Abstract
BACKGROUND:
Toll-like receptors participate in various biological mechanisms, mainly including the immune response and inflammatory response. Nevertheless, the role of TLRs in STAD remains unclear.
OBJECTIVE:
We aimed to explore the expression, prognosis performance of TLRs in STAD and their relationship with immune infiltration.
METHODS:
Student’s t-test was used to evaluate the expression of TLRs between STAD tissues and normal tissues. Kaplan-Meier method was applied to explored the prognosis value of TLRs in STAD. And qRT-PCR validated their expression and prognosis value. Spearman’s correlation analysis and Wilcoxon rank-sum test were used to assess the association between TLRs and immune infiltration in STAD.
RESULTS:
The mRNA level of TLR3 was downregulated in STAD. We summarized genetic mutations and CNV alteration of TLRs in STAD cohort. Prognosis analysis revealed that STAD patients with high TLR3 expression showed better prognosis in OS, FP and PPS. The result of qRT-PCR suggested that TLR3 expression was decreased in STAD tissues and STAD patients with high TLR3 mRNA level had a better OS. Univariate and multivariate cox regression analysis suggested TLR3 expression and clinical stage as independent factors affecting STAD patients’ prognosis. A positive association existed between TLR3 expression and the abundance of immune cells and the expression of various immune biomarkers. Furthermore, key targets related to TLR3 were identified in STAD, mainly including MIR-129 (GCAAAAA), PLK1, and V$IRF1_01.
CONCLUSIONS:
Our result demonstrated TLR3 as a prognosis marker and associated with immune infiltration in STAD.
Keywords
Introduction
Gastric cancer (GC) is considered one of the severe diseases worldwide; gastric cancer ranks fifth among all cancers in terms of incidence and is the third leading cause of cancer-related death [1]. Stomach adenocarcinoma (STAD) is the most common subtype of gastric cancer (GC). Eighty percent of STAD patients are diagnosed at advanced stages because of the lack of symptoms at early stages, endoscopy, and cancer screening [2, 3]. The prognosis of advanced STAD patients is so poor that the 5-year survival rate is less than 30% [4]. Currently, surgery and chemotherapy are still the primary approaches for the treatment of STAD [5]. Although new therapies, such as immune and targeted therapies, have been utilized in recent years, there has still limited improvement in the prognosis of STAD patients [3, 6]. Thus, more survival biomarkers and therapeutic targets for STAD are needed.
Toll-like receptors (TLRs) are a class of single, membrane-spanning, noncatalytic proteins, and this class consists of 10 members (TLR1–TLR10) in human [7]. TLRs are also called pattern-recognition receptors and are associated with various biological mechanisms, such as activating the immune system, inducing inflammatory responses and regulating the proliferation, apoptosis and invasiveness of various types of cancer cells [7, 8]. The potential applications of TLRs in cancer were also focused by clinicians. Numerous studies have revealed that some TLRs (TLR1 to TLR10) could be potential prognostic biomarkers or therapeutic targets in certain types of cancer, mainly including oral squamous cell carcinoma, pancreatic cancer, hepatocellular carcinoma, neuroblastoma, thoracic esophageal squamous cell carcinoma and colorectal cancer [9, 10, 11, 12, 13, 14]. In cancer therapy, some of TLR-targeted agents were under clinical estimation. A multi-center study reported that a combined therapy, including poly-ICLC (a TLR3 agonist), radiotherapy and temozolomide, was utilized in glioblastoma treatment, and the improved effects were presented [15]. In phase clinical trial, OM-174, a TLR2 and TLR4 agonist, was found to benefit with refractory solid tumor [16]. Genetic variations of TLRs also played a crucial role in cancer diagnosis and prevention. Semlali et al. reported two kinds of TLR2 single-nucleotide polymorphisms (SNPs) might predict risk of colon cancer development, and TLR6 SNP rs5743810 might be used in the diagnosis of breast cancer [17, 18]. The above evidences showed potential clinical value of TLRs in various human cancer types. Nevertheless, few studies systematically elucidated the specific functions of TLRs in STAD.
Based on the emergence of a variety of reliable bioinformatics databases and the development of gene sequencing technology, comprehensive analysis of TLRs has become feasible. In the current study, we investigated the transcription level, prognostic performance and relative subgroup analysis of TLRs in STAD to select potential prognostic biomarkers from TLRs. Drug sensitivity and immune cell infiltration analyses conducted on STAD tissues were used to explore the resistance of TLRs to chemotherapy and potential immunotherapeutic targets among the TLRs. Cancer pathway activity analyses, functional enrichment analyses, and protein-protein interaction networks were conducted to validate the biological functions and molecular mechanisms of TLRs in STAD.
Materials and methods
Datasets and preprocessing
The RNA sequencing (RNA-seq) data of STAD (
Mutation analysis, drug sensitivity analysis and pathway activity analysis of TLRs in STAD
The mutation frequency and oncoplot of 10 TLRs in STAD patients was generated by “maftools” packages. The location of CNV alteration of 10 TLRs on 23 chromosomes was drawn by “RCircos” package. Moreover, a PPI network was constructed for 10 TLRs with Search Tool for the Retrieval of Interacting Genes (STRING) [23]. In drug sensitivity analysis, gene expression profiling data of 265 small molecules was obtained from Genomics of Drug Sensitivity in Cancer (GDSC). Pearson correlation analysis was performed to analyze the coefficients between TLRs and drug-targets and the detailed method could be found in Rees et al. [24]. In cancer-related pathway activity analysis, TLRs expression was divided into High expression group and Low expression group by median expression. The difference of pathway activity score (PAS) between groups is defined by Student’s
Prognosis analysis
Kaplan-Meier plotter (
Validation of the expression and prognosis value of TLRs in STAD
Approved by the Ethics Committee of the Second Affiliated Hospital of Nanchang University, we selected 48 STAD tissues and pair-normal gastric tissues from patients don’t receive any local or systemic treatment preoperatively (IRB No.2020(012)). Informed consent was obtained from each patient. All the Total RNA of STAD and normal tissue were extracted with TRIzol reagent (Vazyme, Nanjing). The synthesis of cDNAs corresponding to the mRNAs and qTR-PCR were performed using PrimeScript RT-polymerase (Vazyme). SYBR-Green Premix (Vazyme) with specific PCR primers (Sangon, Shanghai). Glyceraldehyde-3-phosphate dehydrogenase was used as an internal control. The 2
The transcription levels of TLR1 to TLR10 in different types of STAD and normal stomach tissues (Oncomine)
The transcription levels of TLR1 to TLR10 in different types of STAD and normal stomach tissues (Oncomine)
Abbreviation: TLR, toll-like receptor.
TIMER (
The transcription levels of TLRs in STAD. (A) The transcription levels of TLRs in STAD in Oncomine dataset. The numbers of datasets are shown in the figure. The red area represents upregulated expression of TLRs, while the blue area represents downregulated expression of TLRs. (B) The transcription levels of TLRs in STAD in TCGA dataset. Abbreviations: TLR, toll-like receptor; TCGA, the Cancer Genome Atlas; STAD, stomach adenocarcinoma. Note: *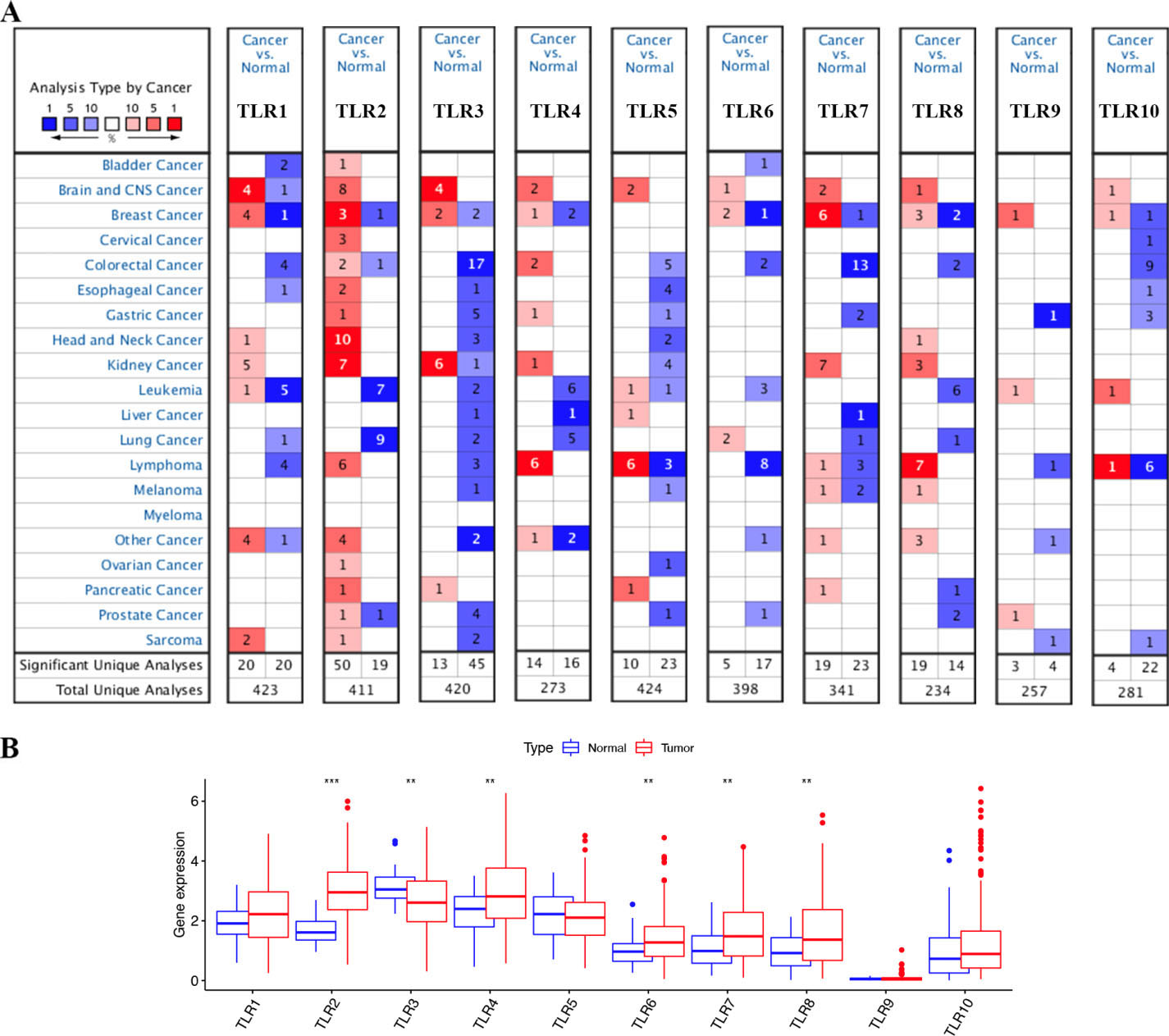
LinkedOmics (
Subgroup analyses of TLR2 expression in STAD. Subgroup analyses based on race (A), age (B), TP53 mutation status (C), gender (D), H. pylori infection status (E), nodal metastasis status (F), individual cancer stages (G), tumor grade (H), and histological subtypes (I) were conducted. Abbreviations: TLR, Toll-like receptor; STAD, stomach adenocarcinoma. Note: *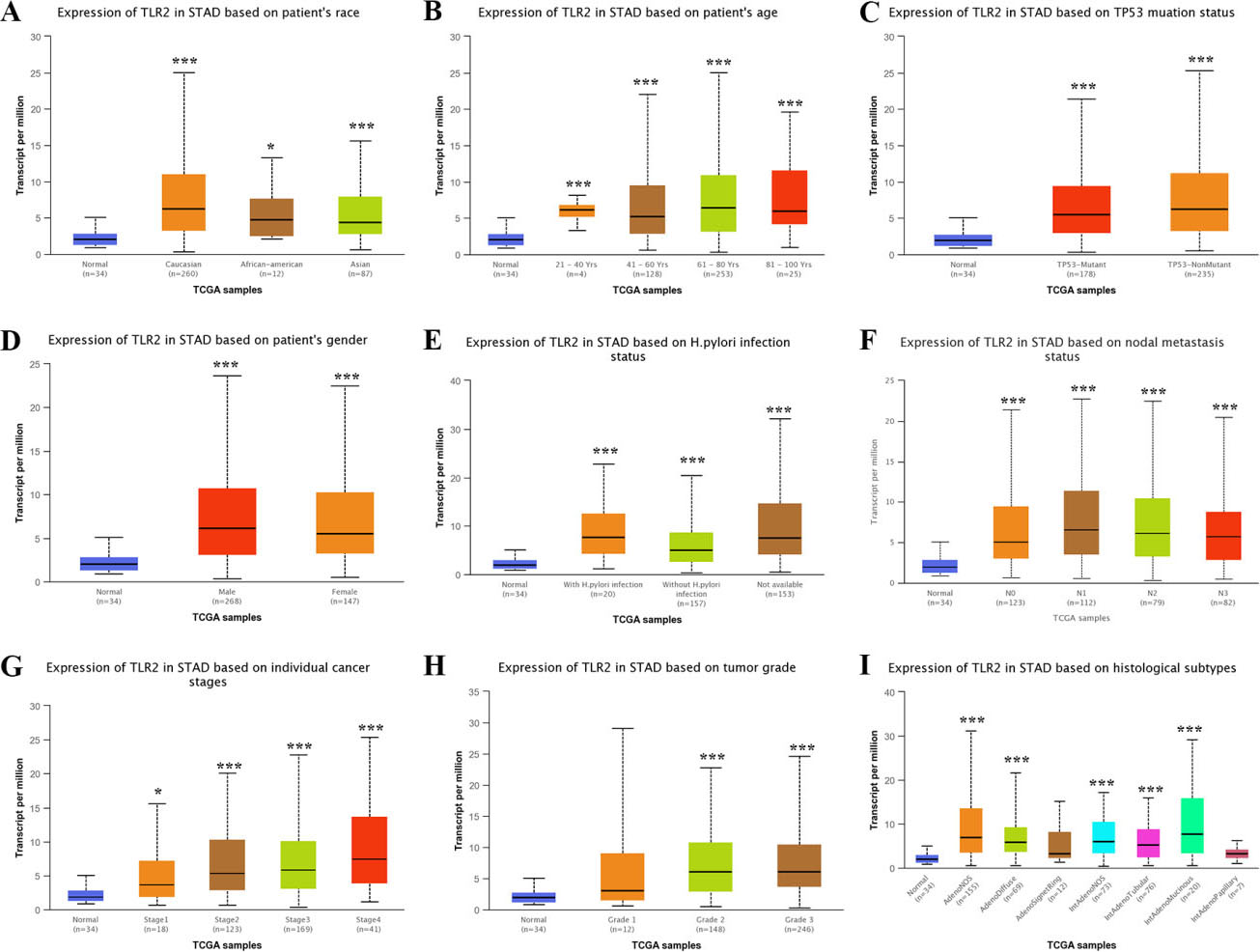
The expression of TLRs in STAD
In the Oncomine database, we explored the transcription level of TLRs (TLR1 to TLR10) in STAD tissues and normal tissues. As shown in Table 1 and Fig. 1A, TLR2 was overexpressed in STAD tissues, while TLR3, TLR5, TLR9 and TLR10 were downregulated in STAD tissues. DErrico et al. [19] showed elevated expression of TLR2 in STAD tissues (fold change
We then explored the mRNA levels of TLR2 and TLR3 in different subgroups of STAD tissues. In the subgroup analyses based on patient race (Fig. 2A), patient age (Fig. 2B), TP53 mutation status (Fig. 2C), patient gender (Fig. 2D), H. pylori infection status (Fig. 2E), nodal metastasis status (Fig. 2F), individual cancer stage (Fig. 2G), tumor grade (Fig. 2H) and histological subtype (Fig. 2I), the transcription level of TLR2 was significantly elevated in STAD tissues. Nevertheless, the transcription level of TLR3 in STAD tissues was significantly decreased in these subgroups (Fig. 3A–I).
Subgroup analyses of TLR3 expression in STAD. Subgroup analyses based on race (A), age (B), TP53 mutation status (C), gender (D), H. pylori infection status (E), nodal metastasis status (F), individual cancer stages (G), tumor grade (H) and histological subtypes (I) were conducted. Abbreviations: TLR, Toll-like receptor; STAD, stomach adenocarcinoma. Note: *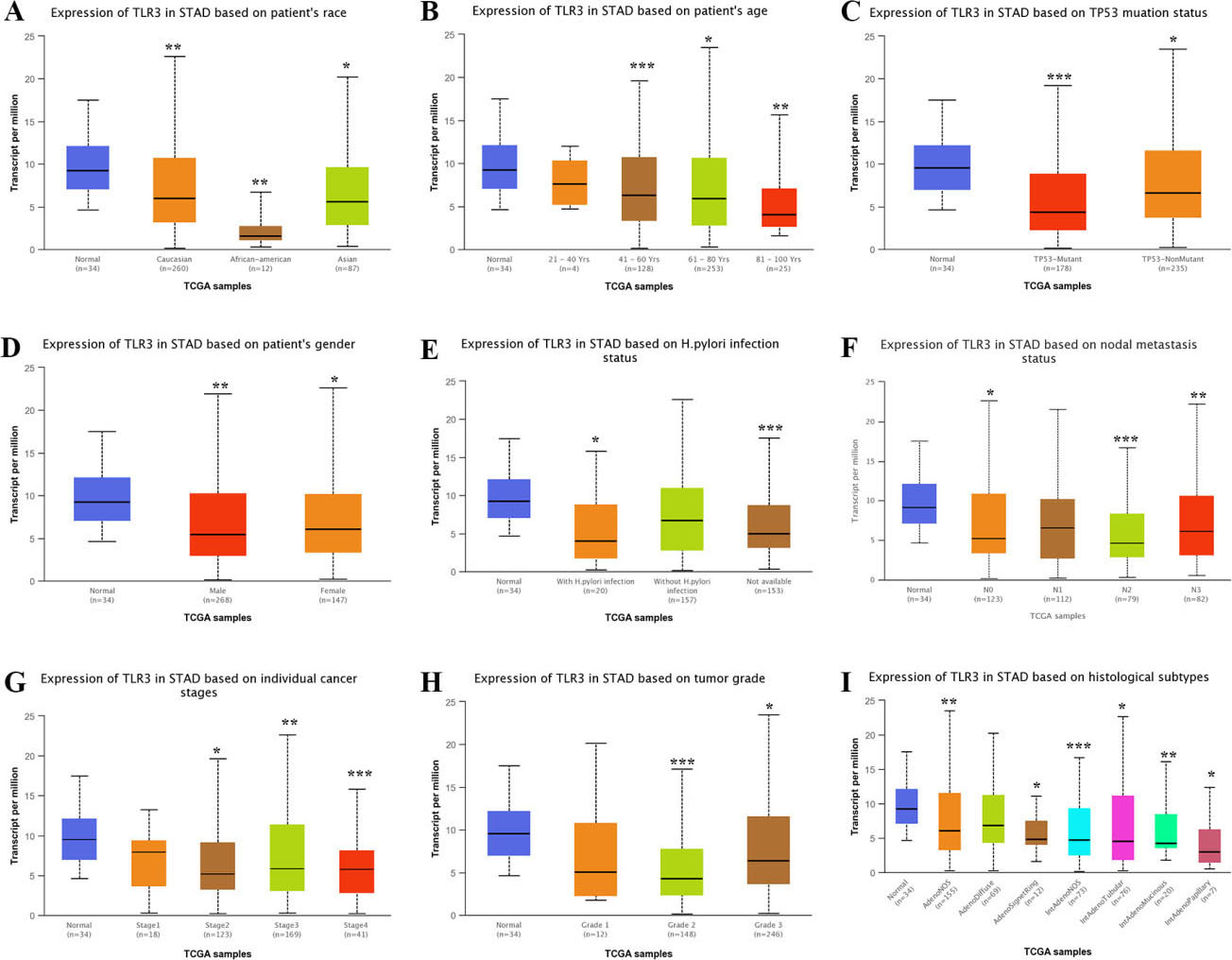
cancer hallmark analysis of TLRs in STAD. (A-B) The mutation frequency and classification of 10 TLRs in STAD. (C) The CNV variation frequency of 10 TLRs in STAD cohort. (D) The location of CNV alteration of 10 TLRs on 23 chromosomes in STAD cohort. (E) PPI network of 10 TLRs. (F) Drug sensitivity analysis of TLRs in STAD. A positive correlation means that a sample with gene expression is resistant to the drug, and vice versa. (G) Cancer pathway activity of TLRs in STAD. The activation and inhibition of TLRs occurs through several common cancer signaling pathways. Abbreviations: TLR, Toll-like receptor; STAD, stomach adenocarcinoma.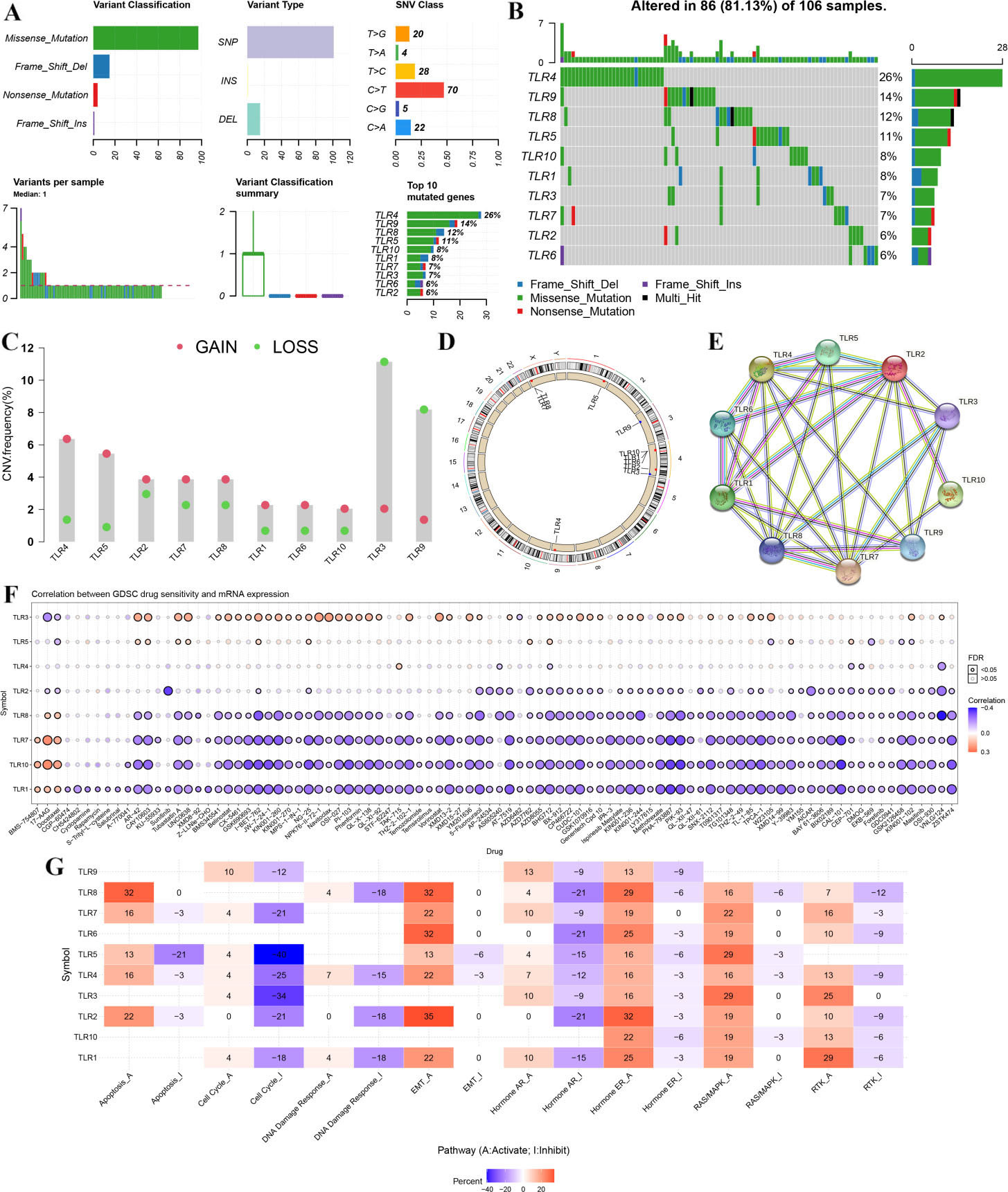
The prognostic value of TLR2/TLR3 in STAD. (A) OS of STAD patients with high/low expression of TLR2. (B) FP of STAD patients with high/low expression of TLR2. (C) PPS of STAD patients with high/low expression of TLR2. (D) OS of STAD patients with high/low expression of TLR3. (E) FP of STAD patients with high/low expression of TLR3. (F) PPS of STAD patients with high/low expression of TLR3. (G-H) OS of STAD patients with high/low expression of TLR3 in GSE29272 and GSE62254 cohort. Abbreviations: TLR, Toll-like receptor; OS, overall survival; FP, first progression; PPS, postprogression survival; HR, hazard ratio; STAD, stomach adenocarcinoma.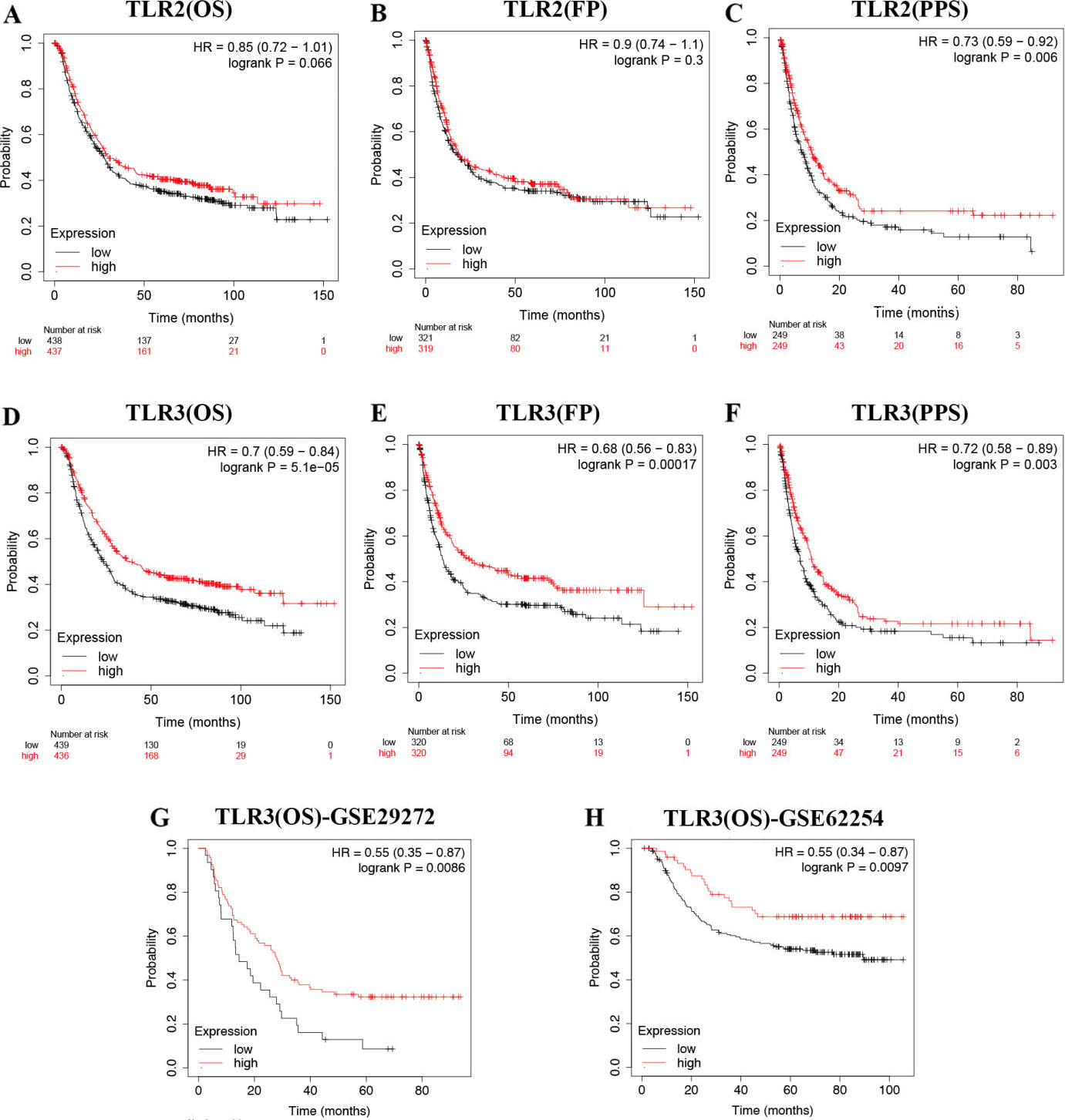
The expression and prognosis value of TLR3 in STAD. (A) The relative expression of TLR3 in STAD tissues and normal tissues. (B) Survival curve revealed the overall survival of STAD patients with high/low TLR3 expression. (C) The ROC curve of TLR3 in predicting the prognosis of STAD. (D-E) Univariate and multivariate analysis of TLR3 and clinical characters in STAD. Abbreviations: TLR, Toll-like receptor; STAD, stomach adenocarcinoma.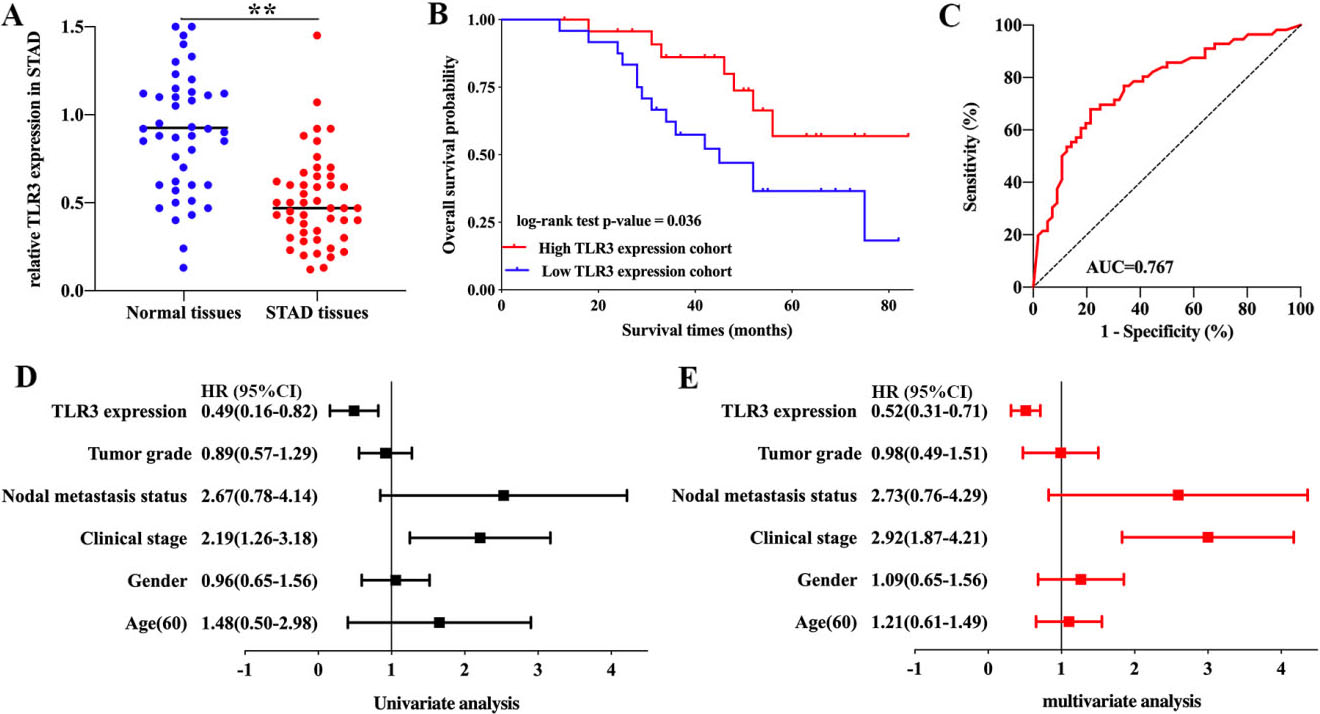
As shown in Fig. 4A and B, 86 of 106 (81.13%) STAD sample experienced genetic mutations. Among all the forms of variant classification, Missense mutation was the most common form (Fig. 4B). The result also suggested that SNP ranked the top variant type and C
Drug sensitivity and pathway activity analysis of TLRs in STAD tissues
To develop a therapy target, it is important to analyze correlation between gene expression and exited drugs. We then explored the correlation between TLR expression in STAD tissues and drug sensitivity based on data from GDSC. As a result, STAD tissues with high TLR3 expression and low expression of TLR1, TLR7, TLR8 and TLR10 were resistant to most small molecules or drugs from GDSC (Fig. 4F). The pathway activity analysis results revealed that TLRs mainly activated the apoptosis pathway, EMT pathway, hormone ER pathway, RAS/MAPK pathway and RTK pathway and inhibited the cell cycle pathway, DNA damage response pathway and hormone AR pathway in STAD tissues (Fig. 4G).
The prognostic value of TLR2 and TLR3 in STAD
Above results found that TLR2 was upregulated while TLR3 was downregulated in STAD. We then examined the prognostic performance of TLR2 and TLR3 in STAD. OS, FP and PPS curves of TLR2 are presented in Fig. 5A–C, respectively. The result indicated a better PPS (
The correlation between TLR3 and immune cell infiltration in STAD. (A) The correlation between TLR3 and the abundance of six types of tumor-infiltrating cells. (B) The correlation between SCNAs of TLR3 and immune infiltration levels in six tumor-infiltrating cells. (C-D) The correlation between TLR3 and TMB or MSI in STAD. Abbreviations: TLR, Toll-like receptor; cor, correlation coefficient; STAD, stomach adenocarcinoma; TMB, Tumor Mutation Burden; MSI, Microsatellite Instability. Note: *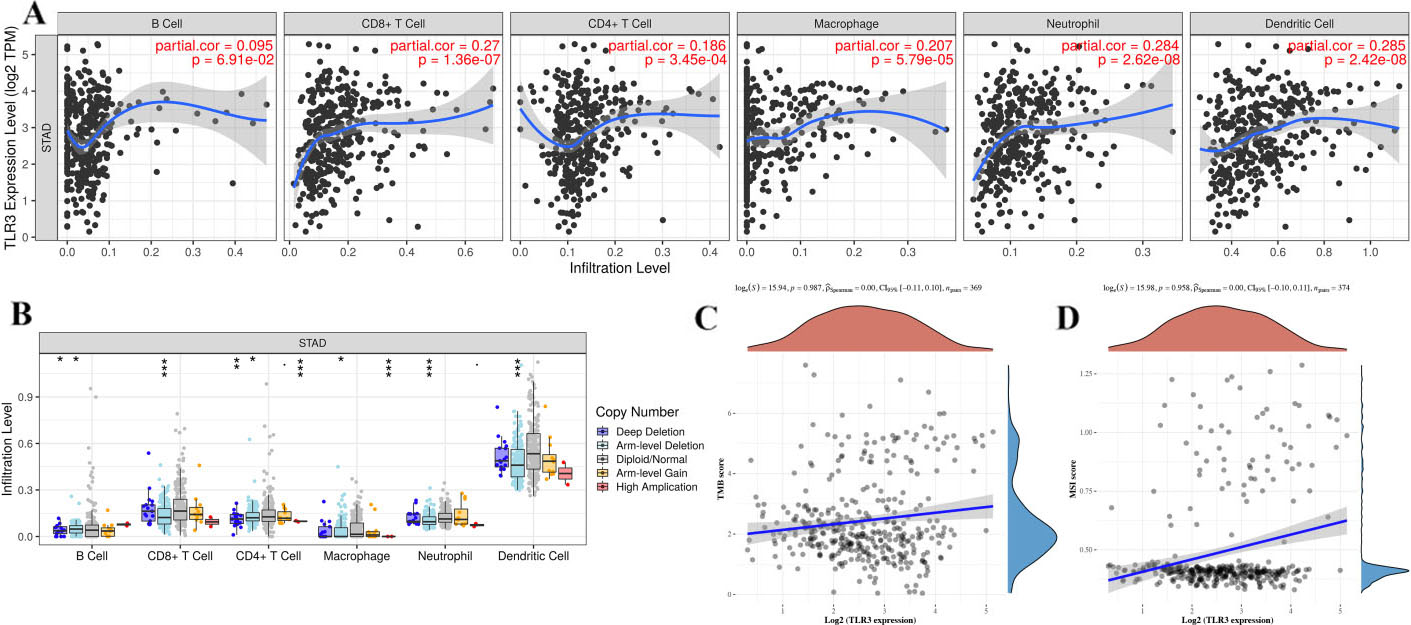
Next, we explored the correlation between TLR3 expression and OS, FP and PPS in STAD with different pathological parameters. We found that high TLR3 expression was associated with better OS in male and female STAD patients and in STAD patients with tumor stage 3, intestinal Lauren classification, lymph nodal metastasis stage 1 and lymph nodal metastasis stage
In order to further verify this result, we also performed prognosis analysis using GSE29272 and GSE62254 dataset, which also suggested that STAD patients with increased TLR3 levels showed a better OS rate in GSE29272 (Fig. 5G,
qRT-PCR was performed to further confirm the expression and prognostic value of TLR3 in STAD. As expected, the data suggested a decreased expression of TLR3 in STAD tissues (Fig. 6A,
Correlation of TLR3 and gene markers on tumor-infiltrating cells in STAD (TIMER)
Correlation of TLR3 and gene markers on tumor-infiltrating cells in STAD (TIMER)
Abbreviations: TLR, Toll-like receptor; STAD, stomach adenocarcinoma; TAM, tumor-associated macrophage; Cor, correlation coefficient. Note: *
Above result showed that TLR3 was downregulated and acted as a prognostic biomarker in STAD and we selected TLR3 for further analysis. Comprehensive analyses were conducted to evaluate the correlation between TLR3 expression and immune cell infiltration. We found that TLR3 expression was positively associated with CD8
Functional enrichment analysis of TLR3 in STAD. (A) The volcano plot of the genes positively (dark red dots) or negatively (dark green dots) associated with TLR3. (B) The heat map of the top 50 genes that positively correlated with TLR3. (C) The heat map of the top 50 genes that negatively correlated with TLR3. GO analyses, including BP analysis (D), CC analysis (E) and MF (F) analysis, were performed by GSEA. (G) KEGG pathway analysis was performed in GSEA. Abbreviations: TLR, Toll-like receptor; GSEA, gene set enrichment analysis; GO, Gene Ontology; BP, biological process; CC, cellular component; MF, molecular function; KEGG, Kyoto Encyclopedia of Genes and Genomes; HR, hazard ratio; STAD, stomach adenocarcinoma.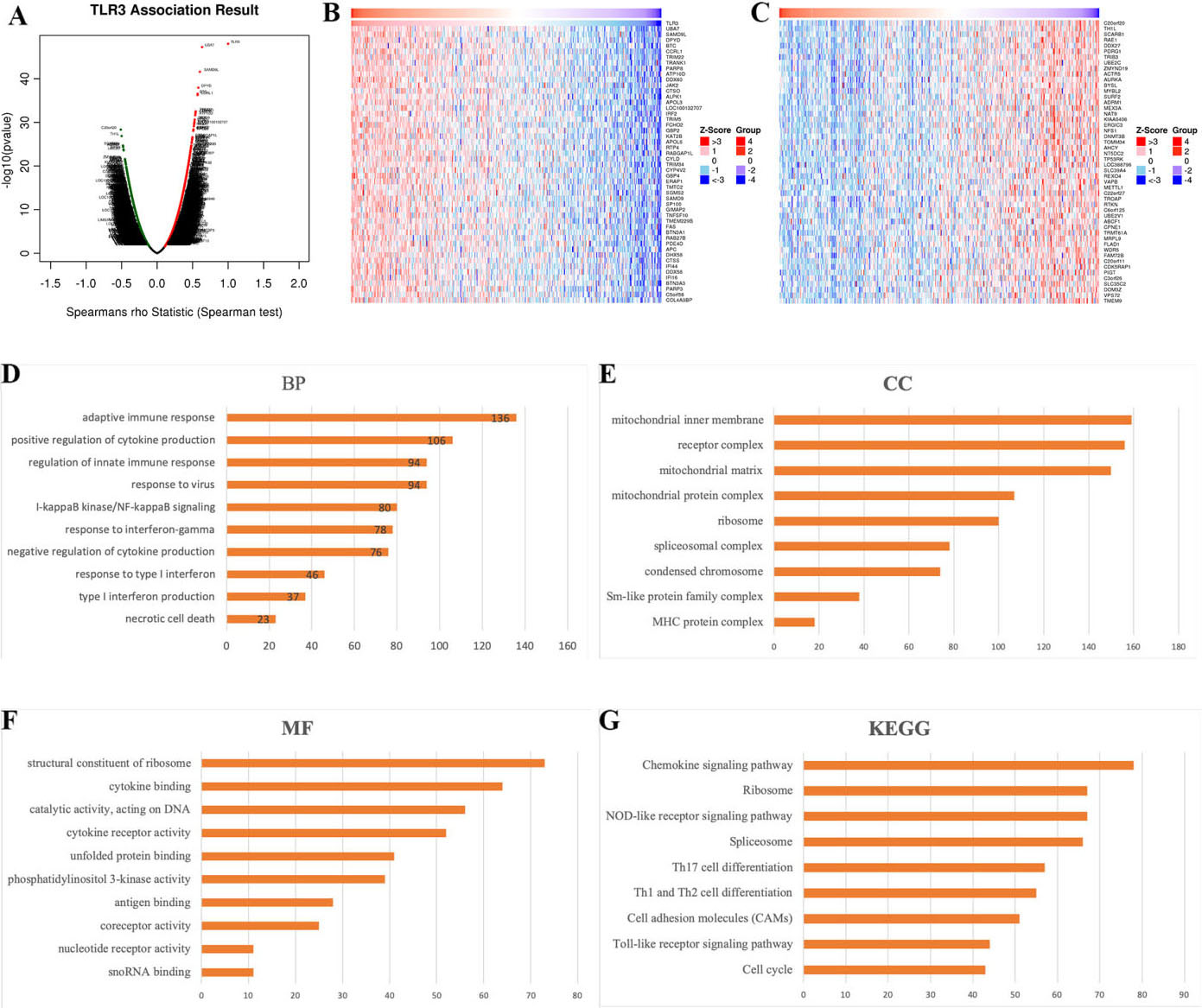
The key miRNA-target, kinase-target and transcription factor-target networks of TLR3 in STAD (LinkedOmics)
Abbreviation: TLR, toll-like receptor; STAD, stomach adenocarcinoma; FDR, false discovery rate.
PPI networks and functional analysis of the genes in the MIR-129 and PLK1 target networks. (A) The PPI networks and functional analysis of MIR-129 target networks. (B) The PPI networks and functional analysis of PLK1 target networks. Different colors of the network edge illustrate the bioinformatics methods applied: coexpression, genetic interactions, predicted, shared protein domains, physical interactions and colocalization. The different colors for the network nodes illustrate the biological functions of the set of enriched genes. Abbreviations: PPI, protein-protein interaction; MIR, microRNA.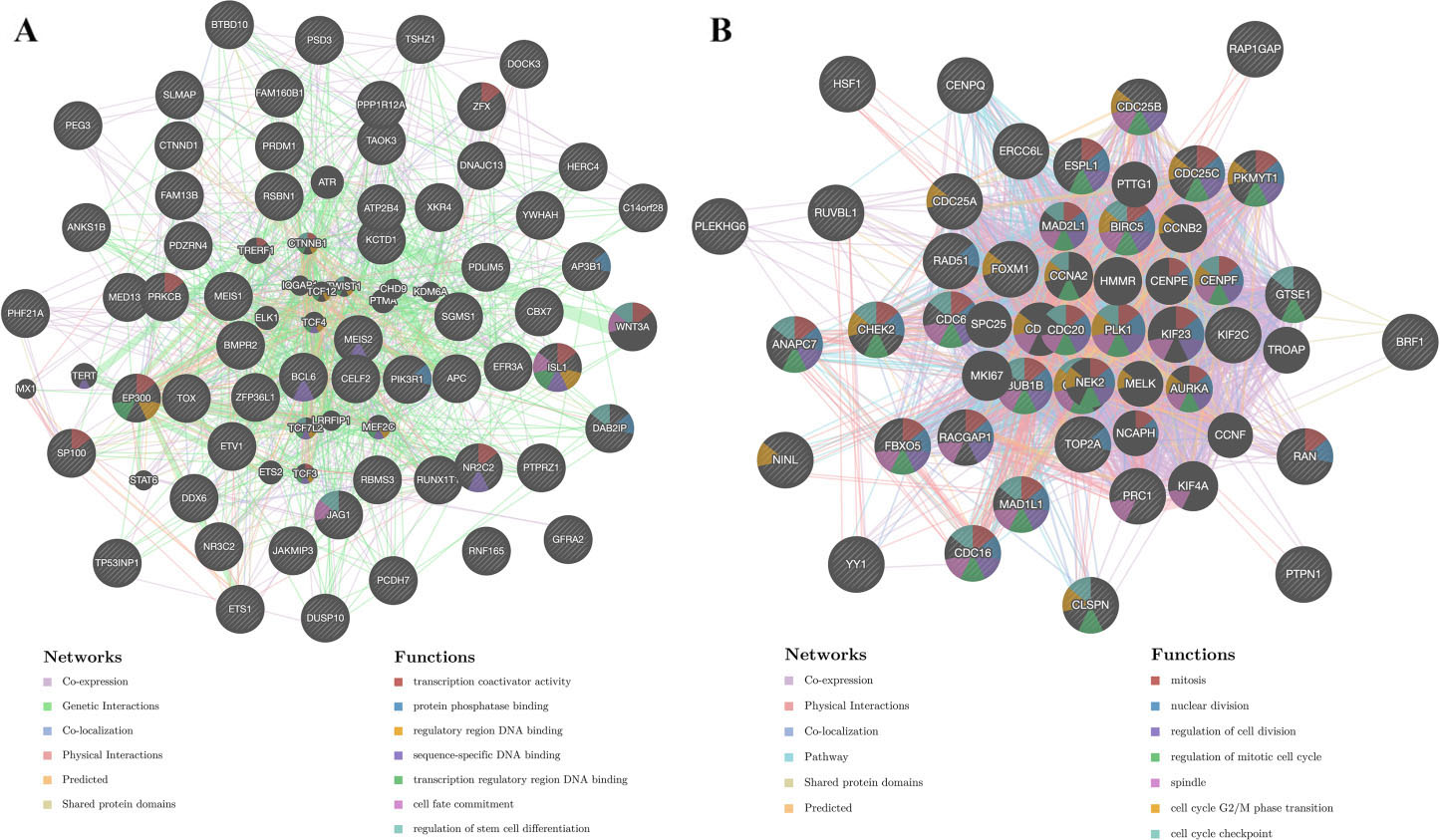
The “LinkFinder” module was used to explore the genes correlated with TLR3 in STAD samples. In total, 10322 genes (dark green dots) were negatively correlated with TLR3, while 9903 genes (dark red dots) were positively correlated with TLR3 (Fig. 8A). The top 50 genes that were positively and negatively associated with TLR3 in STAD samples are shown in Fig. 8B and C. Gene ontology (GO) analysis in the “LinkInterpreter” module demonstrated that the role of TLR3 in STAD primarily participated in the immune response, positive regulation of cytokine production, NF-KappaB signaling, response to interferon-gamma, cytokine binding, and catalytic activity of DNA (Fig. 8D–F). Moreover, KEGG pathway analysis revealed that TLR3 exerted biological functions in STAD mainly via the chemokine signaling pathway, NOD-like receptor signaling pathway, Toll-like receptor signaling pathway, and Th1, Th2 and Th17 cell differentiation (Fig. 8G).
MicroRNA, kinase and transcription factor target networks of TLR3 in STAD
To validate the targets of TLR3 in STAD, we explored the top 5 most significant microRNA (miRNA), kinase, and transcription factor target networks by GSEA in LinkedOmics (Table 3). The top 5 most significant miRNA target networks were MIR-129 (GCAAAAA), MIR-507 (GTGCAAA), MIR-202 (ATAGGAA), MIR-505 (GTGTTGA) and MIR-142-3P (ACACTAC). Similarly, the top 5 most significant kinase target networks were PLK1, LCK, JAK2, JAK3 and HCK13. Furthermore, the top 5 most significant transcription factor target networks were V$IRF_Q6, V$IRF1_01, V$IRF2_01, V$IRF3_01 and V$E2F_Q4. We next constructed protein-protein interaction networks to validate the potential biological functions of MIR-129 (GCAAAAA) target networks, PLK1 target networks and V$IRF1_01 target networks by GeneMIANA. The gene set in the MIR-129 target network mainly participated in sequence-specific DNA binding, regulatory region DNA binding and protein phosphatase binding (Fig. 9A). The gene set in the PLK1 target networks was mainly involved in the regulation of the mitotic cell cycle and cell division (Fig. 9B). The gene set in the V$IRF1_01 target network was mainly correlated with the response to type I interferon, type I interferon signaling pathway, response to interferon-gamma and interferon-gamma-mediated signaling pathway(Supplementary Fig. 2).
Discussion
TLRs, a subset of pattern-recognition receptors (PRRs), are capable of responding to conserved microbial products and endogenous damaged molecules and are involved in the biological mechanism of pathogen invasion, cancer and tissue injury [7, 33]. The TLR signaling pathway can transmit signals from inappropriate immune enhancement and antitumor immunity to modulate the progression, metastasis, recurrence and drug resistance of cancer cells [34]. Accumulating studies have been conducted to determine the clinical value, biological functions and molecular mechanisms of TLRs in various types of cancers. However, we know very little about the specific functions of TLRs in STAD.
The above analysis showed that the transcription level of TLR2 in STAD tissues was significantly elevated. However, the transcription level of TLR3 in STAD tissues was substantially decreased. Some studies have reported that TLR2 is overexpressed in gastric cancer and pancreatic cancer, and TLR2 expression is associated with pathological parameters, such as nodal metastasis stage and microvascular invasion [35, 36]. Furthermore, Bonnin et al. [37] and Yuan et al. [11] reported that TLR3 is downregulated in hepatocellular carcinoma. The above evidence is similar to the results in our study. However, more studies that focus on the expression level of TLR3 in STAD are needed.
Attention has been given to the prognostic value of TLR2/TLR3 in STAD. Unexpectedly, TLR2 seemed not to be a potential biomarker for STAD. We next found that STAD patients with high TLR3 expression had a better prognosis in terms of OS, FP and PPS than those with low TLR3 expression, which indicated that high TLR3 expression might be a protective factor for the survival of STAD patients. Evidence on hepatocellular carcinoma, neuroblastoma and thoracic esophageal squamous cell carcinoma also showed that patients with high TLR3 expression had a favorable prognosis [11, 12, 13]. Some studies have reported that activation of TLR3 induces apoptosis in prostate cancer [38], non-small cell lung cancer [39], ovarian cancer [40], and hepatocellular carcinoma [11]. Inducing apoptosis of cancer cells may be one of the mechanisms by which TLR3 improves the prognosis of STAD patients.
Tumor-infiltrating immune cells play a pivotal role in eliminating cancer cells and hindering cancer progression [41]. The abundance of tumor-infiltrating immune cells affects the proliferation and progression of tumor cells, thus influencing the clinical outcomes of cancer patients [42]. We found that the expression level of TLR3 was positively associated with the abundance of CD8
The main functions of TLR3 in STAD were associated with the immune response, positive regulation of cytokine production, NF-KappaB signaling, response to interferon-gamma, cytokine binding, and catalytic activity of DNA. TLR3 exerts functions in STAD mainly through the chemokine signaling pathway, NOD-like receptor signaling pathway, Toll-like receptor signaling pathway, and Th1, Th2 and Th17 cell differentiation. TLRs are primarily correlated with innate immunity; however, they also participate in the adaptive immune system mainly through activation of dendritic cells and regulation of cytokines, such as IL-1
We constructed PPI networks to validate the potential functions of the MIR-129 (GCAAAAA) target network, PLK1 target network and V$IRF1_01 target network. The functions of the MIR-129 (GCAAAAA) target network mainly involved the process of gene expression. Thus, TLR3 may influence the occurrence and progression of STAD by modulating the process of oncogene expression via the MIR-129 (GCAAAAA) target network. Overexpressed PLK1 causes destruction of checkpoints, which leads to excessive cell division and eventually results in the tumorigenesis and progression of cancer [48]. The functions of the PLK1 target network primarily involved the process of cell division. Hence, TLR3 may regulate the division of STAD cells by the PLK1 target network. Interferon regulatory factors (IRFs) are closely related proteins that mainly participate in regulating interferon-inducible genes in the interferon system [49]. The loss of functional interferon regulatory factors 1 (IRF1) plays a pivotal role in the development of GC [50]. The functions of the V$IRF1_01 target network was primarily responsible for the regulation of the interferon system. Thus, TLR3 may affect the occurrence and progression of STAD by regulating the interferon system via the V$IRF1_01 target network.
Some limitations still exist in the current study. First, our analyses were mainly conducted at the transcriptional level, and some of the results may not apply to studies based on the protein level. Second, relative fundamental and clinical studies that focus on the clinical value and molecular mechanisms of TLR3 in GC or STAD are rare compared to those that focus on TLR3 in other types of cancer. Thus, more studies are required to verify our results.
Conclusions
In the present study, we primarily explored the expression, the prognostic performance of TLRs in STAD, and the relationship between TLRs expression and immune infiltration. As a result, compared with normal stomach tissues, a significantly decreased mRNA level of TLR3 was detected in STAD tissues. And STAD patients in high TLR3 expression group showed a better prognosis in OS, FP and PPS. In addition, TLR3 expression was positively associated with abundance of CD8
Author contributions
Conception: Linquan Wu, Aoxiao He and Zhihao Huang.
Interpretation or analysis of data: Jiakun Wang and Hongcheng Lu.
Preparation of the manuscript: Zhihao Huang and Qian Feng.
Revision for important intellectual content: Wenjun Liao and Qian Feng.
Supervision: Rongguiyi Zhang and Xiaoyun Xu.
Supplementary data
The supplementary files are available to download from
sj-pdf-1-cbm-10.3233_CBM-210354.pdf - Supplemental material
Supplemental material, sj-pdf-1-cbm-10.3233_CBM-210354.pdf
Footnotes
Acknowledgments
This article was supported by National Natural Science Foundation of China (No. 81860431) and National Natural Science Foundation of China (No. 82060447).
Conflict of interest
All authors declare no conflicts of interest in this work.
