Abstract
MicroRNAs are endogenous small noncoding RNAs that negatively regulate gene expression at posttranscriptional level. The discovery of microRNAs has identified a new layer of gene regulation mechanisms, which play a pivotal role in development as well as in various cellular processes, such as proliferation, differentiation, cell growth, and cell death. Deregulated microRNA expression favors acquisition of cancer hallmark traits as well as transforms the tumor microenvironment, leading to tumor development and progression. Many recent studies have revealed altered expression of microRNAs in oral carcinoma with several microRNAs shown to have key biological role in tumorigenesis functioning either as tumor suppressors or as tumor promoters. MicroRNA expression levels correlate with clinicopathological variables and have a diagnostic and prognostic value in oral carcinoma. For these reasons, microRNA has been a hot topic in oral cancer research for the last few years. In this review, we attempt to summarize the present understanding of microRNA deregulation in oral carcinoma, their role in acquiring cancer hallmarks, and their potential diagnostic and prognostic value for oral cancer management.
Introduction
Oral squamous cell carcinoma (OSCC), the most common malignancy in the oral and maxillofacial regions, accounts for over 95% of all head and neck cancers. 1 Oral cancer has an annual global incidence and mortality rates of approximately 5.9 and 3.3 per 100,000 persons, respectively.2,3 The Indian sub-continent remains the global epicenter of oral cancer, with approximately 70,000 cases and more than 48,000 oral cancer–related deaths occurring annually. 4 In India, oral cancer is the most common cancer among men and ranks third among women. 5 Tobacco use, alcohol consumption, and betel quid chewing are the key established risk factors for oral cancer development with the use of smokeless tobacco particularly important in the Indian population.6,7
Despite advances in therapy and increasing knowledge about the development and the molecular mechanisms behind oral cancer, the 5-year overall survival of patients is still only around 50%. 8 Even though oral cavity is easily accessible for direct visual examination, most OSCCs are not diagnosed until an advanced stage. Early stage oral carcinoma is easily curable through surgery, radiation therapy, and chemoradiation; whereas patients with advanced oral cancer at late stages often present high mortality rates with little chance for receiving effective treatment. 9 Hence, it is essential to discover new therapeutic targets that are imperative for oral cancer development and progression.
MicroRNAs (miRNA) have started a revolution in molecular biology and emerged as key players in carcinogenesis. These conserved posttranscriptional regulators of gene expression are integral to almost all known biological processes, including cell growth, proliferation and differentiation, and metabolism and development of organism. 10 Over the past decade, many studies have established the importance of miRNAs in cancer biology by controlling expression of their target messenger RNAs (mRNAs) to facilitate tumor growth, invasion, angiogenesis, and immune evasion.11,12 Moreover, tumor miRNA profiles can define relevant subtypes, patient survival, and treatment response.13–15 Cancer-associated miRNA biomarkers can be detected in biological fluids, allowing less invasive monitoring. 16 For the last few years, miRNA has hence been a hot topic in cancer research owing to its potential value as diagnostic and prognostic marker for cancer management.
In this review, we summarize the present understanding of miRNA deregulation and its underlying molecular mechanisms in oral cancer. We outline how the miRNA alterations correlate with cancer hallmarks as well as how miRNAs transform the tumor milieu of oral cancer. The potential role of miRNAs as diagnostic tools and predictive biomarkers of oral cancer is also discussed.
MiRNAs—biogenesis and function
MiRNAs are an evolutionarily conserved group of endogenous, small, noncoding single-stranded RNAs of 18 to 24 nucleotides that negatively regulate gene expression at posttranscriptional level within diverse cell types. 17 They were first identified in 1993 in Caenorhabditis elegans and revealed to be involved in regulating the timing of larval development. 18 MiRNAs are encoded in the genome in various contexts: They can be expressed from intronic or intergenic transcripts, which may encode a single miRNA hairpin precursor, or clusters of multiple precursors. The biogenesis of miRNAs generally occurs by a well-characterized conserved processing mechanism (Figure 1). Initially, miRNAs are transcribed from miRNA coding sequences (residing principally in intergenic regions or within introns of genes) as a form of long primary transcript (pri-miRNAs) by the RNA polymerase II (Pol II) or Pol III enzyme.19,20 The pri-miRNA is spliced and usually capped with a 5′ 7-methylguanosine cap (m7G) and polyadenylated at the 3′ end, similar to protein-coding mRNAs. The long pri-miRNA transcript is processed by the microprocessor complex (500–650 kDa), comprising a nuclear protein DiGeorge Syndrome Critical Region 8 (DGCR8) and a nuclear RNase, Drosha, to generate an intermediate product, pre-miRNA with an imperfect stem loop hairpin secondary structure. 21 With the help of Exportin-5, the 60–70 nucleotide length pre-miRNAs are exported from nucleus to cytoplasm 22 where they are trimmed into short double-stranded miRNAs (22–25 nucleotide) by the cytoplasmic RNase III, Dicer. 23 A helicase then unwinds this duplex to form the mature miRNA which is then asymmetrically incorporated along with Argonaute proteins into the RNA-induced silencing complex (RISC) and exert their function by binding to the 3′-untranslated regions (3′-UTRs) of their target genes. Each step of the process is highly regulated, and deregulation at any level can result in inappropriate miRNA functions.

Schematic representation of miRNA biogenesis and function. MiRNAs are transcribed from DNA into primary miRNA (pri-miRNA) by RNA polymerase II. The long pri-miRNA is processed by Drosha and DiGeorge Syndrome Critical Region 8 (DGCR8) to generate pre-miRNA which is exported from nucleus to cytoplasm where they are trimmed into short double-stranded miRNAs by Dicer. A single strand of mature miRNA is incorporated along with Argonaute proteins into the RNA-induced silencing complex (RISC) which binds to the target mRNA leading to either mRNA degradation or translational repression. Deregulated miRNA expression causes oncogene activation and/or tumor suppressor gene suppression leading to carcinogenesis.
MiRNAs negatively regulate gene expression by binding to the 3′-UTRs of target mRNAs thereby degrading or destabilizing the mRNA or by inhibiting protein translation;24,25 some reports demonstrate that they may also function as positive regulators.26,27 MiRNA-based regulation is dependent on expression of its mRNA targets, which are not always ubiquitously expressed, and miRNA can have effects specific to cells types and conditions. The posttranscriptional silencing of target genes by miRNAs depends upon the degree of complementarity between miRNA and its target mRNAs. MiRNAs that bind to mRNA sequences with perfect or nearly perfect complementarity induce the RNA-mediated interference (RNAi) pathway, in which mRNA transcripts are cleaved by a miRNA-associated RISC (miRISC). Most animal miRNAs are thought to act by binding to imperfectly complementary sites within the 3′-UTRs of target mRNAs, thus inhibiting the initiation of translation via the miRISC. The imperfect match between miRNAs and their targets open up the possibility for miRNAs to regulate multiple genes. 24
The discovery of miRNAs has identified a new layer of gene regulation mechanisms, which play a pivotal role in almost all known biological processes, such as metabolism, development, proliferation, differentiation, cell growth, and cell death.10,28 Computational predictions and genome-wide identification of miRNA targets estimate that each miRNA can bind to hundreds of different mRNAs, which collectively results in the regulation of more than half of the protein-coding genes in humans. 29 Reflecting their importance in cell biology, it is not surprising that deregulation of miRNA expression is linked to various human diseases, including cancer. 30
MiRNAs altered in OSCC
The expression profiles of miRNAs appear to be tumor and tissue specific. Numerous studies have identified aberrant miRNA expression profiles in oral cancer tissues and/or cell lines relative to the corresponding normal controls.31–33 Several miRNAs have key biological role in the initiation and progression of oral carcinoma functioning either as tumor suppressors or as tumor promoters. 34 MiRNA expression profiles correlate with and could explain the pathogenesis, 35 metastasis,36,37 and chemoresistance 38 of oral cancer. In addition to this, expression levels of a number of miRNAs correlate with clinicopathological variables 33 and have a diagnostic and prognostic value in oral carcinoma. 31 Table 1 shows the commonly deregulated miRNAs in oral cancer along with their target genes.
Commonly deregulated miRNAs and their associated signal pathways/target genes in OSCC/HNSCC.

MiRNA: microRNA; OSCC: oral squamous cell carcinoma; HNSCC: head and neck squamous cell carcinoma; OLP: oral leukoplakia.
Some important candidate miRNAs in oral cancer etiology and progression have been proposed, including loss of miR-34b, miR-100, miR-125b, miR-137, miR-193a, and miR-203, which seems to occur specifically in OSCC compared to normal oral mucosa.32,41 Overexpression of miR-21, miR-181b, and miR-345 has been suggested as a signature of malignant transformation of oral precancerous lesion, leukoplakia, for which histological assessments have limited prognostic value. 35 Downregulation of miR-125a, miR-184, and miR-16 and upregulation of miR-96 in both oral tumors and oral surgical margins suggest that the combinatorial regulation by these miRNAs and their target transcription factors are associated with oral tumorigenesis and are of value in the detection of minimal residual disease in surgical margins. 73
MiRNAs in OSCC—mechanisms of deregulation
More than a few studies point out that single nucleotide polymorphisms (SNPs) in pri-miRNAs and pre-miRNAs influence processing and expression of mature miRNAs, contributing to cancer initiation and progression. Roy et al. 74 studied SNPs at pri-miR and pre-miR regions of eight miRNA and four miRNA processing genes in oral cancer patients and stated that genetic variation at miRNA and processing genes altered the risk of oral cancer. A case control study in individuals with leukoplakia and/or erythroplakia showed that rs7372209 SNP in pre-miR-26a-1 leads to reduced or deleted expression of mature miR-26a-1, causing a 2.5-fold increased risk of oral premalignant lesions (OPLs). 75 In a study of 660 patients, polymorphisms in miR-146, miR-149, miR-196, and miR-499 were found to significantly increase the risk of HPV16-associated oropharyngeal squamous cell carcinoma, particularly in never smokers. 76 Furthermore, there was a significant association between SNPs of these four miRNA genes and an increased risk of OSCC associated with betel nut chewers and smokers. 77
As the genes encoding miRNAs are frequently located in fragile sites, it is plausible that loss or gain of chromosomal material could alter miRNA expression. 78 For instance, the expression of miR-100 and miR-125b, two miRNAs mapping to chromosome 11q, is significantly decreased in OSCC owing to the loss of chromosome material. 41
Deregulation of the key genes and proteins involved in the biogenesis of miRNAs might have profound effects on miRNA expression. For example, reduced expression of Dicer resulted in downregulation of mature miRNAs in a panel of human cancer cell lines. Dicer has also been implicated as a direct target of let-7 family miRNAs. 79 Reduced let-7b expression leads to Dicer upregulation facilitating OSCC cell proliferation. 80 Also, SNPs in miRNA biogenesis pathway genes might cause the deregulation of miRNA-processing proteins and have been linked to the increased risk of OPL, such as the SNP of rs3742330 in Dicer. 75
MiRNAs in OSCC—role of epigenetic modifications
Human cancers, including OSCC, are not only polygenetic but also polyepigenetic in nature. 32 The most common and readily detectable epigenetic event in cancer is inactivation of tumor suppressor genes 81 by DNA hypermethylation of CpG islands, while hypomethylation-associated activation of oncogenes is a less observed phenomenon.82,83 The transcriptional inactivation of miRNAs by CpG island hypermethylation suggests that epigenetic modification is a crucial factor in determining the expression of miRNAs in human cancers. 84 It is also a good marker to explore novel epigenetically silenced tumor suppressive (TS) miRNAs. Members of the miR-34 gene family (miR-34a, miR-34b, and miR-34c) are direct targets of p53, and their ectopic expression in cancer cells induces cell cycle arrest and apoptosis in part by targeting Myc. 85 The promoters of miR-34b and miR-34c genes are targets of CpG island hypermethylation in multiple malignancies including oral cancer. 32 MiRNA profiling of 18 OSCC cell lines indicated that CpG hypermethylation caused the downregulated expression of miR-34b, miR-137, miR-193a, and miR-203. 32 Uesugi et al. 48 identified two TS miRNAs, miR-218 and miR-585, frequently silenced by DNA hypermethylation in OSCC. Among these, miR-218 acts as a suppressor of the mechanistic target of rapamycin (mTOR)–Akt signaling pathway, independently of the phosphoinositide 3-kinase (PI3K)–Akt signaling pathway, in OSCC. The epigenetic silencing of miR-218 and consequent activation of the mTOR–Akt signaling pathway induced by the overexpression of Rictor (a possible target of miR-218) may contribute to oral carcinogenesis. Shia et al. 86 reported that epigenetic silencing of miR-329 and miR-410, two miRNAs from the 14q32.2 cluster, may contribute to Wnt-7b overexpression and activate the Wnt-β-catenin signaling pathway, thus promoting proliferation and invasion of OSCC. Besides, miR-137 is hypermethylated in OSCC and linked to poor survival rates.87,88 However, DNA hypermethylation epigenetically activates miR-200/miR-205 in tumors, which are repressed in the absence of hypermethylation, specifically in CD44 high OSCCs. However, there are indications that these miRNAs are comparatively repressed in disease with poor prognosis. 89
MiRNAs in the regulation of cancer hallmarks
MiRNAs form central nodal points in cancer development pathways. They have multifaceted roles in cancer-stem cell biology, angiogenesis, epithelial mesenchymal transition (EMT), metastasis, drug resistance, and so on. The conventional hallmarks of cancer progression have been described as self-sufficiency in growth signals, ignorance of antigrowth signals, apoptosis evasion, limitless replicative potential, sustained angiogenesis, and tissue invasion and metastasis. 90 Reprogramming of energy metabolism and evading immune destruction have been further considered as emerging hallmarks. 91 Deregulated miRs have been reported in each stage of oral cancer progression, from normal epithelium to metastatic cancer (Figure 2).
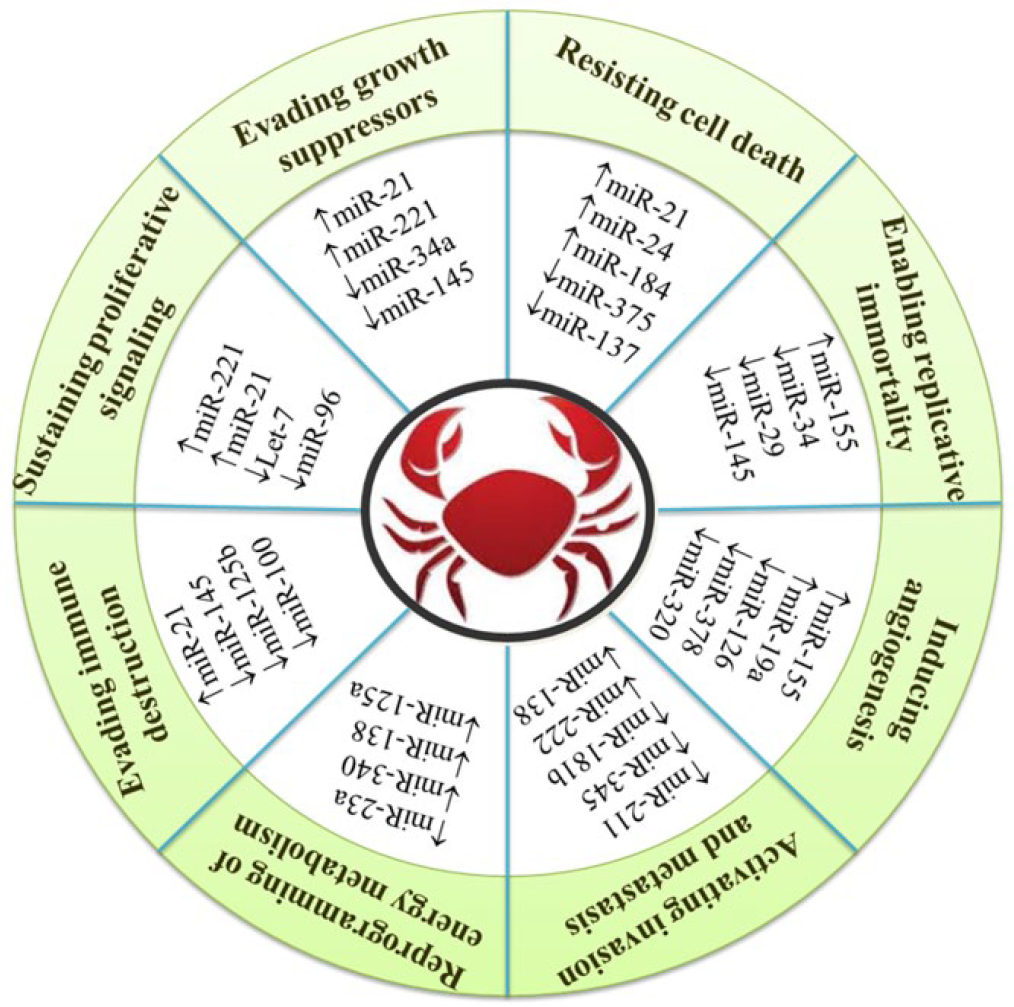
Association between miRNAs altered in oral cancer and various cancer hallmarks. The upregulated and downregulated miRNAs are depicted by up and down arrows, respectively.
Sustained proliferation
Progression through cell cycle is guarded by a series of checkpoints and regulatory proteins including cyclins and cyclin dependent kinases (CDKs). Any imbalance in the cell cycle regulatory pathways can lead to cancer development. Deregulated miRNAs enhance cell proliferation by targeting cell cycle regulators. MiR-137 and miR-193a downregulate the expression of CDK6 and E2F transcription factor 6 (E2F6), respectively, and, as a consequence, reduce OSCC cell growth. MiR-137 and miR-193a inhibit cell growth by G0–G1 arrest and caspase-mediated apoptosis, respectively.32,92 MiR-125b and miR-100 have also been proposed as tumor suppressors in OSCC inhibiting cell proliferation. EGFR3, a fibroblast growth factor receptor and predicted target for miR-100, is mutated in several malignancies including OSCC and is involved in development and proliferation. 41 The tumor suppressor miR-125a reduces proliferation and invasion of OSCC cells by posttranscriptional regulation of estrogen-related receptor alpha (ESRRA) expression. 57 In oral tumor samples, HOX-cluster-embedded miR-196a/b and miR-10b are upregulated and downregulated, respectively, and interfere in cell proliferation through distinct processes, possibly targeting a small set of genes involved in cell cycle progression. 93 In head and neck squamous cell carcinoma (HNSCC), restoring miR-375 expression inhibits cell proliferation and clonogenicity and accumulates cells in sub-G1. 92 MiR-503 and miR-15a, also significantly downregulated in HNSCC, have been confirmed to repress cyclin D142 and cyclin E expression, 40 respectively, resulting in inhibition of DNA synthesis. However, miR-155 has been suggested as an oncogene, and has been implicated in the regulation of cell survival, growth, and chemosensitivity. It downregulates expression of the tumor suppressor CDC73 and is itself regulated by the transforming growth factor-β (TGF-β)/SMAD signaling pathway, which is frequently activated in HNSCC. 71
Apoptosis evasion
MiR-21 is frequently overexpressed in tongue squamous cell carcinoma (TSCC) and is associated with poor prognosis by virtue of inhibiting apoptosis. In contrast, its downregulation is found to be associated with chemoresistance. 38 In TSCC, upregulation of miR-21 was found to be associated with low levels of tropomyosin 1 (TPM1) and phosphatase tensin homologue (PTEN) expression. 33 A large number of other significant oncogenic and tumor suppressor targets have been identified for miR-21 in head and neck and other cancers including Ras, programmed cell death 4 (PDCD4), reversion inducing cysteine rich protein with kazal motifs (RECK), and HNRPK.49,65,94–96 Moreover, Zhang et al. 97 suggested that miR-21 could affect formation of reactive oxygen species (ROS) by directly attenuating superoxide dismutase (SOD) family member 3, SOD3, and by an indirect mechanism that limited tumor necrosis factor (TNF)-α production, thereby reducing SOD2 levels to promote tumorigenesis. Likewise, upregulation of miR-2463 and miR-18462 is associated with enhanced proliferation and reduced apoptosis. 63 Overexpression of miR-184 is anti-apoptotic in nature as it targets c-Myc as well as TNFAIP2, an apoptosis-related gene.47,62 Another study has proposed a different explanation for the effects of miR-184 on epithelial cells and cancer cell lines via Akt signaling as ectopic expression of miR-184 suppresses Akt pathway that is associated with increased cell apoptosis and death. 98 Upregulated miR-24 in OSCC seems to directly target RNA binding protein dead end 1 (DND1) contributing to apoptosis resistance and proliferation through downregulation of cyclin dependent kinase inhibitor 1B (CDKN1B), one of the downstream genes of DND1. 63 Furthermore, aberrant reduction of miR-133a and miR-133b in TSCC is accompanied by overexpressed oncogene pyruvate kinase type M2 (PKM2) causing enhanced cell proliferation and reduced apoptosis. 39
Limitless replicative potential
Acquirement for human telomerase reverse transcriptase (hTERT) expression is an early event in oral carcinogenesis. The hTERT gene encodes the catalytic unit of telomerase that allows the maintenance of telomere length and also an escape from replicative senescence. MiR-31 has been reported to activate hypoxia pathways by targeting the FIH (factor-inhibiting hypoxia-inducible factor (HIF)-1α) gene. 64 Hence, during hypoxic conditions, telomerase is activated by transcriptional activation of hTERT, which is in turn regulated by the transcription factor HIF-1α. In vitro studies also indicate that the exogenous introduction of miR-31 along with hTERT into normal oral keratinocyte cells (NOKs) causes immortalization of oral keratinocytes. 69 Studies in other cancers point out hTERT-mediated tumor metastasis via indirect targeting of ITGB1. 99 In OSCC, miR-124 was reported to inhibit metastasis by the downregulation of ITGB1 expression indicating a plausible association between miR-124 and hTERT. 43 Another TS miRNA, miR-512-5p suppresses tumor growth by targeting hTERT in telomerase positive HNSCC. 100
Sustained angiogenesis
Hypoxia-inducible factor (HIF) regulates many genes involved in tumor angiogenesis and metastasis. Oncogenic miR-31 and its passenger strand miRNA (miR-31*) have been shown to be upregulated in oral leukoplakia (OLP) and OSCC.64,68,69 In HNSCC, aberrantly overexpressed miR-31 directly represses the expression of factor-inhibiting HIF (FIH), an inhibitor of HIF, and activates the HIF pathway under normal growth conditions. Elevated expression of miR-31 relates with upregulation of vascular endothelial growth factor (VEGF) and downregulation of E-cadherin in oral premalignant disease. In addition, miR-31-FIH-HIF-VEGF regulatory cascade affects several biological processes such as cell proliferation, migration, and EMT in OSCC cells. 64 In head and neck cancer cell lines studied under hypoxia conditions, partial regulation of the High Mobility Group A2 (HMGA2) protein, a member of the HMGA family, was identified to be controlled at least in part by miRNA-98. 61 Conversely, miR-320 is downregulated in OSCC, with its expression correlating inversely with the vascularity. Hypoxia suppressed miR-320 expression through HIF-1α and increased that of neuropilin 1 (NRP1) as well as promoted the motility and tube formation ability of endothelial cells via VEGF signaling pathway, resulting in tumor angiogenesis. 58 Another TS miRNA, miR-34a is reported to inhibit tumor angiogenesis by blocking VEGF production as well as by directly inhibiting endothelial cell functions. 50
Activating invasion and metastasis
Despite intensive local treatment, advanced HNSCC often have a poor prognosis due to tumor recurrences and distant metastases. Nearly 50%–60% of patients with advanced disease develop local or regional recurrences after treatment. 101 The spread of neoplastic cells to locoregional lymph nodes is a highly predictive factor in prognosis of HNSCC patients. Routine clinical tumor stage classifications are inadequate in searching for local cervical metastases and micrometastases, because of which miRNAs are considered as new factors with potential prognostic value. MiR-205 is one such plausible lymph node metastasis marker, since it is often abundantly expressed in squamous cell epithelium (but not in normal lymph nodes). 102 MiR-10b also actively participates in oral cancer formation by promoting cell migration and invasion. 103 Furthermore, it is considerably elevated in the plasma of patients with oral cancer and precancer suggesting that miR-10b possesses a high potential to discriminate the normal subjects.
Reduced levels of miR-138 and miR-222 have been shown to be correlated with elevated risk for TSCC metastasis. MiR-138 downregulation enhances metastasis in TSCC, potentially through regulation of RhoC and ROCK2 Rho GTPases. It is substantiated by repression of RhoC and ROCK2 in TSCC cells upon miR-138 restoration, resulting in reorganized cell stress fibers, round bleb like cell morphology, and suppressed cell metastasis.36,37 In sharp contrast, inhibition of miR-138 in TSCC cells enhanced the expression of RhoC and ROCK2, leading to enhanced cell stress fiber formation and elongated cell shape, as well as accelerated cell metastasis. 42 In addition, miR-138 regulates the expression of key EMT-related molecules like Fos like antigen 1 (FOSL1), vimentin (VIM), zinc finger E-box binding homeobox 2 (ZEB2), and enhancer of zeste homologue 2 (EZH2).44,45
Recent studies have shown that G protein–coupled receptor kinase interacting ArfGAP 1 (GIT1), an important scaffold protein for focal adhesion complexes, plays a pivotal role in cancer cell migration/invasion and metastasis. Huang et al. 54 reported miR-491-5p to be a metastatic suppressor in oral cancer cells through regulation of its downstream target, GIT1, resulting in perturbation of FAK/paxillin and extracellular signal–regulated kinases 1/2 (ERK) signaling as well as matrix metalloproteinase (MMP) expression and activity. Hence, they propose that miR-491-5p and GIT1 could potentially serve as metastatic prognosis biomarkers and targets for intervention of OSCC metastasis. Another important protein associated with tumor cell invasion and migration is matrix metalloproteinase 1 (MMP1) which is involved in the degradation of the extracellular matrix (ECM). Liu et al. 37 observed that miR-222 thwarted the expression of MMP1 by both direct targeting of the 3′-UTR of MMP1 mRNA and indirect inhibition of MMP1 expression by targeting the 3′-UTR of SOD2 mRNA, resulting in inhibition of TSCC metastasis. Likewise, miR-29a is downregulated in OSCC tissues and inhibits MMP2 expression by directly binding to its 3′-UTR. Functional studies revealed that transfection with miRNA-29a mimics attenuated invasive potential, increased apoptosis rate, and enhanced chemosensitivity of OSCC cell lines to cis-platinum (CDDP). 56 MiR-211 too is associated with the most advanced nodal metastasis and vascular invasion of OSCC. Besides, miR-211 overexpression strongly enhances cell proliferation, migration, and concomitant anchorage independent colony formation. 104 MiR-204, underexpressed in HNSCC, functions as a metastasis suppressive gene, reintroduction of which generates a significant inhibition of cell motility and adhesion in vitro, and lung metastasis in vivo. 105 Our understanding of the functions of these miRNAs may help us better comprehend their role in oral cancer.
Reprogramming of energy metabolism
Metabolic reprogramming of cancer cells is an indispensable event for tumorigenesis and metastasis. The interplay between oncogenic processes and metabolic reprogramming is finely regulated by miRNAs. 106 Recently, Xu et al. 107 reported the expression of miR-340 to be decreased in OSCC which induced a metabolic switch in oral cancer cells. MiR-340 downregulation was associated with increased glucose transporter-1 (Glut1) expression, and subsequent increase in lactate secretion and glucose uptake rate. Hence, miR-340-induced altered metabolism results in rapid proliferation of oral cancer cells by regulation of glycolysis. Furthermore, miR-23a upregulation controls SMAD4 expression and consequently GLUT4 translocation thereby indirectly regulating glucose transport. 108
Malignant tumors procure high glycolytic phenotype even when adequate oxygenation is present (Warberg effect). In order to attain this, cancer cells deregulate the irreversible steps of glucose metabolism. For instance, miR-143 expression is inversely correlated with hexokinase 2 (HK2) levels, and upregulated HK2 in OSCC promotes the metabolic shift toward aerobic glycolysis. The miRNA-dependent regulation of hexokinase expression is not limited to HK2 as miR-138 targets HK1. 109 The best example for oncogene-driven metabolic alteration is that mutation in Ras oncogene promotes glycolysis. As miR-206 directly targets Ras to inhibit OSCC progression, it can be considered as miRNA-mediated indirect regulation of cell metabolism. 110
Evasion of anti-tumor immune responses
The role of immune system in tumor formation is still an unresolved issue. However, the immune system operates as a significant barrier to tumor formation and progression. Counterintuitively, the infiltration of neoplastic tissues by cells of the immune system serves to promote tumor progression. 91 However, tumor establishment necessitates evasion of immune response by tumor cells wherein miRNA dysregulation appears to make a significant contribution. Apoptosis of circulating T cells seems to be a generalized phenomenon in OSCC patients, which also correlates with Fas ligand (FasL) expression on tumor cells. 111 Fas is reported to be regulated by let-7 family of miRNAs, which are downregulated in OSCC. 80
MiRNAs in the evolution of tumor microenvironment
The tumor microenvironment comprises of fibroblasts, immune cells, endothelial cells, and ECM, proteases, and cytokines. It evolves by complex crosstalk with the cancer cells they harbor, hence facilitating tumor formation and progression. 91 MiRNAs in the tumor cells and microenvironment have distinct functions which are transformation of the tumor microenvironment by paracrine mechanisms and stabilization of cancer hallmarks, respectively. Cancer-associated miRNAs affects three major components of the tumor milieu—vasculature, ECM, and immune cells. Alterations in the miRNA profiles of neighboring cells that lack genetic abnormalities favor the acquisition of cancer hallmark traits. 112 The role of miRNAs in angiogenesis and immune evasion has been discussed in previous sections.
Regulation of tumor stromal cells by miRNAs
The tumor microenvironment is an aggregate of fibroblasts, endothelial cells, pericytes, and immune inflammatory cells, which favors cancer growth. Of these, cancer-associated fibroblasts (CAFs) differ from normal fibroblasts by their high expression of α-smooth muscle actin (SMA) and their pro-tumorigenic properties. They enhance ECM production and secrete cancer-activating cytokines and chemokines for crosstalk with cancer cells, significantly promoting tumorigenesis. 113 A high frequency of CAFs in patients with mobile tongue cancer has a negative impact on recurrence and is associated with patient’s poor survival. 114 Accumulating data indicates that normal fibroblasts and CAFs exhibit distinct miRNA profiles and that miRNAs involved in CAFs contribute to fibroblast activation and cancer cell metastasis. 112 For instance, miR-148a is downregulated in CAFs compared with normal fibroblasts isolated from clinical OSCC tissue. Upon overexpression, miR-148a significantly impaired the migration and invasion of oral carcinoma cells by directly targeting WNT10B. 115 Likewise, miR-21 expression in OSCC is predominately localized to tumor stromal cells with colocalization between miR-21 and α-SMA. 116
MiRNAs in development of drug resistance
MiRNA deregulation is also associated with development of drug-resistant phenotype in OSCC. Cisplatin is an effective chemotherapeutic against OSCC as a single agent as well as in combination with other drugs. However, the intrinsic/acquired chemoresistance to cisplatin is an important clinical issue. Comparative analysis in parental and cisplatin resistant cell lines revealed resistance-specific miRNA signature which modulates cancer stem-cell-like and EMT-type properties in OSCC. 117 In the TSCC cell line UM1, antisense miR-222 transfection combined with cisplatin treatment was found to reduce cell proliferation and promote apoptosis by downregulating PUMA gene thus offering novel rationales for combinational therapy. 118 Another study on cisplatin-sensitive and resistant TSCC cell lines suggested the existence of differential miRNAs with chemosensitivity and chemoresistance. Among the altered miRNAs, miR-21 was identified as chemosensitive, while miR-214 and miR-23a were chemoresistant. 38
MiRNA signatures classify cancer
Each tumor type manifests a distinct miRNA signature that differentiates it from normal tissues and other tumor types. Most cancers can be further sub-classified into prognostic groups based on these signatures. 13 Jiang et al. 119 employed real-time quantitative polymerase chain reaction (PCR) to successfully identify miRNA deregulations in 32 cancer cell lines among which five were from the head and neck/oral cavity. Clustering analysis based on the miRNA precursor expression values enabled most of the cancer cell lines to be clustered based on the tissue of origin. This suggests that each cancer exhibits a unique miRNA expression profile/signature based on its specific tissue of origin. Avissar et al. 31 claimed that the expression ratio of miR-221:miR-375 showed a 92% sensitivity and 93% specificity to classify OSCC. Barker et al. 120 found that miR profiles were distinct and specific for HNSCC in the tonsil, base of tongue, and postnasal space. In addition, the miR expression profiles between primary cancer and its nodal metastatic disease were consistent implying that miR profiles may be used as a diagnostic tool to determine whether the nodal metastasis is from the oral cavity, particularly when the primary tumor cannot be identified on the basis of tumor biopsy samples. On the contrary, Hui et al. 92 did not find distinct expression profiles between the three subsites investigated—the hypopharynx, oropharynx, and larynx. This conflicting data may be due to technical problems as well as differences in stage, grading, and sampling from multiple anatomical sites but signifies the need for further analysis.
MiRNAs as diagnostic tools of oral cancer
Since distinct miRNA expression profiles exist for OSCC and normal tissues, miRNA expression analysis offers an opportunity for early stage diagnosis of OSCC. Correspondingly, a number of individual miRNAs have been advocated as putative specific biomarkers for OSCC diagnosis, such as aberrantly overexpressed miR-21 and miR-205. 121 In addition, miRNAs seem to hold significant potential as diagnostic tools to detect metastatic disease. Abundantly expressed in HNSCC cells, miR-205 could sensitively and accurately detect cervical lymph node metastases from HNSCC, including OSCC. This is reinforced by the finding that miR-205 expression level examined using quantitative reverse transcription polymerase chain reaction (qRT-PCR) could detect metastatic HNSCC in each positive lymph node specimen. In addition, HNSCC positive lymph nodes could be distinguished from metastases from other primary tumors and, intriguingly, as little as one HNSCC cell could be detected from a background of 1 million lymphocytes using an in vitro lymphoid tissue model. 102 More strikingly, miRNA-based classifier has been demonstrated to be more accurate than mRNA-based classifier to diagnose metastatic cancer of unknown primary origin. 13
MiRNAs as predictive biomarkers in oral carcinoma
Our progressively greater understanding of the molecular alterations underlying cancer has created opportunities for more accurate and meaningful diagnosis and prognosis than was previously possible. Now, entering into the era of personalized medicine, patient-management decisions increasingly depend on molecular analyses. In addition to specific genetic alterations, additional molecular features including DNA methylation, gene expression, and miRNA expression can provide vital clinical information. MiRNAs may have greater utility than mRNAs as prognostic indicators owing to their stability within clinical samples and their robust expression patterns. 13 Many signatures have been proposed, and their use is being investigated in clinical trials. Until fairly recently, miRNA analysis was performed using qRT-PCR and microarray-based approaches. Next-generation sequencing (NGS) has emerged as a cost-effective option, and the barriers to bioinformatics analysis are no longer a daunting prospect to the non-specialist laboratory. 122
The relative levels of a small number of miRNAs have been revealed to have profound prognostic values in determining the survival of patients with OSCC. By multivariate analyses, miR-21 expression is proposed as an independent predictor of poor survival for patients with TSCC. There is a strong association between high-level expression of miR-21 and significantly decreased 5-year survival of patients with HNSCC. 33 Moreover, low-level expression of miR-205 was found to significantly correlate with locoregional relapse of HNSCC, independent of disease severity at diagnosis and treatment. 89 Downregulated miR-205 and let-7d combined is further suggested as a robust predictor of shorter HNSCC survival. 89
Cervigne et al. 35 demonstrated that miR-21, miR-181b, and miR-345 expression levels were recurrently increased and linked to higher severity during OSCC progression. They identified miRNA expression profiles that could separate progressive leukoplakia and OSCC from non-progressive leukoplakia and normal tissues, showing the potential of using miRNAs as reliable indicators of leukoplakias at high risk of malignant transformation. Other reports also indicated that miR-21 and miR-221/miR-222 are involved in progression from OLP to oral cancer.33,123 Furthermore, miRNA expression level seems to change with different malignancy grades and reflect the risk of OPL or the biological characteristics of OSCC, such as the metastatic potential, possibility of relapse, and chemosensitivity/chemoresistance. 104 It supports the notion that miRNA-based staging might offer adequate therapeutic biological information, ultimately advancing the clinical management of OPL and OSCC.
In the past few years, there have been several reports indicating the presence of salivary miRNAs and their potential as noninvasive biomarkers in oral cancer detection. How salivary mRNA and miRNA biomarkers are safeguarded and shuttled from the source of the tumor to the saliva is intriguing. Exosome, a macromolecule, protects salivary mRNAs and miRNAs from ribonucleases present in the saliva. Exosomes are small, cell-secreted vesicles around 30–100 nm in length that are known to package and transport mRNAs and miRNAs. By being localized inside exosomes, salivary mRNAs were found to be remarkably stable. 124 This stability has also been exhibited by endogenous salivary miRNAs which degrade at a much slower rate than exogenous miRNAs. 125 Exosomes have been shown to transfer mRNA from different cell types and activate or modulate gene expression in oral keratinocytes. 124 MiR-125a and miR-200a are two salivary miRs that have been shown to be substantially reduced in oral carcinoma patients versus healthy controls. 125
Likewise, miRNAs in plasma samples exhibit high stability and are resistant to RNase activity. This stability is due to their inclusion in lipid or lipoprotein complexes, such as apoptotic bodies, microvesicles, or exosomes, which prevent their degradation by RNases. 126 The stability combined with the good accessibility make circulating miRNAs attractive biomarker candidates. Circulating miR-142-3p, miR-186-5p, miR-195-5p, miR-374b-5p, and miR-574-3p represent the most promising markers for prognosis and therapy monitoring in the plasma of HNSCC patients. 127 In tongue carcinoma, the plasma level of miR-184 is found to be amplified when compared to control. 62 Similarly, amplified plasma levels of miR-31 and miR-24 have been detected in oral carcinoma.64,128 Moreover, plasma miRNAs are shown to be reduced after tumor resection, implying that they may be released from cancerous tissues into circulation and hence is a potential marker for disease progression.62,64 This was seen in case of plasma and salivary miR-3147 and plasma miR-184. 62
Concluding remarks and future perspectives
Complexity of cancer progression mechanisms in higher organisms is thought to also be achieved through controlled and coordinated gene regulatory networks of miRNAs. Alterations in miRNA function and regulation have rapidly emerged as important players in cancer pathogenesis. In comparison to mRNA-based studies, miRNA expression analysis is more effective in cancer classification and posttranscription level regulation studies. Recent work has revealed intriguing changes in the global state of miRNA expression in cancers including oral cancer. Actuating evidence has identified miRNA deregulation in OSCC and OPLs to be also regulated by cellular and epigenetic factors. Emergence of such multiple regulatory networks, with their potential interactive action, contributes to cellular transformation. Despite remarkable recent progress in understanding of functional mechanisms of miRNA in various human cancers, the mechanisms, targets, and extent of miRNA-mediated gene expression in oral cancer is unclear. An expanding knowledge of specific roles of deregulated miRNAs could further contribute to our understanding of the complexity of tumor behavior and progression. Future studies focusing on the correlated expression of TS and oncogenic miRNAs in OSCC and OPLs as well as associating them with tumor stages could probably initiate a revolution in prognostic analysis. The frequency and potential reversibility of miRNA alterations make them potential adjuvants to chemotherapy and radiotherapy. Research prospects in circulating miRNA detection and analysis may pave way to development of novel minimally invasive or noninvasive cancer biomarkers. Furthermore, analysis of the miRNA expression pattern in surgical margins and OPLs could give important information about residual tumor extension and dysplasia to malignancy transition, respectively. Consideration of this information would lead to the development of novel therapeutic targets and agents for targeted therapy and could potentially enhance our abilities for customized patient-centered treatment especially when other established therapies have failed.
Footnotes
Acknowledgements
The first author, V.G.M., would like to acknowledge for the fellowship provided by University Grants Commission, New Delhi, India. The authors also acknowledge the Department of Biotechnology, Government of India, for their financial assistance.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
