Abstract
Objective
To determine whether the Arg753Gln polymorphism of the toll-like receptor 2 (TLR2) gene and the Asp299Gly polymorphism of the TLR4 gene in critically ill patients affect their clinical outcomes.
Methods
Medical and surgical patients in three intensive care units (ICU) were enrolled in this prospective study. TLR2 and TLR4 gene polymorphisms were determined using restriction fragment length polymorphism analysis.
Results
A total of 145 patients were included in this study: 28 patients carried heterozygous mutations (10 in the TLR2 gene, 19 in the TLR4 gene, and one combined) and 117 patients were wild type. Severe sepsis was observed in 33% of wild types (n = 38), 60% of the TLR2 group (n = 6), and 63% of the TLR4 group (n = 12); the difference was significant between the TLR4 and wild type groups. Both TLR groups demonstrated a shorter time-to-onset of severe sepsis or septic shock. Only the TLR4 group demonstrated significant progression towards septic shock compared with the wild type group. Length of ICU stay was significantly prolonged in the TLR4 group compared with the wild type group, but not in the TLR2 group.
Conclusions
Two common SNPs of the TLR2 and TLR4 genes – Arg753Gln and Asp299Gly – were associated with a shorter time-to-onset of severe sepsis or septic shock in patients admitted to the ICU.
Keywords
Introduction
Infections remain a serious concern in patients requiring critical care. A serious complication of infection in the intensive care unit (ICU) is a severe sepsis and its maximal manifestation, septic shock.
Even if the initial clinical presentation of Gram-positive, Gram-negative and fungal sepsis is frequently similar, the host response to these pathogens is largely different and relies on innate immunity. The innate immune system represents the first line of defence in recognition of microbial structures. Reliable detection of microbial components is based on pathogen-associated molecular patterns (PAMPs) detected by pattern recognition receptors (PRRs) that warn the host about the presence of infection. 1
The toll-like receptor (TLR) family is the best characterized class of PRRs in mammalian species; at least 10 TLRs have been identified in humans as being responsible for the detection of multiple PAMPs. 2 TLR2, originally recognized as a receptor responsible for lipopolysaccharide-mediated cellular signalling, 3 is now recognized as being a receptor for bacterial lipoproteins and lipoteichoic acids. In contrast, TLR4 is identified as the ultimate receptor for the lipopolysaccharide (endotoxin) of Gram-negative organisms. 1 A different manifestation of infection in critically ill patients suggests that host genetics influence, via the modulation of the expression and function of the TLRs, the extent of the innate immune response and the severity of sepsis. 4
Various single nucleotide polymorphisms (SNPs) of the TLRs have been identified and these are common variants within a population, found at a frequency >1%. 5 Analysis of the Asp299Gly and Thr399Ile polymorphisms in the human TLR4 gene found an increased susceptibility to Gram-negative infection in a small patient group with the Asp299Gly mutation. 2 A previous study intended to evaluate the clinical outcome of TLR4 gene polymorphisms in critically ill Japanese patients, but no Japanese participants carrying the TLR4 gene Asp299Gly or Thr399Ile polymorphisms were detected. 6 The link between the Arg753Gln SNP and the GT-repeat microsatellite in the TLR2 gene and the occurrence of severe staphylococcal infection was investigated and no associations with disease or mortality were found. 7 Some evidence from animal studies, however, suggests that TLR4 function induces TLR2 expression and vice versa and that a close interaction exists between these two receptors.8,9
This present study investigated whether there is a difference in the severity and the course of sepsis in critically ill patients with the Arg753Gln SNP in the TLR2 gene or the Asp299Gly SNP in the TLR4 gene compared with wild type patients.
Patients and methods
Patient population
This prospective, controlled, noninterventional, clinical, observational study collected data during serial periods of 3 months between August 2006 and December 2008 at the Department of Anaesthesiology and Intensive Care Medicine, Charité – Universitätsmedizin Berlin, Campus Charité Mitte and Campus Virchow-Klinikum, Berlin, Germany. After an initial period of 1 year, the prevalence of TLR gene polymorphisms was lower than expected so the study proceeded with inclusion of consecutive patients for this study.
The Charité is a 3200-bed, acute-care university hospital in Berlin, Germany, providing tertiary medical treatment. Three ICUs under anaesthesiological management (one cardiosurgical, one neurosurgical and one interdisciplinary unit) with a total of 61 beds were involved in the study. Postsurgical patients and those with medical or neurological conditions were admitted to these three ICUs. Patients of both sexes ≥18 years were assigned to study inclusion following ongoing intensive care treatment of >36 h. Exclusion criteria were age <18 years and pregnancy.
This study was approved by the institutional Ethics Committee Board of Charité – Universitätsmedizin Berlin (no. EA 1/117/06) and the data safety authorities. The study was registered with the German clinical trials register (no. DRKS00003171). All patients or their next of kin signed informed consent before inclusion in the study.
Data collection
Data from eligible patients were recorded during the daily rounds from the medical records, and from the hospital mainframe computer and the patient data management system (Computer Organized Patient Report Assistant [COPRA]; COPRA System GmbH, Berlin, Germany). Data were collected every day for the previous 24-h period. Records of vital signs, laboratory findings, microbiological and radiological diagnostics, anti-infective, vasopressive and immunosuppressive agents, ventilation data, pulmonary gas exchange, urine output and fluid balance were entered into a study database (Microsoft® Access®; Microsoft Corporation, Redmond, WA, USA).
Data on immune suppression, alcohol, and drug or nicotine abuse status were taken from the patient’s data file. Immune suppression was defined as the patient being on corticosteroids or other immunosuppressive agents, having human immunodeficiency virus or leukaemia, or having had chemotherapy. Any dose of corticosteroids above the prednisolone equivalence dose of 7.5 mg daily was counted as immune suppression. Types of immune suppressive agents in use during the period of this study for transplant recipients of solid organs (e.g. liver or kidney) were tacrolimus (PROGRAF®; Astellas Pharma, Deerfield, IL, USA) and mycophenolate mofetil (CellCept®; Roche, Nutley, NJ, USA).
TLR2 and TLR4 analysis and genotyping
A 2-ml sample of peripheral venous blood was collected from eligible patients into ethylenediaminetetra-acetic acid tubes (BD Vacutainer®; Becton Dickinson, Franklin Lakes, NJ USA), which were stored in a central freezer at −20℃ and transported once a week to a central laboratory (Central Institute for Laboratory Medicine, Clinical Chemistry and Pathobiochemistry, Labor Berlin, Charité – Universitätsmedizin Berlin, Campus Virchow-Klinikum, Berlin, Germany) for processing. Genomic DNA was extracted from the peripheral venous blood with a purification kit (QIAamp® DNA Blood Mini Kit; QIAGEN, Hilden, Germany) according to the manufacturer’s protocol and stored at −20℃ until analysed. Primer design and method validation were carried by one of the authors (A.W.) for use in routine analysis in the clinical chemistry laboratory employed in this study.
Genotyping for the polymorphism R753Q (Arg753Gln) in the TLR2 gene and the polymorphism D299G (Asp299Gly) in the TLR4 gene was performed using a polymerase chain reaction (PCR) in a Biometra Trio Block thermal cycler (Biometra, Göttingen, Germany). PCR was carried out in a volume of 20 µl, with a concentration of 1 µM of the respective sense and antisense primer, 0.2 mM of each deoxynucleotide triphosphate, 1 U AmpliTaq Gold® DNA polymerase (Applied Biosystems, Foster City, CA, USA), and 35 ng of genomic DNA in 1x AmpliTaq Gold® PCR buffer (Applied Biosystems). The primers used for the TLR2 gene were: 753TLR2F, 5′-TTGACTCCATTGAAAAGAGC-3′; and 753TLR2R, 5′-TAAATATGGGAACCTAGGAC-3′ (TIB Molbiol, Berlin, Germany). For the TLR4 gene, the primers were: 299TLR4F, 5′-GGCTTCATAAGCTGACTTT-3′; and 299TLR4R, 5′-CATCCGAAATTATAAGAAAAG-3′ (TIB Molbiol). The thermal cycling conditions for both PCRs involved preliminary denaturation at 95℃ for 10 min followed by 34 cycles of denaturation at 95℃ for 20 s, annealing at 54℃ for 20 s, and elongation at 72℃ for 30 s, followed by a final elongation step at 72℃ for 5 min. The products were 300 base pairs (bp) for the Arg753Gln polymorphism and 316 bp for the Asp299Gly polymorphism. Following the PCR, restriction endonuclease digests were performed on the PCR products. For the Arg753Gln polymorphism, 10 µl of the PCR reaction were digested with 1 U PstI restriction endonuclease (New England Biolabs, Frankfurt am Main, Germany) in 1 x NEB buffer 3 at 37℃ overnight. For the Asp299Gly polymorphism, 10 µl of the PCR reaction were digested with 1 U BccI restriction endonuclease (New England Biolabs) in 1 x NEB buffer 1 at 37℃ overnight. The final volumes of the reactions were 20 µl each. The digestion of the TLR2 gene PCR product resulted in a single band of 300 bp when arginine was present at position 753 in the gene and two bands (190 and 110 bp) when glutamine was at position 753. The digestion of the TLR4 gene PCR product yielded two bands (222 and 94 bp) when asparagine was present at position 299 in the gene and three bands (113, 109, 94 bp) when glycine was at position 299. Following this, the digests were analysed by polyacrylamide gel electrophoresis on a 12% polyacrylamide gel stained with silver nitrate.
Regarding the TLR2 gene, a loss of function single nucleotide polymorphism (SNP) within the TLR2 gene results in an amino acid exchange Arg753Gln (SNP ID: rs5743708, mRNA accession: NM_003264, protein accession: NP_003255, MAF/minor allele count: A = 0.0119/26); possible functional relevance: compromised in its ability to activate NFKB).
Regarding the TLR4 gene, a loss of function SNP within the TLR4 gene results in an amino acid exchange Asp299Gly (SNP ID: rs4986790, mRNA accession: NM_138554, protein accession: NP_612564, MAF/minor allele count: G = 0.0441/96, possible functional relevance: impaired LPS-induced phosphorylation of p38 (MAPK14; 600289) and TBK1 (604834), activation of NFKB and IRF3, and induction of IL8 and IFNB mRNA expression).
Group assignment
Genotypes were assigned and confirmed by independent laboratory investigators who were unaware of the patients’ phenotypes. Patients were then divided into three groups according to the assigned polymorphism: Arg753Gln heterozygous carriers (TLR2 group), Asp299Gly heterozygous carriers (TLR4 group) and wild type carriers (wild type group). One patient carried both heterozygous Arg753Gln and Asp299Gly polymorphisms and was included in the analysis as described specifically in the Results section. No rare variants of the studied polymorphisms in the homozygote state were observed.
Outcome parameters
The primary outcome parameter was severity of sepsis (severe event), which was measured using the following surrogate markers: Acute Physiology and Chronic Health Evaluation (APACHE II) score, Simplified Acute Physiology Score (SAPS II), Sequential Organ Failure Assessment (SOFA), and Therapeutic Intervention Scoring System (TISS-28).10–13
The following secondary outcome parameters were also measured: duration of mechanical ventilation, duration of sepsis, severe sepsis, septic shock, mortality, duration of ICU stay, and time-to-onset of infection. Duration of mechanical ventilation was defined as the period the patient required mechanical ventilation support during the ICU stay with the trachea intubated (or via tracheostomy). The ICU stay was defined as the number of days a patient remained in the ICU during one hospital stay, including readmissions from other wards but not after discharge home. Infections were defined according to criteria of the US Centers for Disease Control and Prevention (Atlanta, GA, USA). 14 The following bacteria associated with potential intrinsic or acquired resistance based on local surveillance data were considered difficult-to-treat pathogens: methicillin-resistant Staphylococcus aureus, extended-spectrum beta-lactamase building bacteria, vancomycin-resistant S. aureus, and nonfermenters such as Pseudomonas species, Stenotrophomonas maltophilia, Acinetobacter baumannii, Citrobacter species, Enterobacter cloacae, Enterococcus faecium and Bacillus cereus.
The clinical conditions of systemic inflammatory response syndrome, sepsis, severe sepsis and septic shock were classified according to the American College of Chest Physicians/Society of Critical Care Medicine consensus criteria and the Surviving Sepsis Campaign guidelines. 15
Statistical analyses
Data are expressed as follows: qualitative data as absolute values and proportions (n, %), continuous data as mean ± SD or median and 25%–75% quartiles (25%, 75%) (the latter for categorical and non-normally distributed continuous variables). All numerical calculations were performed with the PASW statistical package, version 18 (SPSS Inc., Chicago, IL, USA) and SAS version 9.1 (SAS Institute, Cary, NC, USA). To test for differences between the groups (because of deviations from normality, small sample sizes and greater differences in sample sizes), the nonparametric Wilcoxon–Mann–Whitney test and Fisher’s exact χ2-test were performed. Survival data (time course of severe events) were estimated according to the Kaplan–Meier methods and compared univariately with log rank statistics. The clinical course of disease severity based on scoring systems as surrogate parameters was evaluated using a nonparametric longitudinal data analysis in a two-factorial design (first factor: group; second factor: time). First, all consecutive days following ICU admission were compared up to day 8 to include into the analysis a minimum of 50% of the specific study population having scores. Secondly, the onset day of a patient defined as the first ICU day with an infection diagnosed was taken as the starting point. To compare the changes in scores during infection, the following ICU days were included into the analysis using the same pattern as described above. This led to longitudinal data analyses of five consecutive ICU days from infection onset. In all statistical tests, a two-tailed P-value of <0.05 was considered statistically significant. Against the background of an observational study, all tests should be understood as constituting exploratory data analysis, such that no adjustments for multiple testing have been made.
Results
A total of 149 Caucasian unrelated patients were included in this study (Figure 1). Four patients were excluded from the analyses because they had ICU stays <36 h. Ten patients were heterozygous carriers of Arg753Gln (TLR2 group) and 19 patients were heterozygous carriers of Asp299Gly SNPs (TLR4 group). One patient carried both heterozygous alleles of these polymorphisms and was included in the analyses of the TLR2 and TLR4 groups as indicated specifically below. A total of 117 patients were assigned to the wild type group regarding each SNP. The genotype frequencies were consistent with the Hardy–Weinberg equilibrium and complied with those previously reported elsewhere.16,17
Flowchart for this prospective, controlled, noninterventional, clinical, observational study to determine if there is a difference in the severity and the course of sepsis in critically ill patients with the Arg753Gln polymorphism of the toll-like receptor 2 (TLR2) gene or the Asp299Gly polymorphism of the TLR4 gene compared with wild type. One patient carried both heterozygous alleles of the Arg753Gln and Asp299Gly polymorphisms and was excluded in the separate analyses of the TLR2 versus TLR4 group to validate the results.
Clinical characteristics of patients admitted to the intensive care unit who were included in this prospective, observational study to determine if there is a difference in the severity and the course of sepsis in critically ill patients with the Arg753Gln polymorphism of the toll-like receptor 2 (TLR2) gene or the Asp299Gly polymorphism of the TLR4 gene compared with wild type control patients.
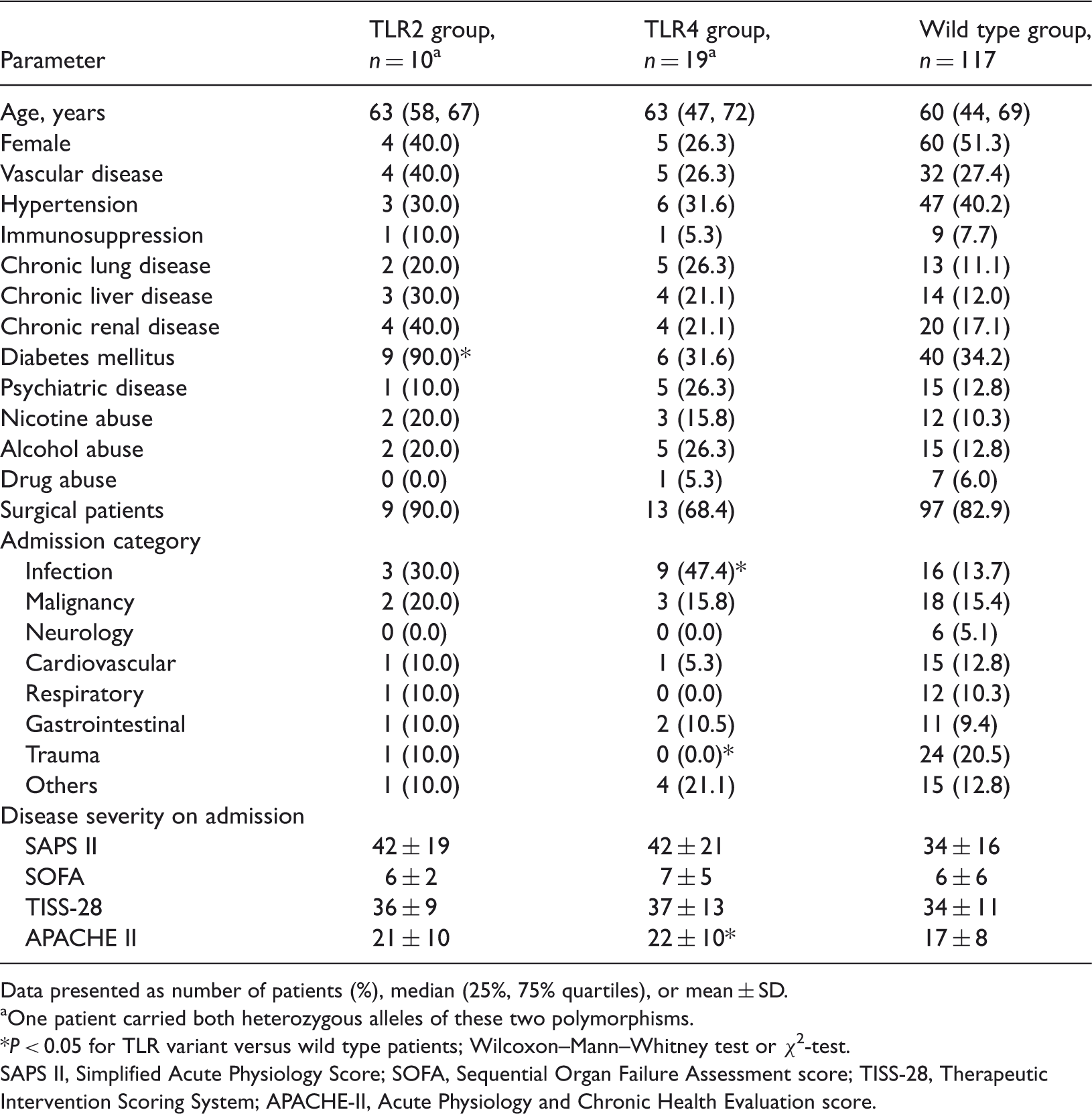
Data presented as number of patients (%), median (25%, 75% quartiles), or mean ± SD.
One patient carried both heterozygous alleles of these two polymorphisms.
P < 0.05 for TLR variant versus wild type patients; Wilcoxon–Mann–Whitney test or χ2-test.
SAPS II, Simplified Acute Physiology Score; SOFA, Sequential Organ Failure Assessment score; TISS-28, Therapeutic Intervention Scoring System; APACHE-II, Acute Physiology and Chronic Health Evaluation score.
Distribution of infections and infectious conditions in patients admitted to the intensive care unit who were included in this prospective, observational study to determine if there is a difference in the severity and the course of sepsis in critically ill patients with the Arg753Gln polymorphism of the TLR2 gene or the Asp299Gly polymorphism of the TLR4 gene compared with wild type control patients.

Data presented as number of patients (%) or mean ± SD.
One patient carried both heterozygous alleles of these two polymorphisms.
P < 0.05 for TLR variant versus wild type patients; Wilcoxon–Mann–Whitney test or χ2-test.
SIRS, severe inflammatory response syndrome.
A greater proportion of patients in the TLR2 group experienced sepsis at some stage of their stay in the ICU compared with the wild type group (P = 0.03) and the TLR4 group (Table 2). In both TLR groups, the progression of sepsis to septic shock occurred more frequently than in wild type patients, affecting 52.6% (n = 10) in the TLR4 group (P = 0.014) but only 40.0% (n = 4) in the TLR2 group.
Analysis of the changes in the APACHE II scores during the first 8 days in the ICU revealed stable higher baseline scores in the TLR2 group compared with the wild type group (P < 0.001) (Figure 2a). However, when the four consecutive days after infection onset were compared, the APACHE II scores in these two groups reached the same baseline level by day 4 (Figure 2b). The APACHE II scores in the TLR4 group showed significantly higher baseline scores, which progressed to higher values at day 8 compared with the wild type group (P = 0.024) (Figure 3a). Similarly, after infection onset in the TLR4 group, the scores further progressed until day 4 compared with the wild type group (P = 0.012) (Figure 3b).
Changes in the Acute Physiology and Chronic Health Evaluation (APACHE II) scores in patients with the Arg753Gln polymorphism of the toll-like receptor 2 (TLR2) gene (TLR2 group; n = 10) during the intensive care unit (ICU) stay (a) and at sepsis onset (b) compared with the wild type control patients. Nonparametric longitudinal data analysis based on the factors of group and time. Changes in the Acute Physiology and Chronic Health Evaluation (APACHE II) scores in patients with the Asp299Gly polymorphism of the toll-like receptor 4 (TLR4) gene (TLR4 group; n = 19) during the intensive care unit (ICU) stay (a) and at sepsis onset (b) compared with the wild type control patients. Nonparametric longitudinal data analysis based on the factors of group and time.

Kaplan–Meier analysis of the time-to-onset of a severe event (i.e. time-to-onset of severe sepsis or septic shock) in patients with either TLR polymorphism (n = 28) demonstrated a significantly shorter time-to-onset of severe sepsis or septic shock compared with the wild type group (n = 117) (P = 0.004; log rank test; Figure 4). When the Kaplan–Meier analysis of the time-to-onset of a severe-event was undertaken for each TLR group compared with the wild type group (Figures 5 and 6), both groups demonstrated a shorter time-to-onset of severe sepsis or septic shock, although only the TLR4 group demonstrated a significant decrease compared with the wild type group (P = 0.003; log rank test). Since one patient carried both TLR polymorphisms, the Kaplan–Meier analyses for the two TLR groups were also undertaken by excluding this one patient. The findings were the same as those obtained when the data from this patient were included in both groups (data not shown). A Cox proportional hazards model using TLR status, age, sex and metabolic diseases as covariates was applied to the data and no significant time-dependent interaction was found for TLR status, fulfilling the proportional hazard condition. Thus, in this present study the effect of the TLR gene status over time remained constant.
Kaplan–Meier analysis of the time-to-onset of a severe event from admission to the intensive care unit (ICU) for both toll-like receptor (TLR2 and TLR4) polymorphism groups (n = 28) compared with the wild type control patients (n = 117) (P = 0.004; log rank test). Kaplan–Meier analysis of the time-to-onset of a severe event from admission to the intensive care unit (ICU) for patients with the toll-like receptor 2 (TLR2) polymorphism (n = 10) compared with the wild type control patients (n = 117). Kaplan–Meier analysis of the time-to-onset of a severe event from admission to the intensive care unit (ICU) for patients with the toll-like receptor 4 (TLR4) gene polymorphism (n = 19) compared with the wild type control patients (n = 117) (P = 0.003; log rank test).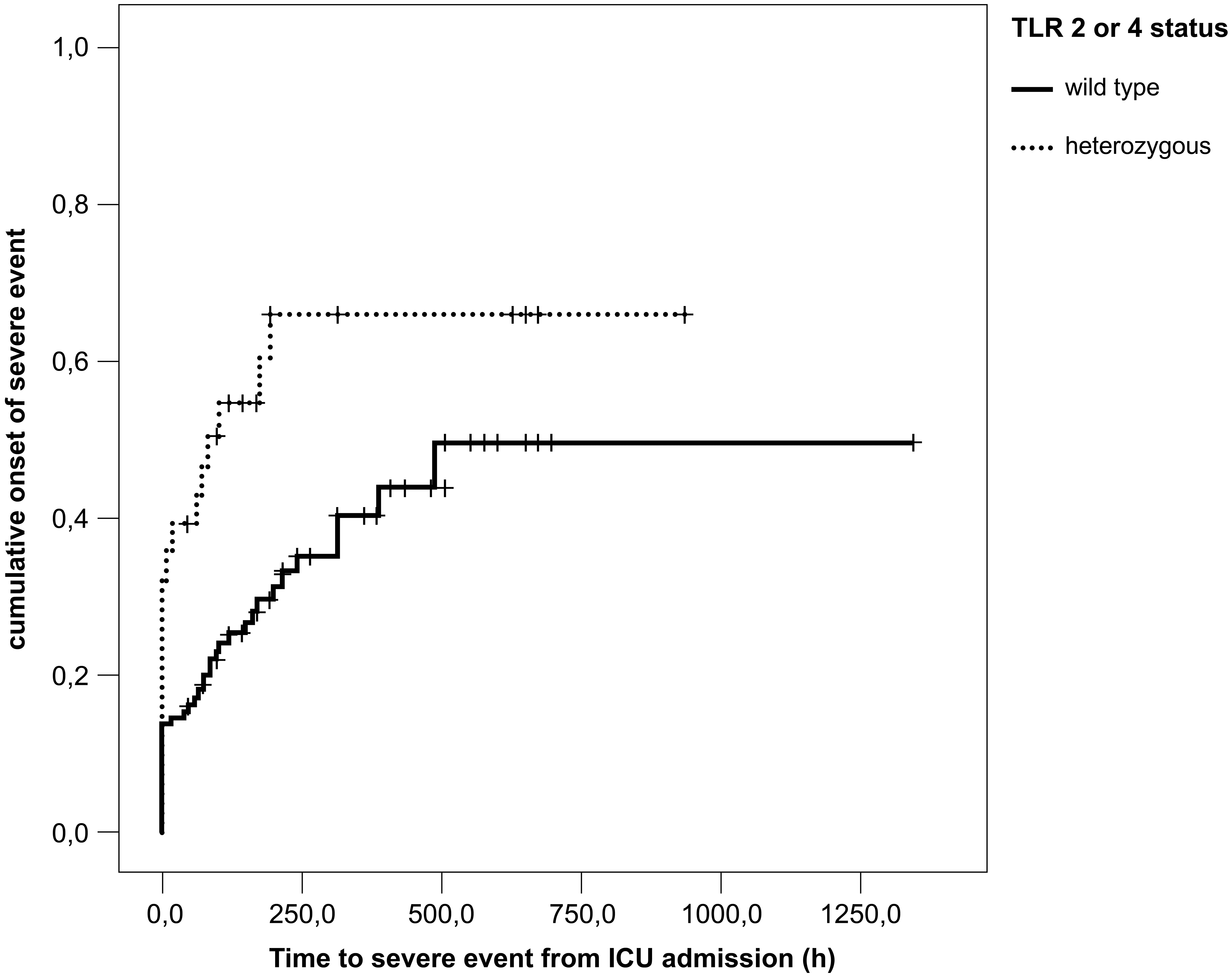


There was no significant difference in the rate of mortality between the three groups. One patient died in the TLR2 group (10.0%) and three in the TLR4 group (15.8%) compared with eight patients (6.8%) in the wild type group. The length of ICU stay was significantly prolonged in the TLR4 group compared with the wild type group (median [25%, 75%]: 24 [7, 40] versus 10 [5, 23] days, respectively; P = 0.047), but there was no significant difference between the TLR2 and wild type groups (median [25%, 75%]: 23 [8, 28] versus 10 [5, 23] days, respectively). The duration of mechanical ventilation was significantly longer in the TLR4 group compared with the wild type group (mean ± SD: 18.8 ± 22.1 versus 6.3 ± 11.6 days, respectively; P = 0.027), but there was no significant difference between the TLR2 and wild type groups (mean ± SD: 10.6 ± 11.5 days versus 6.3 ± 11.6 days, respectively).
Discussion
The most important finding of this present study was that the presence of two common SNPs of the TLR2 and TLR4 genes – Arg753Gln and Asp299Gly – was associated with a shorter time-to-onset of severe sepsis or septic shock in medical and surgical patients admitted to the ICU. Previous research has demonstrated that genetic variations influence the frequency and alter the course of infectious conditions. For example, a study of 91 patients with septic shock and 73 healthy control subjects found that mutations in the TLR4 gene were associated with an increased risk for septic shock and a higher incidence of Gram-negative bacterial infections. 2 Kiechl et al. 18 suggested that the diminished innate immune defence observed in subjects with the Asp299Gly TLR4 gene polymorphism was a result of reduced intravascular inflammation and results in the enhanced risk of severe bacterial infections. Moreover, functional cross-talk between the different TLRs has been shown. 19
A recent study that examined critically ill patients with the TLR4 wild type, TLR4 SNP (for Asp299Gly and Thr399Ile) and TIRAP/Mal SNP alleles found that severe sepsis and septic shock occurred in six out of nine patients with ventilator-associated pneumonia who were carriers of the combination of the TLR4 and TIRAP/Mal SNPs. 20 The authors postulated that a lack of TLR4 might be associated with a worse outcome, 20 which was in accordance with the findings of this present study. Controversially, another study observed no correlation between TLR2 and TLR4 gene SNPs and the development and outcome of sepsis in surgical critically ill patients. 21
One animal study suggested that the TLR pathway might be involved in the stress hormone axis because the results showed that steroidogenesis was altered in TLR4-deficient mice in response to lipopolysaccharide challenge. 22 This type of altered stress response might provide one explanation for the different severity of sepsis. In contrast, another study that investigated whether there is a difference in cytokine production as sepsis progresses, found similar levels of TLR2 and TLR4 proteins on monocytes both in septic patients and in control subjects. 23
Another explanation relies on the capability of TLR4 proteins to transduce proinflammatory signals produced by various types of reactive oxygen metabolites. For example, several studies have demonstrated in various settings that TLR4 proteins might regulate intracellular signalling pathways in this way, independently of lipopolysaccharide activation.24–26 Patients lacking receptor activity, as in the case of those with SNPs, might not exhibit these proinflammatory characteristics.
The incidence of Gram-negative infection with polymicrobial infections increases depending on the genotype of the patient; being 100% in patients with the TLR4 gene Asp299Gly allele, compared with 50% in wild types and 33.3% in patients carrying the TLR4 gene Asp299Gly/Thr399Ile allele. 2 In a study that investigated patients with TLR4/CD14 gene SNPs in surgical ICUs compared with healthy volunteers, 14 of 77 patients with TLR4-deficient alleles showed a significantly higher rate of Gram-negative infection. 27 In line with this present study, Kumpf et al. 20 found no association between the TLR4 gene 299/399 polymorphisms and the incidence or type of microorganisms in surgical infections.
The number of infections per patient did not differ significantly between the TLR4 and wild type groups in the present study. A study of patients after major visceral surgery found no correlation between the TLR4 gene Asp299Gly polymorphism and the development of sepsis, cytokine level at onset and the outcome of sepsis. 28 A previously published study identified a trend toward a higher rate of infections at admission in patients with TLR4 gene SNPs. 21 This present study found no difference in the distribution of abdominal infections between all three groups. Different susceptibility to bacterial infections has been shown to be dependent on the presence of the Thr399Ile SNP of TLR4. 18
In contrast, the TLR2 group in the present study exhibited a significantly higher number of infections per patient and incidence of difficult-to-treat pathogens compared with the wild type group. This finding was not surprising because the TLR2 gene is highly expressed in respiratory epithelium 29 and pulmonary infections represented a major source of infection in the patients in this present study. Research on normal human bronchial epithelial cells showed that stimulation of TLR2 proteins led to a higher activation of the epidermal growth factor receptor cascade, resulting in interleukin-8 and vascular endothelial growth factor production. 30 It is possible that the recruitment and stimulation of neutrophils, as well as the repair of injured tissue, are insufficient in patients with TLR2 gene SNPs.
A recent case–control study found no association between various TLR2 gene polymorphisms and outcomes in severe staphylococcal disease. 7 In an in vitro study, the heterozygous TLR2 alleles produced a full cytokine response to S. aureus ligand. 31 However, it should be noted that Moore et al. 7 included a large percentage of patients with community-acquired staphylococcal infections. The present study was mainly affected by difficult-to-treat nosocomial infections in this patient group, experiencing species other than S. aureus (i.e. the present setting was not restricted to staphylococcal infections). It should be noted that TLR2 is not the only molecule responsible for the recognition of Gram-positive bacterial structures and TLR4, as other members of TLR family and their associated mediators are at least partly involved in signal cascade. 32 Tlr2 gene knockout mice are still highly susceptible to S. aureus infection, suggesting that other pathways might have been implicated in staphylococcal infections. 33 For example, TLR4 but not TLR2 is essential for the recognition of lipoteichoic acid, the main ligand from Gram-positive bacteria, in mice. 33
In the present study, patients in the TLR2 group were significantly more likely to have diabetes mellitus compared with the wild type group. TLR2 gene SNPs were recently associated with the development of type 1 diabetes. 34 Another study demonstrated higher levels of TLR2 on the cell surface of monocytes from patients with type 1 diabetes compared with healthy control subjects. 35 An animal study showed that increased TLR2 signalling contributes to the prolonged inflammation observed in the wounds of mice with experimental diabetes. 36 Currently, it remains unclear how TLR2 gene SNPs might result in a higher occurrence of diabetes, which might, in turn, be responsible for a significantly higher rate of infections per patient in the TLR2 group in the present study.
In the present study, the highest mortality rate was seen in the TLR4 group, followed by the TLR2 group, and the lowest rate was observed in the wild type group, but the differences were not significant. However, this was not the primary aim of the present study because mortality in ICU patients, particularly in those with sepsis, is affected by a variety of factors, so the mortality data should be interpreted cautiously. Both the duration of mechanical ventilation and the length of ICU stay were prolonged in patients with TLR gene polymorphisms in the present study, reaching statistical significance only in the TLR4 group compared with the wild type group. A previously published study demonstrated that elevated TLR2 and TLR4 mRNA levels were associated with a longer duration of mechanical ventilation and a longer duration of ICU stay in paediatric patients. 37 It should be noted that in the present study the severity of disease was significantly greater, as measured by the APACHE II score, in patients in the TLR4 group at baseline compared with the wild type group, which might have impacted negatively on the subsequent clinical outcomes of this group.
This present study had a number of limitations. First, as for most studies on genetic polymorphisms, this present study only included a relatively small number of patients due to the rarity of the TLR2 and TLR4 gene SNPs in the general population. This restricts the conclusions that can be drawn from the available data. Secondly, this present study did not investigate the downstream signalling pathways, which could provide additional information on the pathogenesis of sepsis evolution in patients with TLR gene polymorphisms. Finally, several mediators are known to modulate signalling through the TLR cascade, including opioids, proton-pump inhibitors, statins, vitamin D complex and other widely used medications. 38 The impact that the concomitant administration of these agents might have in patients with sepsis who have different TLR gene polymorphisms remains unknown.
In conclusion, the present study demonstrated that the presence of two SNPs of the TLR2 and TLR4 genes – Arg753Gln and Asp299Gly – were associated with a shorter time-to-onset of severe sepsis or septic shock in medical and surgical patients admitted to the ICU.
Footnotes
Acknowledgements
We specially thank Drs Tanja Trefzer, Dionisios Mavrodis, Tanja Sawatzki and Daniela Pfaff for their enormous help in the recruitment of patients for this study. We also thank Dr Felix Balzer and our team of medical students: their support in data acquisition was invaluable.
Declaration of conflicting interests
A.T. received funding unrelated to this study from SIRS-Lab GmbH and Orion Pharma GmbH. I.N. and S.T. received funding unrelated to this study from Roche GmbH. C.S. received funding unrelated to this study from Abbott, Aspect, Baxter, BBraun, Essex Pharma, Fresenius Kabi, GSK, and Wyeth. A.W., S.H., A.R. and K.D.W. declare that they have no conflicts of interest.
Funding
This study was funded from internal university grants from Charité – Universitätsmedizin Berlin and received no specific grant from any funding agency in the public, commercial or not-for-profit sectors.
