Abstract
BACKGROUND:
Head and neck squamous cell carcinoma (HNSC) is one of the most common cancer types in the world. The study in molecular markers for HNSC prognosis is of great significance. We hypothesized that Aminoacyl tRNA Synthetase Complex Interacting Multifunctional Protein 1 (AIMP1), a gene that encodes a cytokine, is a critical biomarker for HNSC.
METHODS:
We acquired clinical data, mRNA expression data, protein staining data, and single-cell expression data of HNSC from open databases and evaluated the clinical prognostic value of AIMP1, and explored the potential roles of AIMP1 in HNSC biology and tumor immune microenvironment.
RESULTS:
AIMP1 was overexpressed in HNSC compared to normal tissues. Higher AIMP1 expression was associated with a worse survival rate. A survival nomogram was constructed for HNSC patients. One thousand two hundred and eighty-one genes were identified as positively associated with AIMP1 and enriched in proliferation-related terminologies, while 303 genes were identified as negatively associated with AIMP1 and enriched in terminologies related to skin development and immune cell regulation. AIMP1 was positively correlated with stemness, cell cycle, and DNA repair, and negatively correlated with angiogenesis, quiescence, metastasis, hypoxia, inflammation, EMT, DNA damage, and invasion in single cells. AIMP1 was expressed higher in malignant cells than immune cells and there was no difference in AIMP1 expression among immune cell types. AIMP1 high group had a lower immune score, stroma score, and microenvironment score.
CONCLUSION:
AIMP1 is a potential diagnostic and prognostic biomarker for HNSC patients and can potentially affect the proliferation and tumor immune microenvironment of HNSC cells. This study provided a novel molecular marker for the improvement of clinical HNSC treatment.
Introduction
Head and neck squamous cell carcinoma (HNSC) is one of the most common cancer types in the world. Each year, more than five hundred thousand new HNSC cases occur, which lead to about 3 hundred thousand deaths worldwide [1]. Traditional treatment methods for HNSC include tumor surgical treatment, radiotherapy, and chemotherapy, but the therapeutic effect of these therapies is not ideal [2]. Although in recent years, researchers have developed several novel methods to improve the therapy of HNSC, these efforts failed to improve the overall survival rate of HNSC patients over the past several years [2]. One of the most challenging issues in HNSC treatment is the late diagnosis of the disease, in which cases most patients have occurred lymph node metastasis and more recurrences that result in a high mortality rate of patients [3]. Therefore, the study in molecular markers that can improve the reliability and effectiveness of HNSC diagnosis and prognosis is of great significance. Understanding the role of these genes in the biological process of HNSC can also facilitate the development of HNSC therapy.
Aminoacyl tRNA Synthetase Complex Interacting Multifunctional Protein 1 (AIMP1) is a gene that encodes a cytokine that is specifically induced by apoptosis [4]. AIMP1 protein is a member of the small inducible cytokine subfamily E and can regulate wound healing, inflammation, and angiogenesis [5, 6, 7]. The precursor protein of AIMP1 is identical to the p43 protein, which is involved in the multi-tRNA synthetase complex and the modulation of aminoacylation activity of tRNA synthetase [8, 9, 10]. These indicated that AIMP1 was involved in the expression of proteins.
Recent studies revealed additional roles of AIMP1 in cancer cells. Studies showed that the release of the AIMP1 cytokine renders the tumor-associated vasculature sensitive to tumor necrosis factor [5]. AIMP1 has also been reported to regulate multiple immune cells in cancers, including NK cells, Macrophages, T-cells, etc. [11, 12, 13]. A study has suggested that AIMP1 is an exogenous anti-tumor agent in a lung cancer xenograft model [14]. As for HNSC, a mass spectrometric analysis revealed that AIMP1 can bind to fascin actin-bundling protein 1 (FSCN1), a critical protein for squamous cell carcinoma which enhances cell proliferation, migration, and invasion [15]. Although the roles of AIMP1 in cancers have not been clearly identified, the expression and the presence of AIMP1 in cancer can be potentially valuable for clinical cancer treatment. AIMP1 has been previously identified as one of the prognostic signatures that might be associated with prognosis and might promote cell proliferation and tumor progression [16]. However, so far, the clinical application of AIMP1 in cancers remains lacking. The exact role of AIMP1 in HNSC required further studies.
Here we identified AIMP1 as a critical biomarker for clinical HNSC. In this study, we tested AIMP1 as a prognostic biomarker for HNSC patients. We also proposed that AIMP1 is critical for the cancer biology and tumor immune microenvironment of HNSC cells. Thus, we evaluated the clinical prognostic value of AIMP1 and explored the potential roles of AIMP1 in HNSC biology and tumor immune microenvironment. This study provided a novel molecular marker for the improvement of clinical HNSC treatment.
Methods
RNA-seq and clinical data acquisition
Clinical and genomic data of HNSC were downloaded from The Cancer Genome Atlas (TCGA,
RNA-seq data analysis
All the analysis methods and R package were implemented by R foundation for statistical computing (2020) version 4.0.3 and ggplot2 (v3.3.2). The xCell algorithms were used to estimate the immune cell infiltration levels [17]. A
Multiple data overexpression analysis
Multiple data sets of the mRNA overexpression and DNA copy number gain of AIMP1 in HNSC were accessed and analyzed using the Oncomine [18].
AIMP1 associated genes enrichment analysis
Differentially expressed genes between AIMP1 high (75–100%) and low (0–25%) groups were identified. The cutoff values of 1.5 and 0.001 were set for fold change and
Immunohistochemistry staining
Immunohistochemistry staining images of AIMP1 in normal squamous epithelial tissues, normal lymph nodes, and HNSC tissues were accessed from Human Protein Atlas (HPA,
Single-cell sequencing data acquisition and analysis
Correlations between AIMP1 expression and functional states in HNSC single-cell datasets GSE103322 were accessed and analyzed using the CancerSEA (
Immune cell infiltration analysis
The immune cell infiltration level was calculated using the TCGA cohort. We calculated the immune cell infiltration score of TCGA data using the xCell algorithms. The immune cell infiltration levels between AIMP1 high (75–100%) and low (0–25%) groups were compared.
Statistical analysis
T-test and one-way ANOVA were applied to compare gene expression differences. Chi-Squared tests were used to compare the distribution of HNSC patients with different clinicopathological variables in the AIMP1 high and low groups. Kaplan-Meier analysis, log-rank test, and multivariate cox regression test were used to conduct survival analysis. Pearson’s correlation test was conducted to evaluate the correlation of two variables. A
Results
AIMP1 was overexpressed in HNSC
In this study, we investigated the mRNA expression of AIMP1 genes in HNSC. Results showed that in both the non-paired and paired comparison, the expression of AIMP1 was significantly higher in HNSC than in normal tissues (Fig. 1A and B). We also calculate the diagnostic ROC, results showed that the AUC of AIMP1 diagnostic ROC was 0.759, indicating that AIMP1 was a promising diagnostic biomarker for HNSC. To validated the overexpression of AIMP1 in HNSC, we also compared the mRNA overexpression or DNA copy number gain of AIMP1 in HNSC in 11 data sets using the Oncomine. Results showed that AIMP1 was significantly overexpressed or obtain copy number gain in HNSC compared to normal tissues among all of these data set (Fig. 1D). To investigate whether AIMP1 was overexpressed in HNSC compared to normal tissue at the protein levels, we compared immunohistochemistry staining images of AIMP1 in normal squamous epithelial tissues and HNSC tissues from Human Protein Atlas for reference (Fig. 2). To avoid false-positive resulted from inappropriate antibodies, we compared staining data of two different AIMP1 antibodies. The stainings of both of the antibodies showed that the staining intensity of AIMP1 in normal oral squamous epithelial tissues and normal esophagus squamous epithelial tissues were between weak to moderate (Fig. 2B1–B2, C1–C2), whereas the staining intensity of AIMP1 in HNSC tissues was between moderate to strong (Fig. 2A1–A2).
The overexpression of AIMP1 in HNSC. A. Unpaired comparison of expression of AIMP1 in HNSC and normal tissues. B. Paired comparison of expression of AIMP1 in HNSC and normal tissues. C. Diagnostic ROC curve of AIMP1 for HNSC. TCGA data were plotted. D. The mRNA overexpression or DNA copy number gain of AIMP1 in HNSC. Multiple data sets were accessed and analyzed using the Oncomine. Heatmap and overall copy number difference analysis between HNSC and normal tissues. The references of the data were listed in the legend.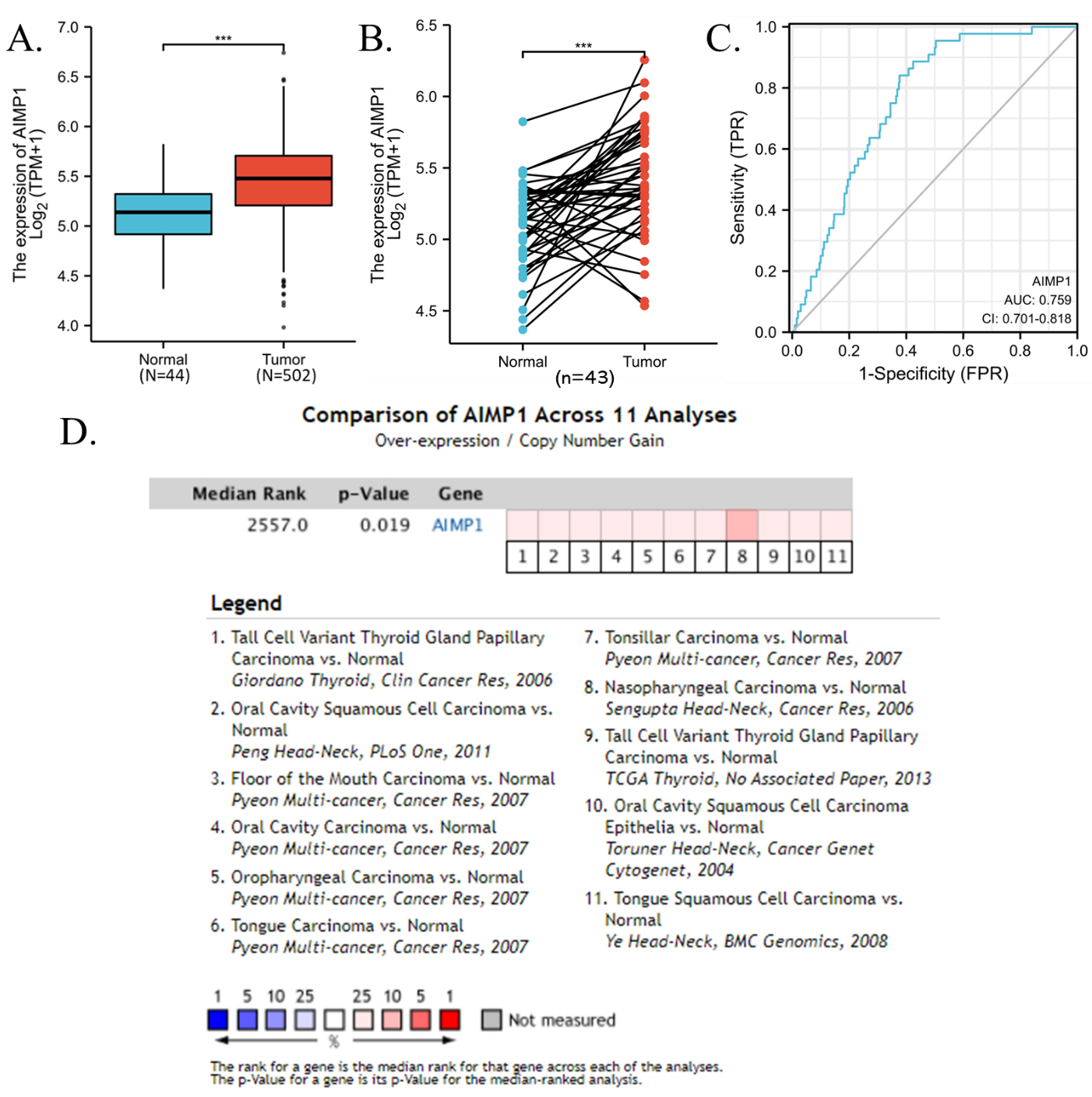
Representative immunohistochemistry staining images of AIMP1 from Human Protein Atlas. Antibody HPA018476 (A1B1C1) and CAB017618 (A2B2C2) were used to stain AIMP1. A. HNSC tissues, B. normal oral squamous epithelial tissues. C. normal esophagus squamous epithelial tissues. Sample information and histochemistry results were shown on the right side of each image.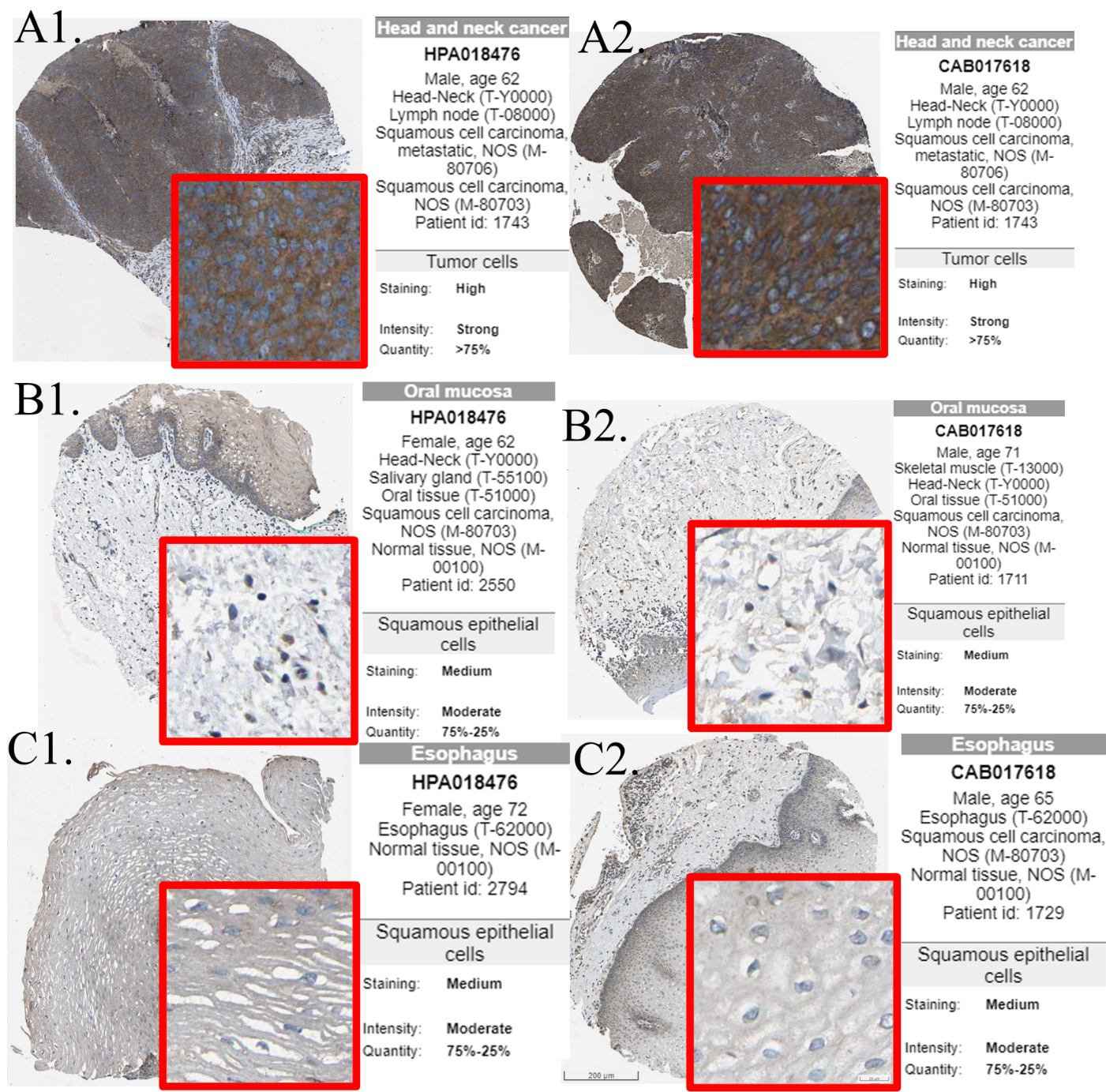
In order to further explore the association between AIMP1 expression and clinicopathological features in HNSC, multiple groups of analyses were performed within the tumor. In this study, we found that grade III HNSC expressed significantly higher AIMP1 than grade I HNSC (Fig. 3A1). Although there was no significant difference between grade I and II or grade II and III, we can see a general trend that AIMP1 expressed higher in higher grade HNSC (Fig. 3A1). In addition, we also compared different pTNM stages, different pT stages, different pN stages, gender, different races, whether smoke, whether drink, whether with radiotherapy and whether HPV positive, and we found no significance in all these comparisons except for a slight difference between pTNM I and TNM IV (Fig. 3A2–9). We also plotted a Sankey diagram for AIMP1 expression and clinicopathological features of HNSC patients to present the basic clinical information of the patients included in this study (Fig. 3B). Detailed distribution data of AMIP1 expression median divided groups were presented in Table 1.
The association between AIMP1 expression and clinicopathological features in HNSC. TCGA data were plotted. A1–10. AIMP1 distribution in HNSC and different sub-groups (A1. tumor grade, A2. pTNM stage, A3. pT stage, A4. pN stage, A5. gender, A6. race, A7. smoke, A8. drink, A9. radiotherapy, A10. HPV status) of HNSC. B. Sanguini diagram of age, gender, race, AIMP1, and survival.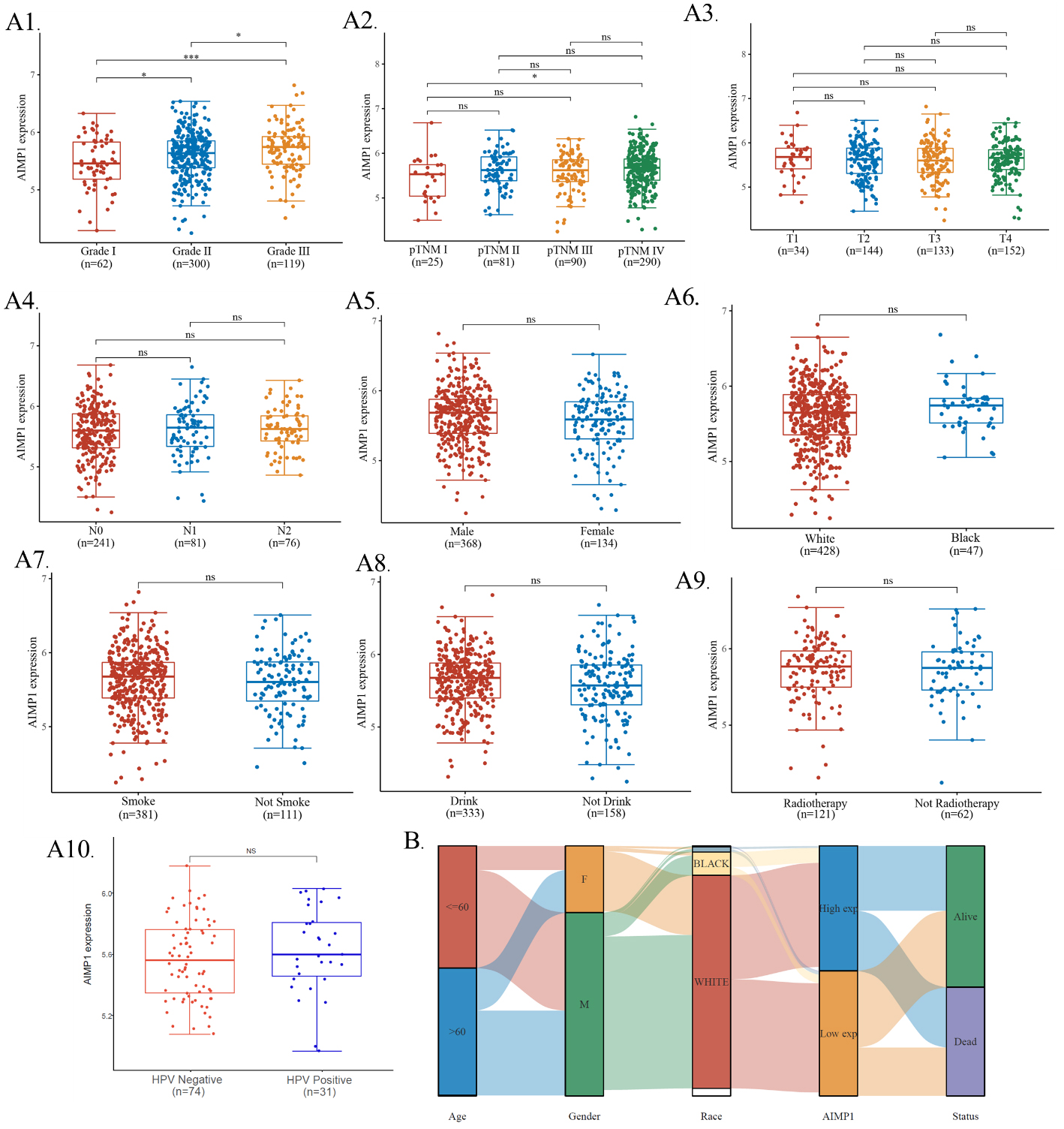
Distribution of HNSC patients with different clinicopathological variables in AIMP1 high and low groups
To explore the prognostic power of AIMP1 for clinical HNSC patients, we analyzed the association of AIMP1 expression and the overall survival of HNSC patients. Results show that there was significance between the high AIMP1 group and the low AIMP1 group (Fig. 4A). To further evaluate the prognostic value of AIMP1 for HNSC patients, we calculated the time-dependent receiver operating characteristic (ROC) analysis of the AIMP1 for overall survival prediction. Results showed that the area under the curve (AUC) for 3-, 5-, 10-year overall survival prediction were 0.651, 0.653, and 0.756 respectively (Fig. 4B). Therefore, we suggested that AIMP1 was a promising predictive factor for the overall survival of HNSC, especially for long-term prognosis. To apply the AIMP1 expression to the clinical prognosis of HNSC, we constructed a survival nomogram. Variables pT_stage, pN_stage, pTNM_stage, AIMP1 expression, gender, age, race, smoker, alcohol history, radiation therapy were taken into account (Fig. 4C). Detailed Cox regression results were displayed in Table 2. The prediction results of the nomogram calibration curves of 1-, 5-, 10-year overall survival were consistent with all patients’ observation results (Fig. 4D).
Cox regression analysis of AIMP1 in HNSC
Cox regression analysis of AIMP1 in HNSC
Prognostic analysis of AIMP1 gene in HNSC. A. Kaplan-Meier survival analysis of High (50–100%) and low (0–50%) AIMP1 groups. log-rank test p-value was shown. B. Time-dependent receiver operating characteristic (ROC) analysis of the AIMP1 overall survival prediction. The area under the curve (AUC) and 95% confidential interval (95% CI) were shown. C. Nomogram for prediction of 1-, 5- and 10-year survival. D. Calibration plots of the nomogram.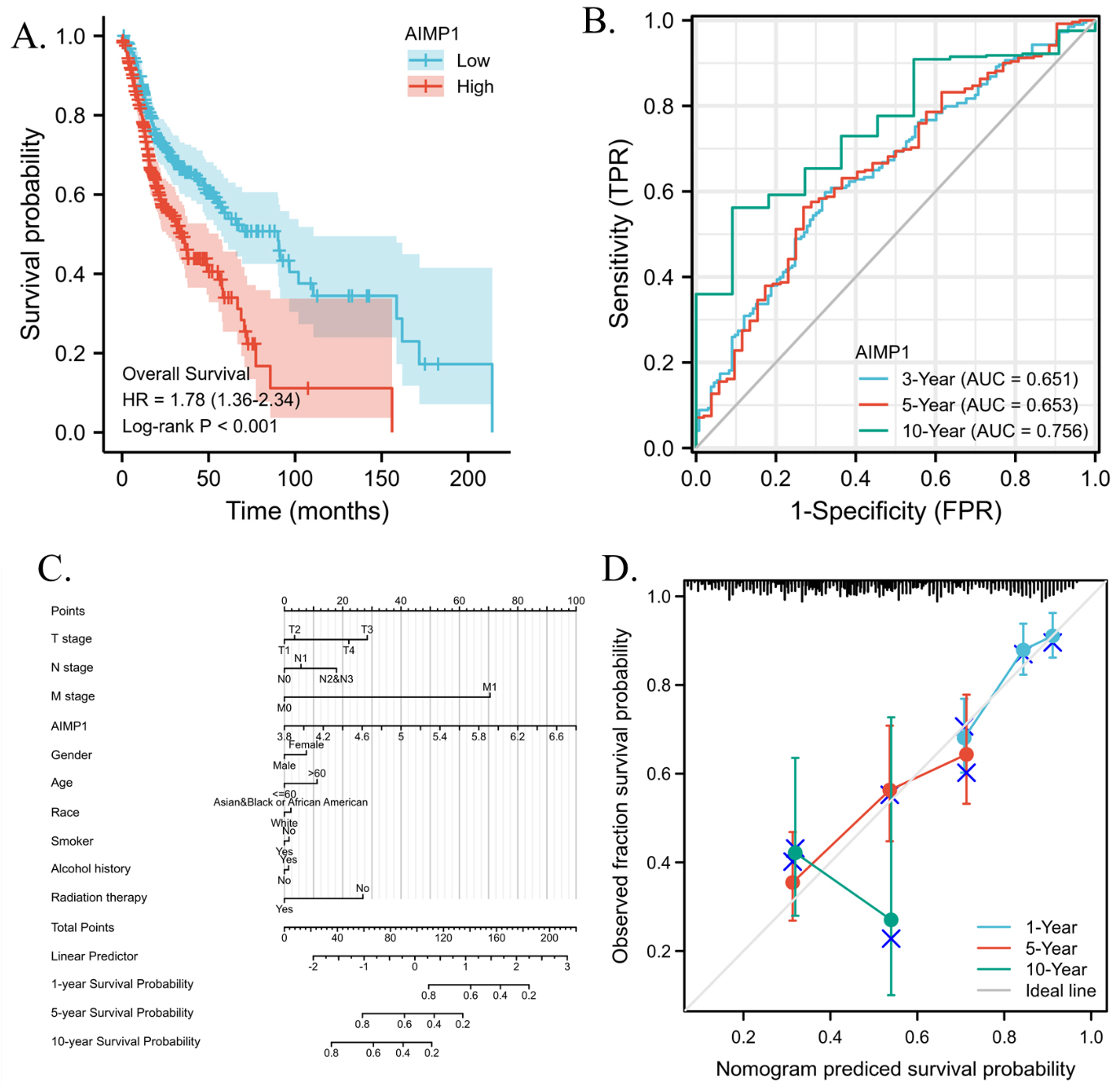
AIMP1 associated genes enrichment analysis. A. Differential gene analysis in HNSC AIMP1 high (75–100%) and low (0–25%) groups. Left panel: Volcano plots were constructed using fold-change 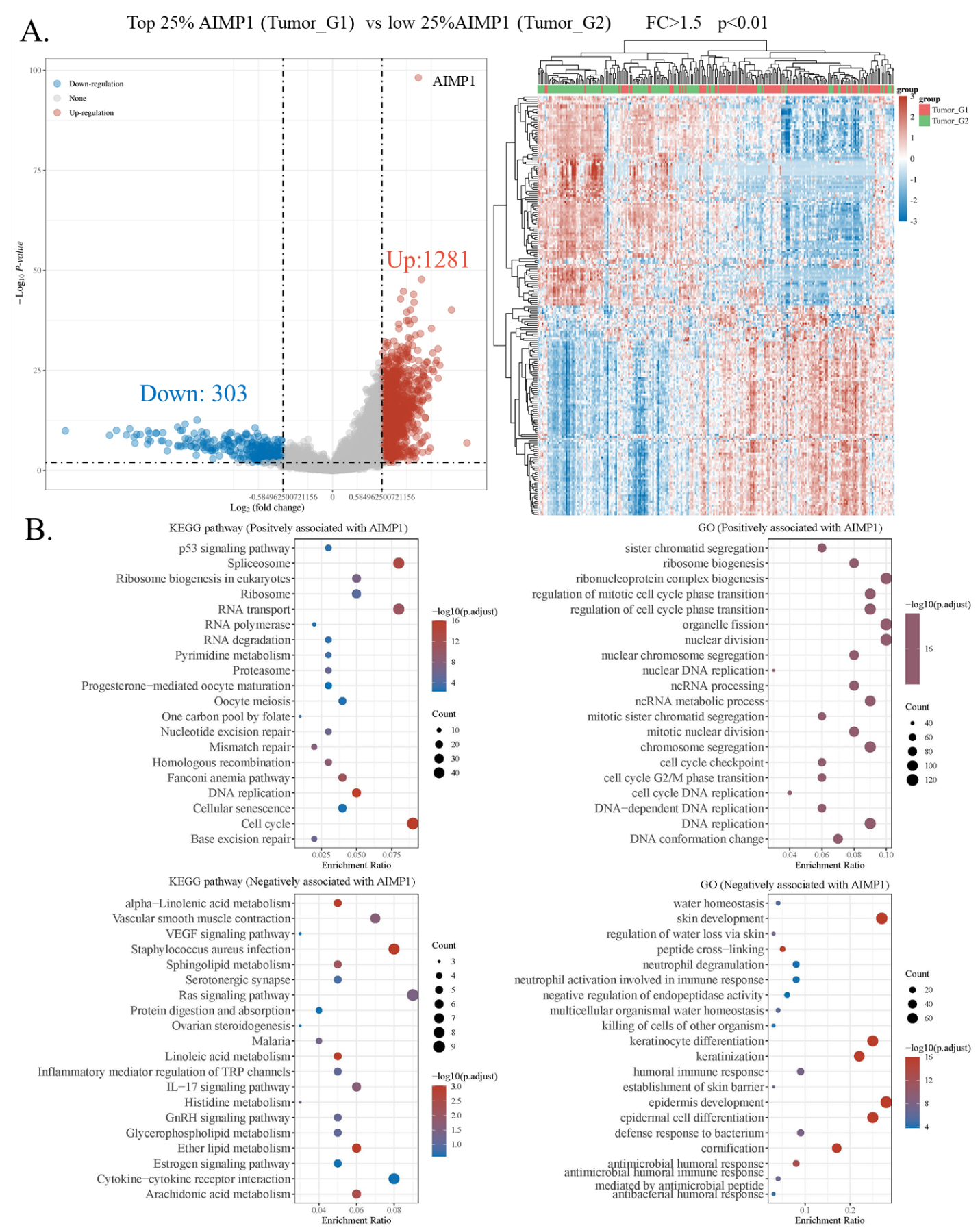
To explore the potential mechanisms involved in the effect of the AIMP1 gene on HNSC, we identified differentially expressed genes between AIMP1 high and low groups. Results showed that 1281 (up) and 303 (down) genes were identified as differentially expressed genes positively and negatively associated with AIMP1 in HNSC respectively. The heat map showed the clusters of samples and significant genes (Fig. 5). These genes were further enriched in GO terminologies and KEGG pathways. Results of KEGG enrichment showed that genes positively associated with AIMP1 were enriched in “Spliceosome”, “RNA transport”, “Cell cycle”, and “DNA replication”, while genes negatively associated with AIMP1 were enriched in “Ras signaling pathway”, “Cytokine-cytokine receptor interaction”, “IL
AIMP1 associated functional states in HNSC
To further explore the potential role of AIMP1 in HNSC, we investigated a single-cell data set of HNSC. We analyzed the correlation of AIMP1 expression and cancer functional state scores of 2150 single HNSC cells and identified 11 functional states that significantly correlated with AIMP1 (
Correlations between AIMP1 expression and functional states in HNSC single-cell datasets. The GSE103322 data (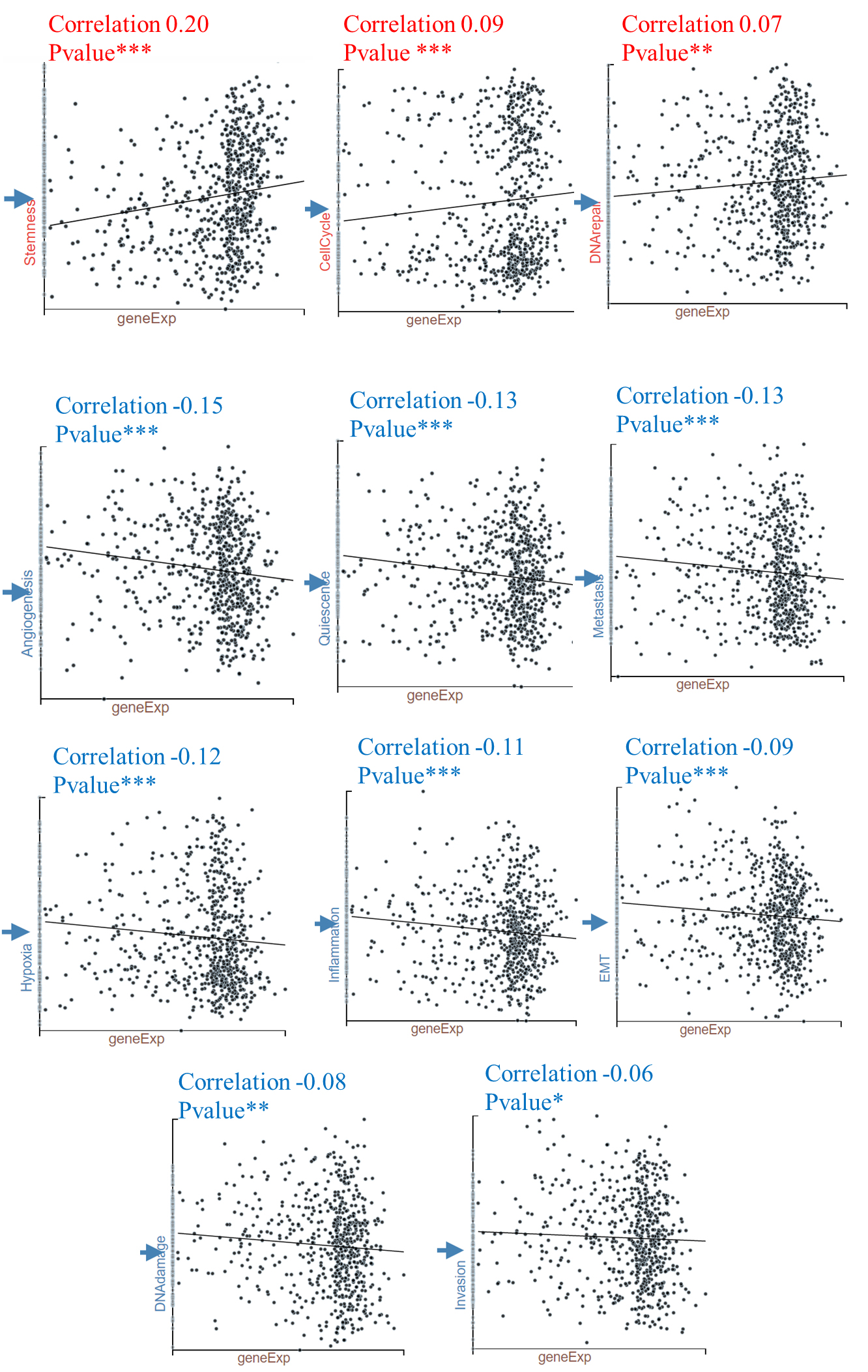
The distribution of AIMP1 expression in different types of cells in HNSC samples. The data were accessed and analyzed using TISCH (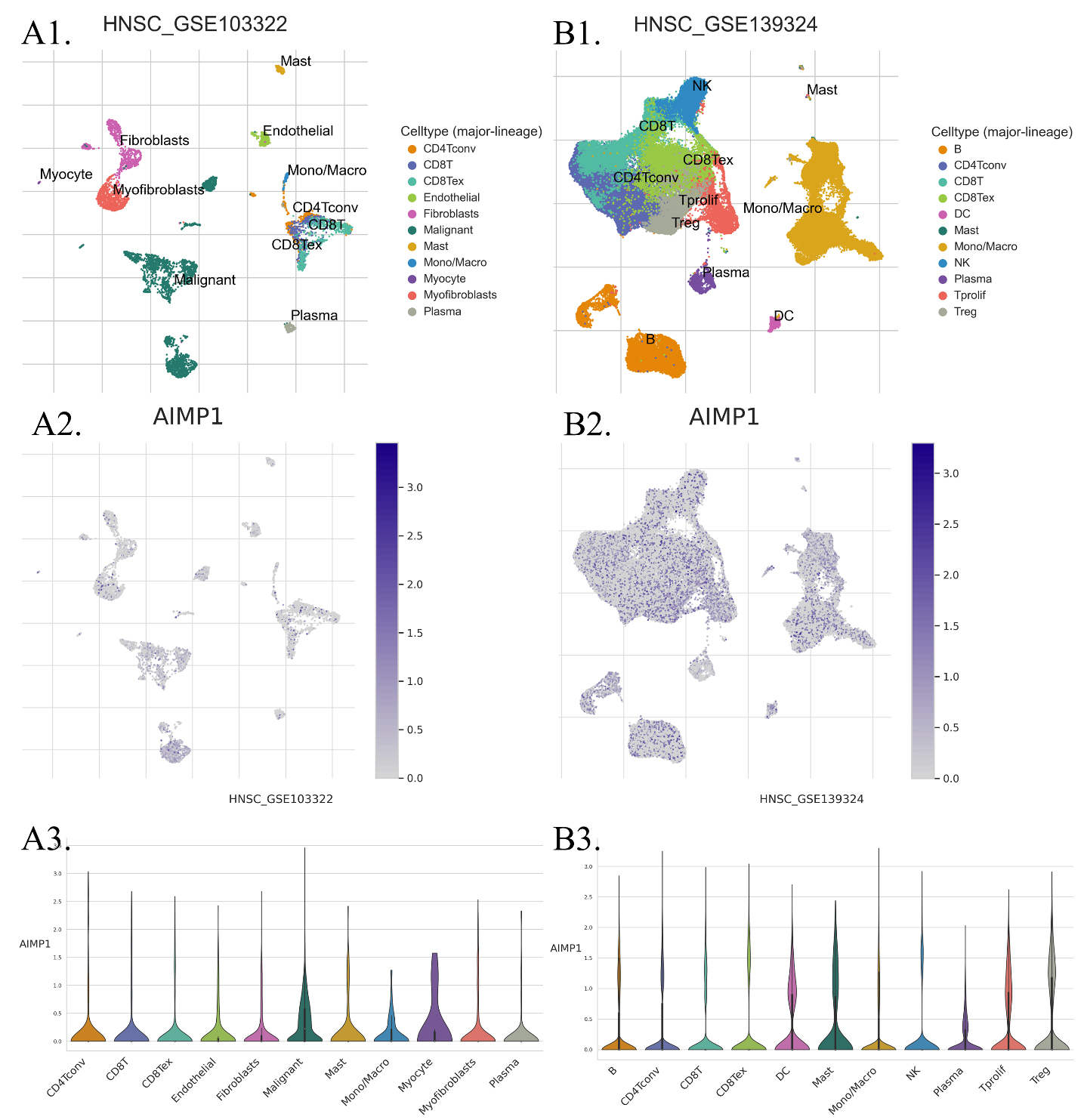
Immune cell infiltration levels of high AIMP1 (75–100%, G1) vs. low AIMP1 (0–25%, G2) groups. A. Immune cell score (xCell) heat map for AIMP1 high and low groups. The significance was analyzed using the Wilcox test. 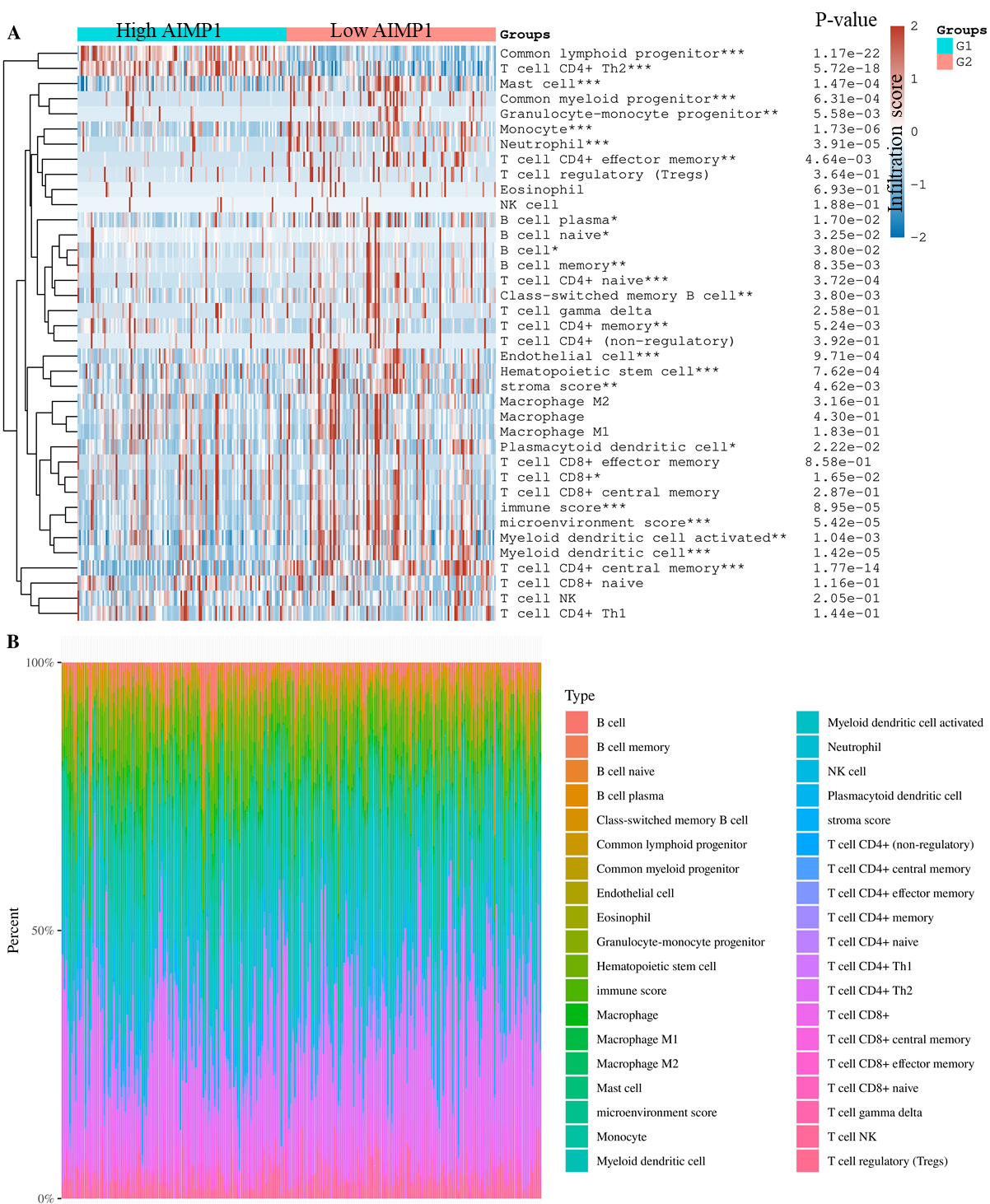
To study the impact of AIMP1 expression on immune cells in HNSC, we first compared AIMP1 expression in different types of cells in HNSC using single-cell seq data. We analyzed two data sets that cover most of the immune cell types in HNSC. Results showed that all of the cell types in HNSC expressed AIMP1. The violin plot of AIMP1 expression in different cell types revealed that malignant (tumor cells) was one of the cell types highly expressing AIMP1 and there was no remarkable difference among immune cell types (Fig. 7). Therefore, we suggested that the expression level of AIMP1 in HNSC samples was mainly dependent on the expression of AIMP1 in tumor cells. Subsequently, we calculated the immune cell infiltration score of TCGA data using the xCell algorithms. We compared immune cell infiltration levels between AIMP1 high (75–100%) and low (0–25%) groups. Results showed that AIMP1 high group had a significantly lower total immune score, stroma score, and microenvironment score. Specifically, compared to the AIMP1 low group, the AIMP1 high group was only significantly higher in the level of Common lymphoid progenitor and T cell CD4
Discussion
This study, for the first time, revealed that AIMP1 was overexpressed in HNSC compared to normal squamous epithelial tissues at both mRNA level and protein levels. The expression of AIMP1 is also associated with higher grade HNSC. Hence, AIMP1 was identified to have not only potential diagnostic but also prognostic values for clinical HNSC treatment. However, a previous study reported that AIMP1 was downregulated in gastric and colorectal cancer [19]. This did not agree with the overexpression of AIMP1 in HNSC. Thus, we suggested the expression pattern and role of AIMP1 might vary from cancer types to cancer types. So far, the expression pattern of AIMP1 in other types of cancers has not been studied, and more effort to study the pan-cancer expression pattern of AIMP1 is required in the future. Nevertheless, the overexpression of AIMP1 in HNSC suggested that it can be a potential diagnostic biomarker for HNSC. A ROC is to evaluate the sensitivity, specificity, and accuracy of a factor. In general, for a diagnostic ROC, an AUC of 0.5 suggests no discrimination (i.e., ability to diagnose patients with and without the disease or condition based on the test), 0.7 to 0.8 is considered acceptable, 0.8 to 0.9 is considered excellent, and more than 0.9 is considered outstanding. The AUC of AIMP1 diagnostic ROC was 0.7 to 0.8, which is acceptable for diagnosis.
Another clinical valuable finding of this study was that AIMP1 expression is significantly associated with HNSC patient’s overall survival. This was demonstrated in both Kaplan-Meier survival analysis and Cox regression analysis. The Cox regression analysis also indicated that AIMP1 expression was independent of age, gender, grade, race, and other clinical factors. In this study, ROC analysis revealed that the AUC of AIMP for HNSC overall survival was over 0.65, and up to 0.75. Generally speaking, for a single prognostic biomarker, an AUC of over 0.65 is acceptable. To increased the accuracy of the biomarker, multiple clinical factors should be also considered. Hence, we not only identified AIMP1 association with survival but also constructed a nomogram with multiple clinical factors to demonstrate the feasibility of using AIMP1 for survival prediction. The nomogram showed considerable confidence, but more variables, such as other reliable biomarkers, should be included to optimize the accuracy of the prediction.
AIMP1 has been suggested to be an exogenous anti-tumor agent for lung cancer [14]. A later study claimed that AIMP1 might suppress gastric and colorectal cancer [19]. Although our results showed that AIMP1 was associated with worse survival of HNSC patients and AIMP1 expression might increase during the development of HNSC, it was not clear what role AIMP1 played in HNSC. Nevertheless, the enrichment study provided some hints for the potential roles of AIMP1 in HNSC. The increased AIMP1 was found associated with multiple proliferation-related terminologies and pathways, such as cell cycle and DNA replication. These results suggested that AIMP1 might be involved in the proliferation of HNSC cells. In addition, the decreased AIMP1 was negatively associated with several terminologies related to skin development and immune cells, which suggested that AIMP1 might deteriorate normal skin development and inhibited immunity. This is rational as AIMP1 is associated with more severe cancer, where cancer might inhibit normal cells.
Single-cell data also revealed some potential roles of AIMP1 in HNSC. The AIMP1 expression was significantly correlated with stemness, cell cycle, and DNA repair which was consistent with the enrichment analysis, further confirming that AIMP1 was associated with cancer proliferation. At the same time, the single-cell data also showed that AIMP1 was negatively correlated with angiogenesis, quiescence, metastasis, hypoxia, inflammation, EMT, DNA damage, and invasion, we suggested some of these states were correspond to the weakened immunity. However, to be admitted, the coefficients of most of these correlations were mostly lower than 0.2, indicating that the correlation of AIMP1 was not very strong. In addition, whether the changes in these states were the reasons or the results of AIMP1 was not clear. To date, the exact role of AIMP1 in HNSC has not been investigated, we urged more functional experiments in the future to validate our conclusion.
Another interesting finding of this study was that AIMP1 might affect the tumor immune microenvironment. Tumor immune single-cell data showed that HNSC tumor cells tended to express a higher level of AIMP1 than immune cells, and there was no difference in AIMP1 expression among different types of immune cells. These results indicated that a higher level of AIMP1 in HNSC samples of TCGA should result from higher levels of AIMP1 in tumor cells, but not any certain types of immune cells. This can partly rule out the bias in the comparison of immune infiltrations because the immune cells will not affect the division of the high and low AIMP1 groups. The immune infiltration analysis revealed that AIMP1 was negatively associated with the immune cell infiltration level, which was consistent with the enrichment results. The only two positive associated immune cell types were lymphoid progenitor and T cell CD4
Conclusion
AIMP1 is a potential diagnostic and prognostic biomarker for HNSC patients and can potentially affect the proliferation and tumor immune microenvironment of HNSC cells. This study provided a novel molecular marker for the improvement of clinical HNSC treatment.
Availability of data and materials
The source of the raw data was provided in the paper and the raw analysis data of this study are provided by the corresponding author with a reasonable request.
Consent for publication
All authors had given final approval of the version to be published.
Funding
This study received funding from Shenzhen Traditional Chinese Medicine Hospital.
Ethical approval
Not applicable.
Authors’ contributions
Hengrui Liu contributed to the design of the study. Yixue Li and Hengrui Liu contributed to the data acquisition, data analysis, and the drafting of the manuscript. Hengrui Liu supervised the project.
Author information
The corresponding author of this study, Hengrui Liu, is a Principal Investigator in Biocomma Limited.
Footnotes
Acknowledgments
The authors thank the support of Biocomma Limited. We also thank Weifen Chen, Zongxiong Liu, Yaqi Yang, and Zirui Tian for their help.
Conflict of interest
The authors claimed that there is no conflict of interest.
