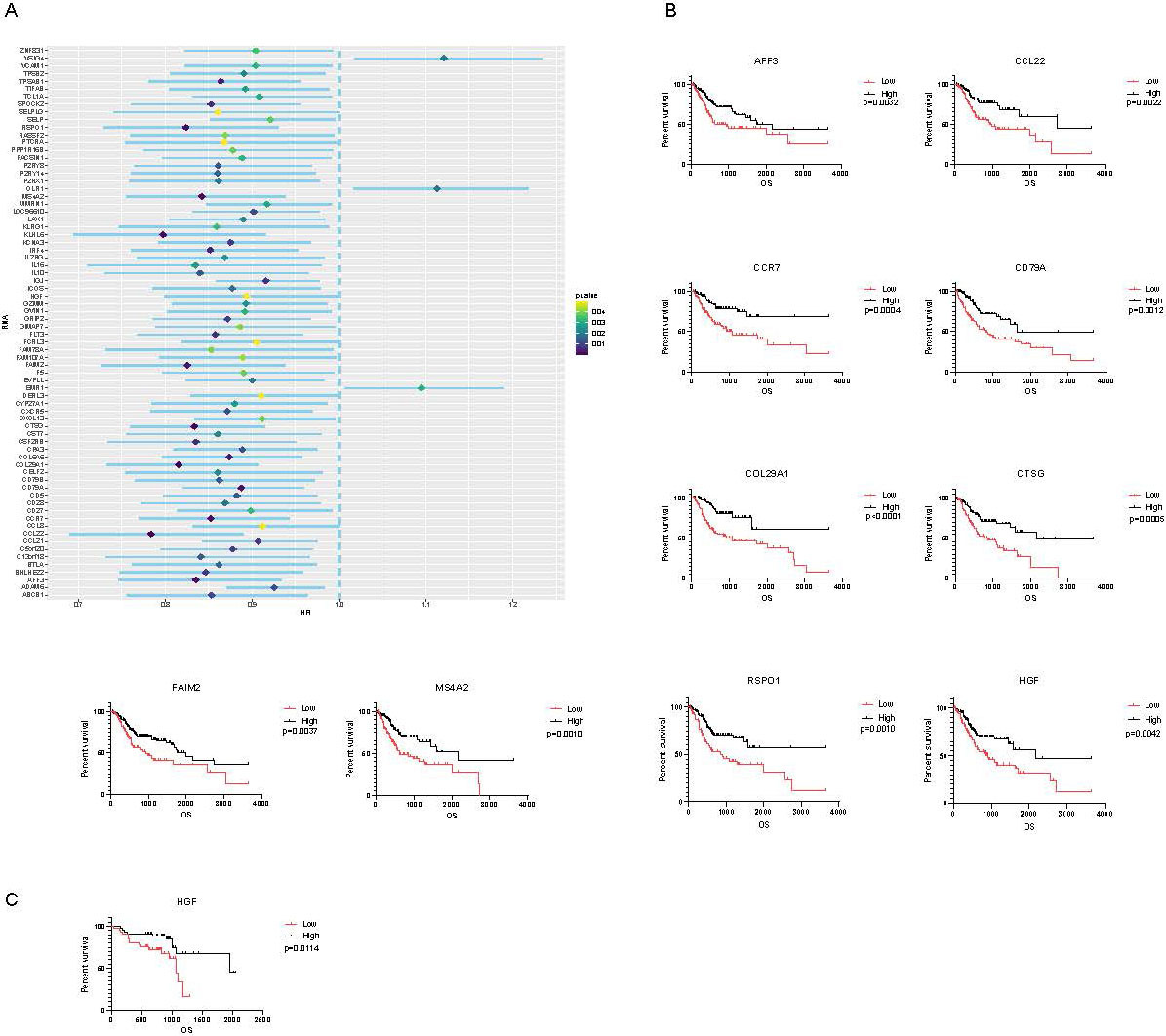
(A) prognostic nomogram for OS prediction. (B) ROC curve (single gene-based model and gene combined with clinical characteristics model) indicating 1-year and 5-year overall survival in the TCGA cohort. (C) ROC curve (single gene-based model and gene combined with clinical characteristics model) indicating 1-year and 5-year overall survival in the GEO cohort. ROC, receiver operator characteristic; AUC, area under the curve; OS, overall survival.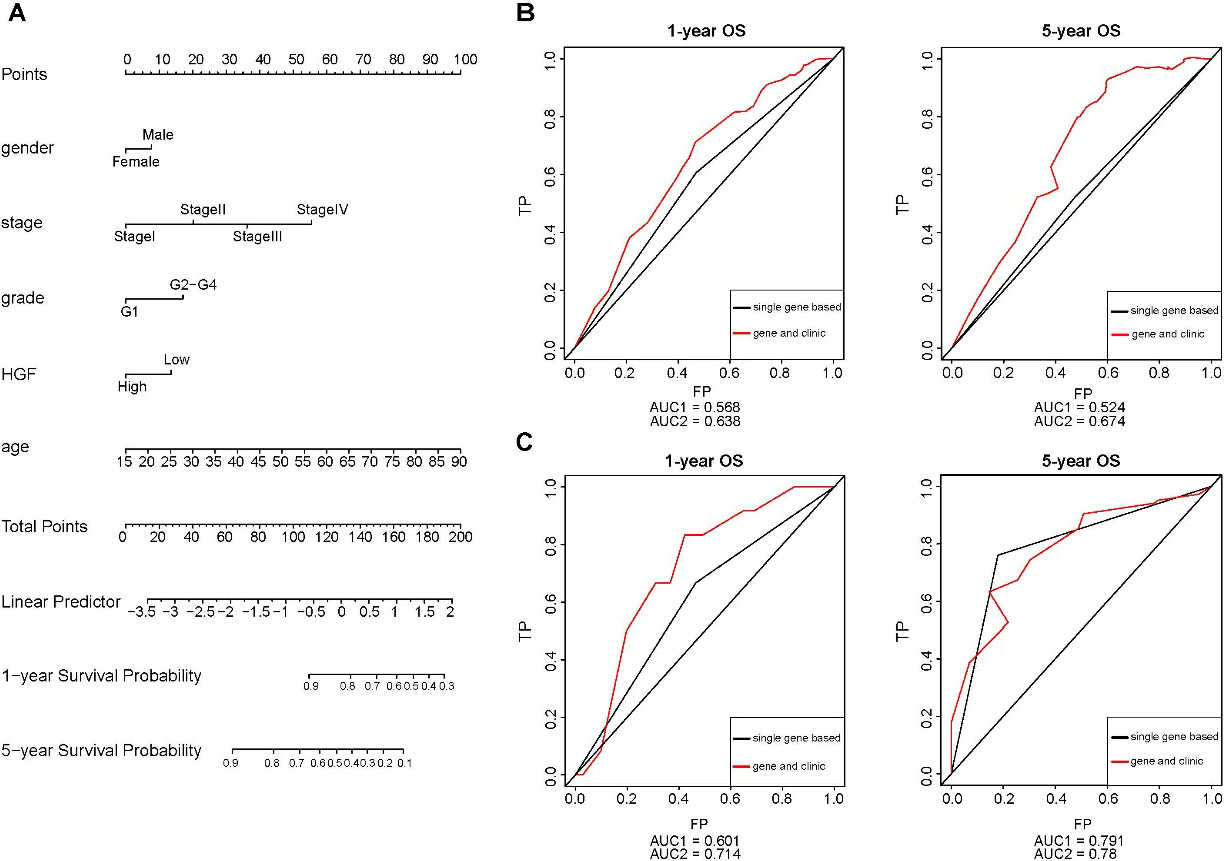
(A–C) HGF was associated with immune cells infiltration. Immune cells included B cell, CD4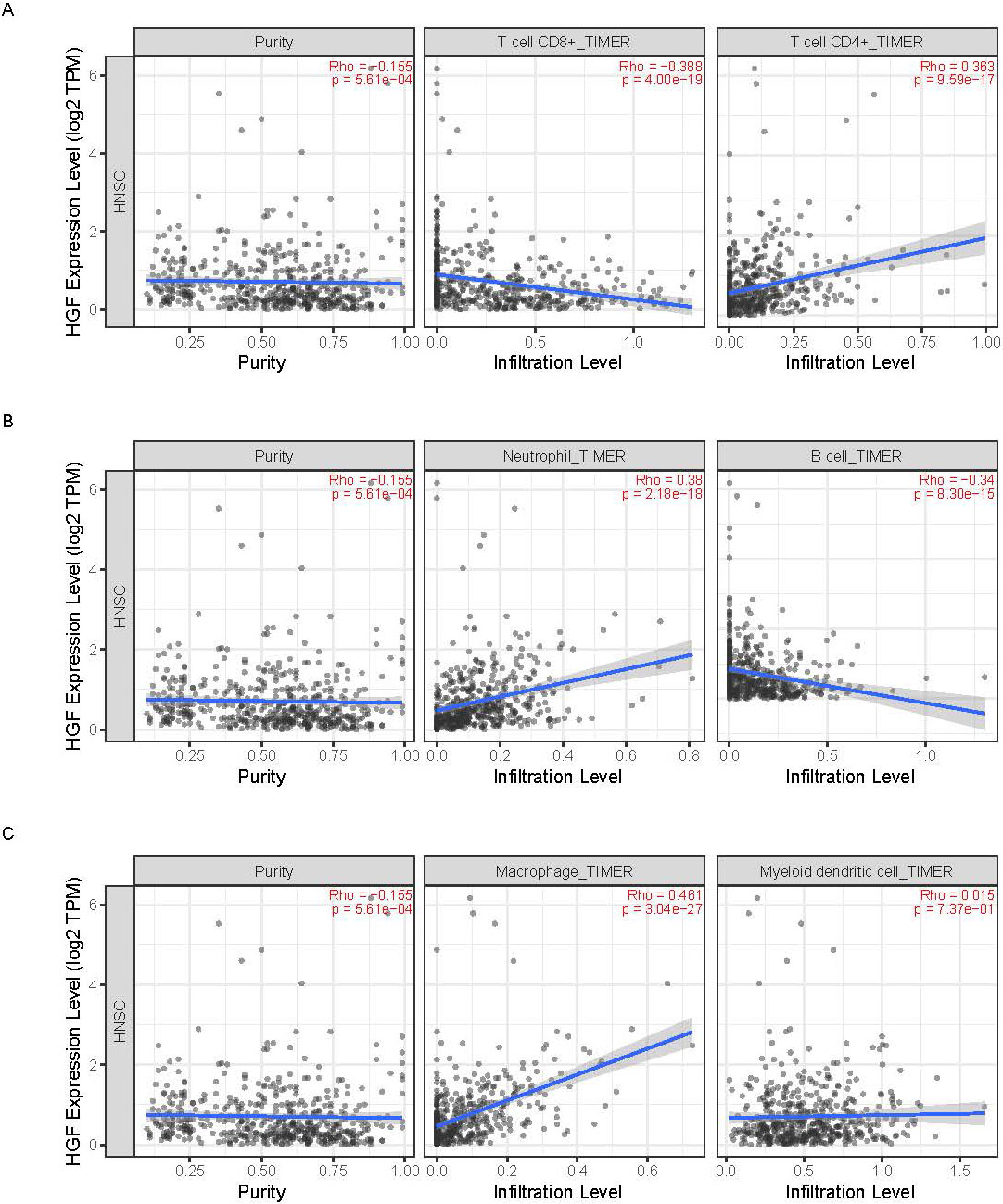
OSCC patients were divided into low-level group (
GO and KEGG analysis of DEGs
The KEGG pathway analysis of the 533 DEGs showed on Fig. 3A, cytokine-cytokine receptor interaction in the top 10 pathways. Gene ontology (GO) terms including cellular component, molecular function and biological process and pathway analyses were listed in Fig. 3B to D. The GO analysis revealed that DEGs might be linked to positive regulation of T cell proliferation, T-cell co-stimulation, MHC class II protein complex and MHC class II receptor activity.
Identification of prognostic gene based on TCGA database and validated on GEO database
A total of 72 DEGs were associated with overall survival which were identified by univariate cox regression analyses and the results were listed in Fig. 4A and Supplementary Table 5. We exhibited top 9 significant DEGs in log-rank test (AFF3, CCL22, CCR7, CD79A, COL29A1, CTSG, FAIM2, MS4A2, RSPO1) and HGF (Fig. 4B). HGF which was associated with overall survival in GEO database as well (
Predicting prognosis of OSCC by nomogram and ROC curve
We constructed a nomogram including age, gender, TMN stage, tumor grade and the expression value of HGF to predict OS at 1 and 5 years after diagnosis in the TCGA cohort (Fig. 5A). The area under the curve (AUC) of clinical characteristics combined HGF was 0.638 and 0.674 corresponding to 1-year and 5-year OS (Fig. 5B) in the TCGA cohort, basing on the risk score (RS) calculated as: RS
Key gene were associated with immune cell infiltration
We explored the relationship between HGF and six kinds of immune cells in OSCC patients by TIMER database. The results illuminated that HGF was significantly related with B cell, CD4
Discussion
The current study by integrating the public databases TCGA and GEO to evaluated the TME of OSCC, a total of 533 DEGs were identified in immune cells and stromal cells. Further functional enrichment analysis showed that those DEGs were involved in immune related pathways including the positive regulation of T cell proliferation pathway, the T-cell co-stimulation pathway, the MHC class II protein complex pathway and the MHC class II receptor activity related pathway. HGF was identified as a candidate gene closely related to the overall survival of OSCC by survival related analysis and the prognostic value of HGF was accessed by nomogram and ROC analysis. The TIMER database analyses also provide evidence for the strong correlation between HGF and immune cells infiltration, indicating the significant prognostic value of HGF in OSCC.
The TME is the cellular environment around tumor cells including endothelial cells, lymphocytes, macrophages, NK cells, other cells of the immune system, fibroblasts, mesenchymal stem cells (MSCs), and the extracellular matrix (ECM), contributing to the development, growth, and metastatic spread of malignancies [11]. The immune cell infiltrates in the environment are associated with better outcomes in early-stage cancer patients [12, 13]. The stromal cells build the growth and survival capacity of tumor cells during tumor initiation and progression by producing growth factors, chemokines and cytokines [14].
Hepatocyte growth factor (HGF), also called scatter factor, is a heterodimer protein that is normally synthesized by mesenchymal cells [15]. Combined with its sole physiological ligand: MET(c-MET, mesenchymal-epithelial transition factor) [16], HGF contributed to both the early and later stages of the cancer development by directly or indirectly stimulating proliferation, invasion, metastasis processes and angiogenesis [17, 18]. Studies have demonstrated that overexpression of HGF and/or MET is characteristic of several epithelial cell derived cancers and for some is an independent prognostic factor associated with aggressive nature of the tumor type and poor clinical outcome [19]. Wislez et al. reported that neutrophils may be attracted to the microenvironment generated by tumors such as bronchioloalveolar carcinoma, releasing bioactive HGF and promoting the invasion of cancer cells [20].
The four functional pathways including: positive regulation of T cell proliferation, T-cell co-stimulation, MHC class II protein complex and MHC class II receptor activity, which are important components of immune response to cancer [21]. A number of studies have shown that CD27/CD70, the members of the tumor necrosis factor receptor (TNFR) family, can enhance tumor or graft rejection, with co-stimulatory effects on T cells enhancing both NK-dependent and T cell-dependent mechanisms of tumor elimination [22]. The MHC class II protein complex transfer antigens derived by cancer cells to tumor-infiltrating CD4 T cells, stimulating direct cytotoxicity toward tumor cells [23]. The positive regulation of T cell proliferation in tumors are related to improved prognosis [24], for the cytotoxic activity of CD8
Finally, there are still some limitations existing in our study. Firstly, the lack of the clinical samples may affect the validation of the results. Secondly, less than 1/3 patients provided information on treatment, the effect of treatment on prognosis could not be evaluated. Moreover, further functional experiments of HGF should be conducted to understand the potential mechanism in the process of the carcinogenesis of OSCC.
Author contribution
Interpretation or analysis of data: Longbiao Zhu and Xinyu Zhang.
Preparation of the manuscript: Donglin Yan and Yan Chen.
Revison for important intellectual content: Donglin Yan and Xinyu Zhang.
Supervision: Longbiao Zhu and Jing Han.
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-210432.
sj-docx-1-cbm-10.3233_CBM-210432.docx - Supplemental material
Supplemental material, sj-docx-1-cbm-10.3233_CBM-210432.docx
Footnotes
Acknowledgments
This work was supported by National Natural Science Foundation of China (81602920, 81702686), The young talents program of Jiangsu Cancer Hospital.
