Abstract
Gastric cancer (GC) is a common cancer with high mortality and morbidity rates worldwide. Although medical and surgical treatments have improved, the mechanisms of the progression of GC remain unclear. Platelet-derived growth factor receptor-
Introduction
Gastric cancer (GC) is one of the most common cancers worldwide [1]. Although medical and surgical treatments have improved, survival rates remain poor because in many cases, GC is already at an advanced stage when diagnosed [2]. Immunotherapy has emerged as a promising cancer treatment with major successes in breast cancer [3], prostate cancer [4] and melanoma [5]. Recent studies have reported that several types of tumor-infiltrating immune cells are associated with the disease outcomes of various human cancers [6, 7, 8]. Immune cell infiltration in human gastrointestinal cancers is associated with cancer progression and is a reliable prognostic predictor [9, 10, 11, 12, 13]. Both cellular composition and organization of tumor-infiltrating immune cells are crucial for inhibiting cancer progression and are implicated in the success of cancer immunotherapy [9, 14].
Platelet-derived growth factor receptor beta (PDGFRB) encoded by PDGFRB gene [15], is a typical transmembrane tyrosine kinase receptor that promotes blood vessel formation when activated by platelet-derived growth factor (PDGF) [16] and controls many important cellular processes such as growth, proliferation, movement and survival [17, 18, 19, 20]. Its dysregulation is closely related to carcinogenesis, as well as to cardiovascular diseases [20]. Raja et al. (2017) suggested that PDGFRB may serve as a prognostic factor in GC by immunohistochemical expression [21]. Wang et al. (2019) reported that NRP1 and PDGFRB were significantly correlated with tumor malignant phenotypes serving as potential prognostic biomarkers for GC patients [22]. Although a series works revealed that the overexpression of PDGFRB correlates with the progression of gastric carcinoma [23] and high expression of epithelial PDGFRB is associated with poor disease-free survival (DFS) and overall survival (OS) in GC patients [24], there was rare study to explore the relationship between PDGFRB expression and the tumor immune microenvironment. This study aimed to explore the relationship of PDGFRB expression with clinicopathologic characteristics, immune cell infiltration status, and prognosis in GC through a bioinformatics analysis.
Material and methods
Oncomine database analysis
Oncomine1
Oncomine:
GEPIA2
GEPIA:
UALCAN3
UALCAN:
Kaplan Meier plotter4
Kaplan Meier plotter:
TIMER5
TIMER:
Expression levels of PDGFRB in gastric cancer. (A) PDGFRB expression in different cancers compared to normal tissues in Oncomine. (B–H) Differential expression (DE) of PDGFRB in Oncomine. (I) DE of PDGFRB in GEPIA using TCGA datasets. (J) DE of PDGFRB in microarray data with accession number of GSE79973. (K) DE of PDGFRB in microarray data with accession number of GSE118916.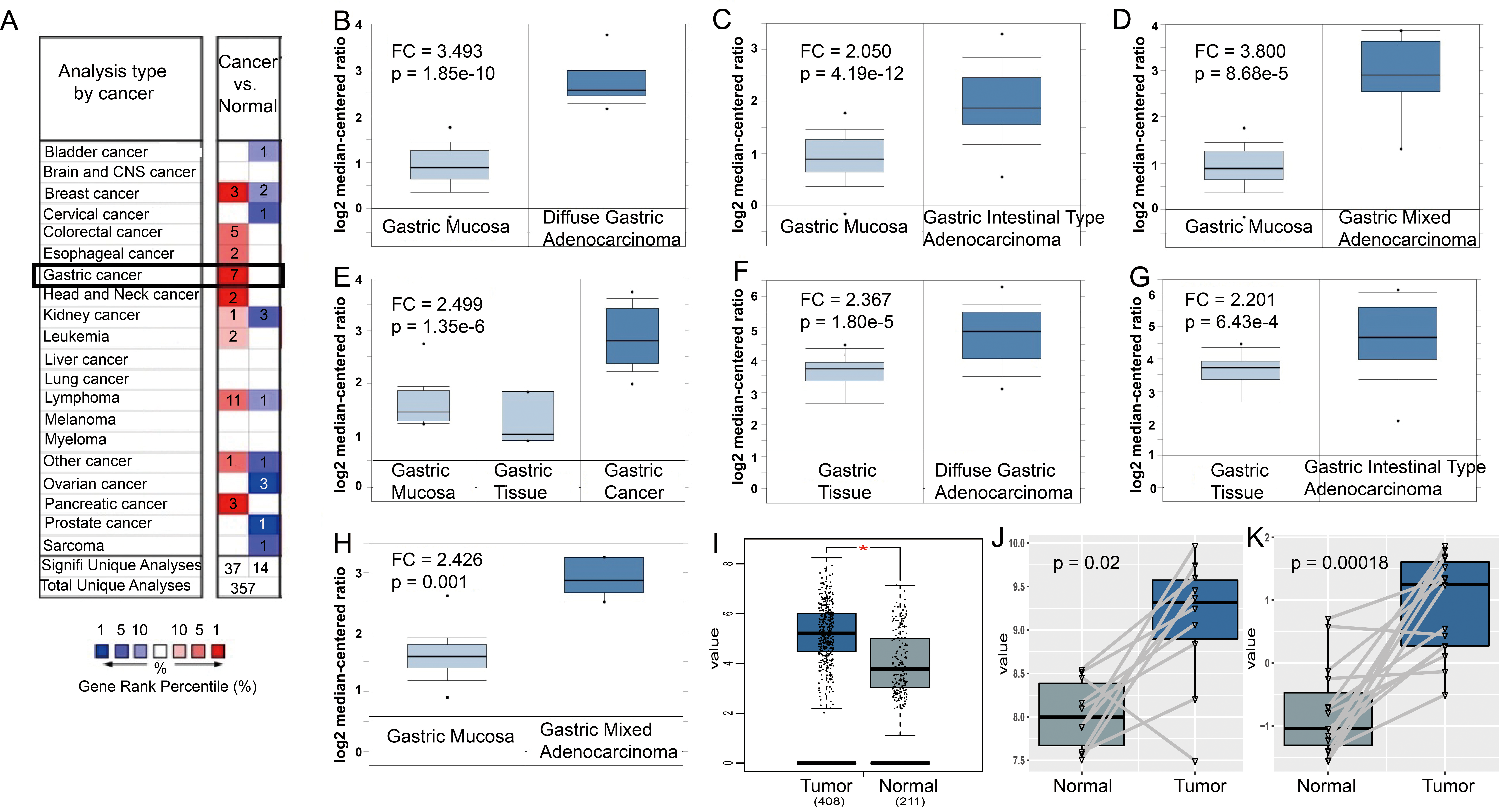
PDGFRB transcription in various subgroups of GC patients (UALCAN). (A) Normal and STAD tissue samples. (B) Normal samples of both genders and male and female STAD patients. (C) Normal samples of any age and STAD patients aged 21–40, 41–60, 61–80, and 81–100 years. (D) Normal samples and STAD patients with tumor grades 1, 2, and 3. (E) Normal samples and African American, Caucasian, and Asian STAD patients. (F) Normal samples and STAD patients with regional lymph node metastasis status N0, N1, N2, and N3. (G) Normal samples and STAD patients with cancer stages 1, 2, 3, and 4. (H) Normal samples and patients with and without H. pylori infection. (I) Normal samples and patients with and without TP53 mutation (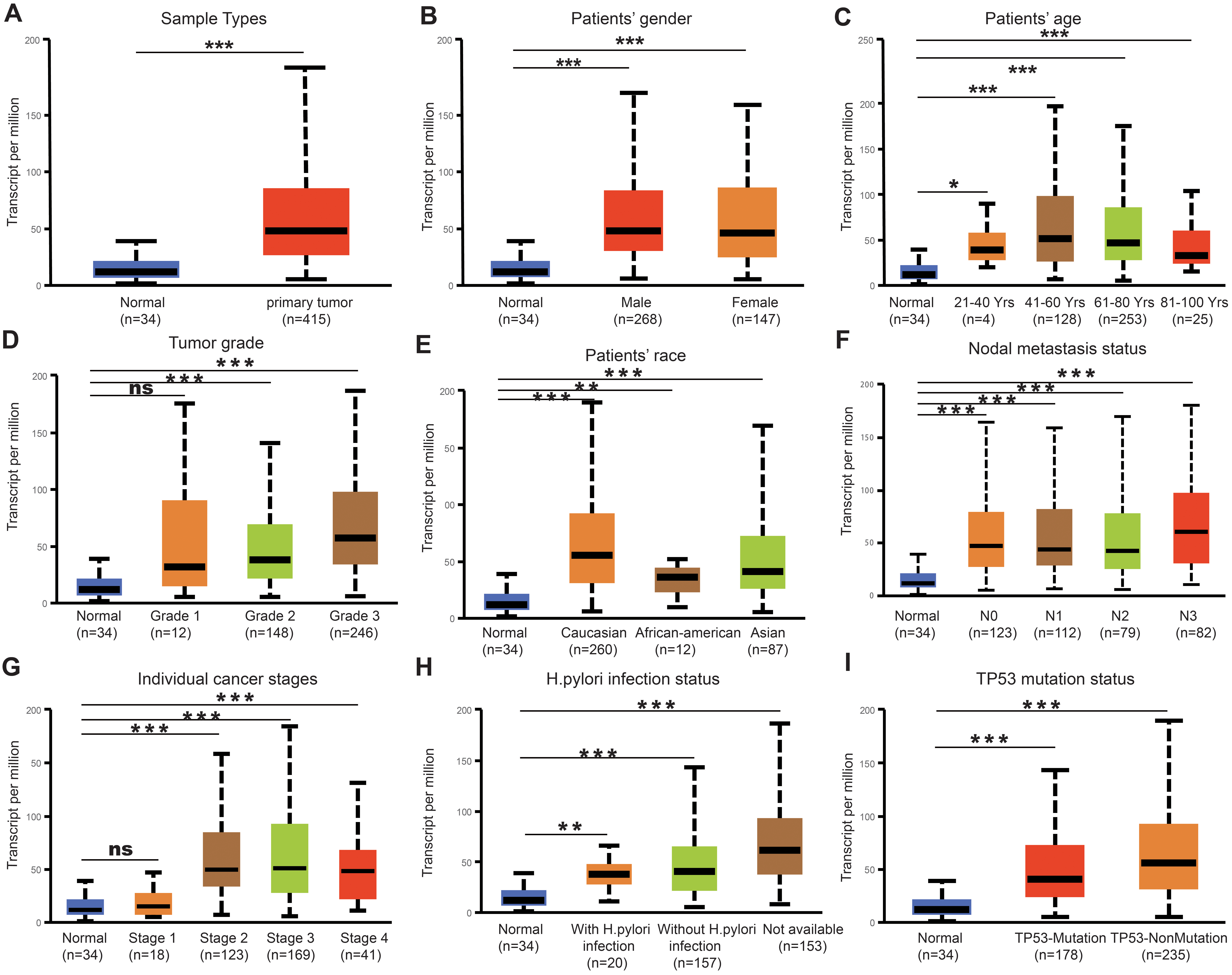
Comparison of Kaplan-Meier and GEPIA survival curves of high and low PDGFRB expression in STAD. Low expression was associated with favorable OS in Kaplan-Meier Plotter using (A) GeneChip and (B) RNA-seq data. (C) Low expression was associated with favorable progression-free survival in Kaplan-Meier Plotter using GeneChip data. (D) Low expression was associated with favorable OS in GEPIA.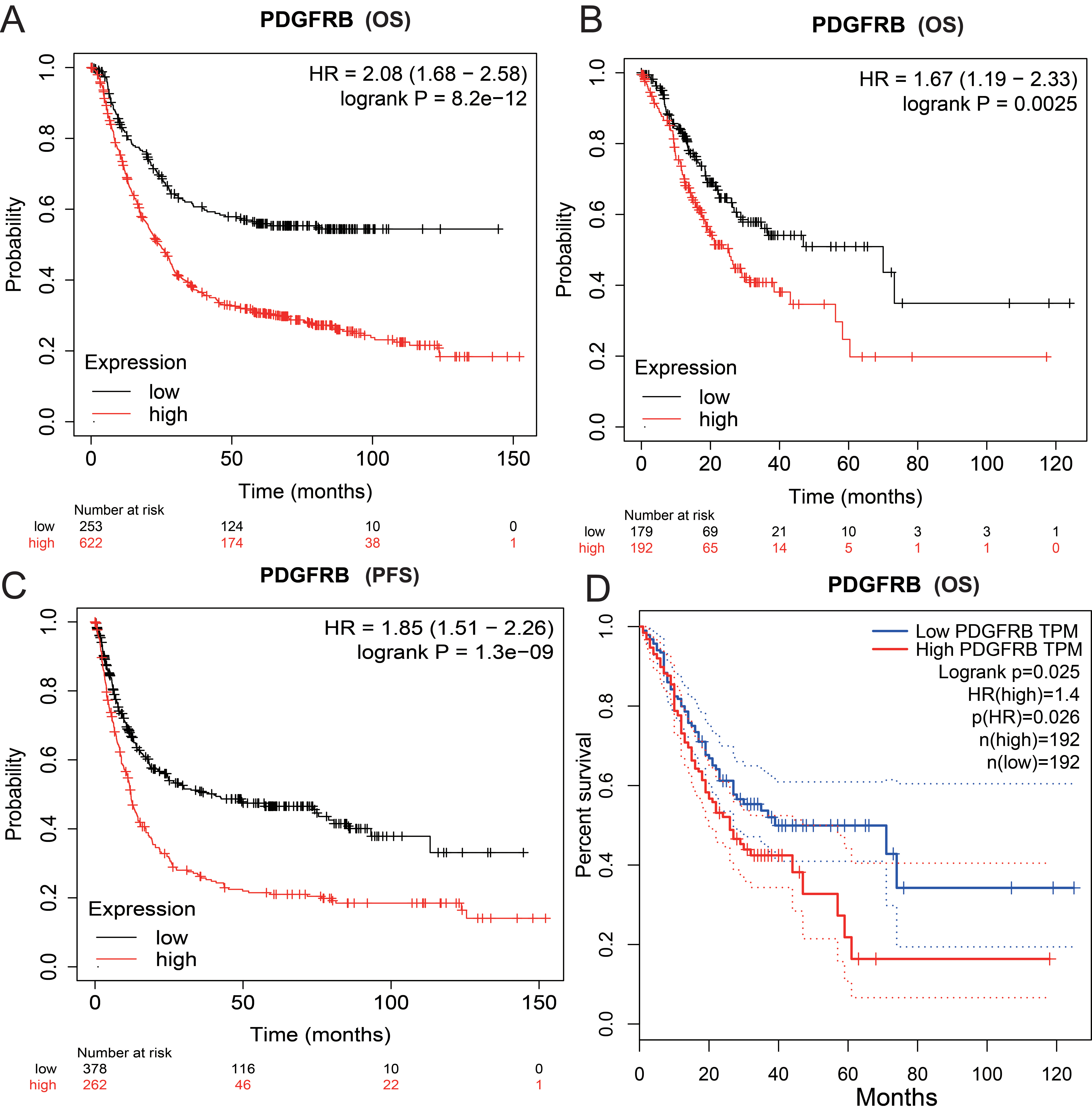
The CIBERSOFT method quantifies the relative scores for 22 human immune cell types with gene-based deconvolution algorithm and the characteristics of 547 marker genes. To enhance the robustness of the results, the algorithm applies Monte Carlo sampling to obtain the deconvoluted
Function exploration for PDGFRB
LinkedOmics6
External datasets with accession numbers of GSE118916 and GSE79973 were downloaded from the Gene Expression Omnibus (GEO) database. Differentially expressed genes were identified using R package limma [38] with fold change (FC)
Results
Elevated expression of PDGFRB in GC
We searched Oncomine to study the differences in PDGFRB expression between different tumor tissues and normal tissues. The results showed significantly low expression levels of PDGFRB in bladder, cervical, ovarian and prostate cancer and sarcoma but significantly high expression levels of PDGFRB in colorectal, esophageal, pancreatic, head and neck cancer, GC, and leukemia (Fig. 1A). There were 7 datasets supporting the expression elevation for PDGFRB in GC (Fig. 1B–H). Then GEPIA with the RNA-seq datasets (Fig. 1I) and GEO with two independent microarray datasets of GSE79973 and GSE118916 (Fig. 2J and K) were used to evaluate differences for PDGFRB expression between GC and normal tissues. All the results showed significant overexpression of PDGFRB in GC. Furthermore, subgroup analysis of multiple clinicopathologic features of TCGA STAD samples in UALCAN consistently showed elevated transcriptional levels of PDGFRB. The expression of PDGFRB was significantly higher in STAD than in normal tissues in subgroup comparisons by gender, age, race, disease stage, tumor grade, Helicobacter pylori infection, and TP53 mutation (Fig. 2).
PDGFRB is a prognosis predictor in GC
The link between PDGFRB expression and patient outcomes in GC was investigated using Kaplan-Meier Plotter in which the genechip data (Fig. 3A and C) and RNA-seq data (Fig. 3B) were applied. Furthermore, GEPIA was also used to validate the survival analysis results using TCGA datasets (Fig. 3D). The results showed that high PDGFRB expression was associated with poor survival.
Correlation of PDGFRB expression with clinicopathological features of GC
High PDGFRB expression was found to be associated with poor prognosis in GC. The underlying mechanisms were explored using Kaplan-Meier Plotter. The correlations between PDGFRB and patient clinicopathologic characteristics were calculated. The results showed that PDGFRB upregulation was associated with worse OS and progression-free survival (PFS) in male and female patients, certain stages and TNM stages (stage N categories represent lymph node involvement), and two types of Lauren classification and differentiation (
Correlation of PDGFRB mRNA expression and clinical prognosis in gastric cancer with different clinicopathological factors by Kaplan-Meier Plotter
Correlation of PDGFRB mRNA expression and clinical prognosis in gastric cancer with different clinicopathological factors by Kaplan-Meier Plotter
We investigated the functions and pathways in which PDGFRB might participate. LinkedOmics was used to identify significantly and positively correlated genes correlated to PDGFRB using 415 STAD samples. The top 50 correlated genes including LRRC32, FBN1, COL6A3, COL8A1 and CDH11, were listed (Fig. 4A). Functional enrichment analysis for the top 50 genes showed that they were involved in extracellular matrix organization, cell adhesion, blood vessel development, angiogenesis, PI3K-Akt signaling pathway, protein digestion and absorption and pathways in cancer (Fig. 4B).
Genes in significantly positive correlation with PDGFRB in gastric cancer. (A) Heatmap showing genes positively correlated with PDGFRB in STAD (top 50). (B) Functional enrichment for the top 50 coexpressed genes.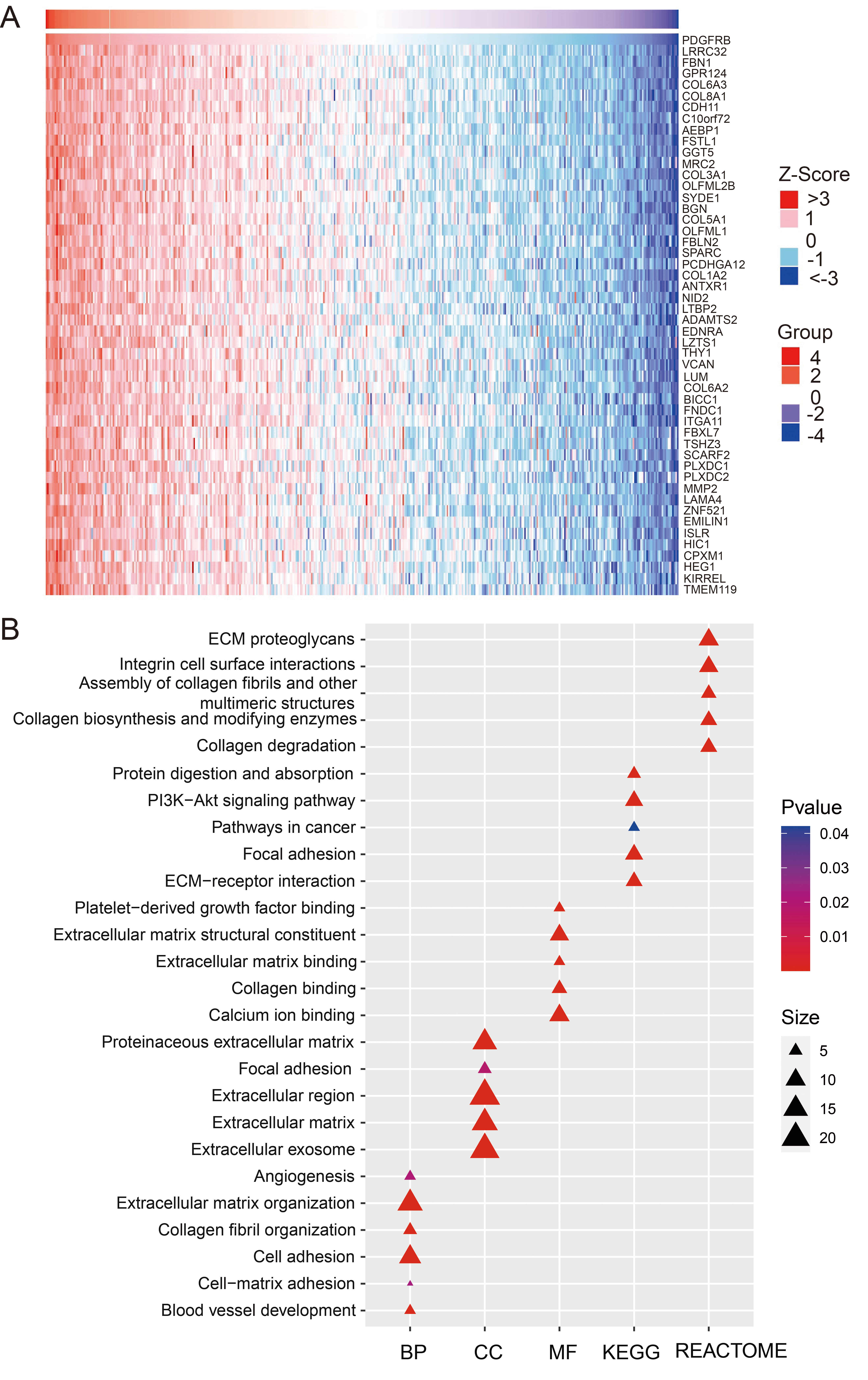
Correlations between PDGFRB expression and (A) B cell, (B) CD8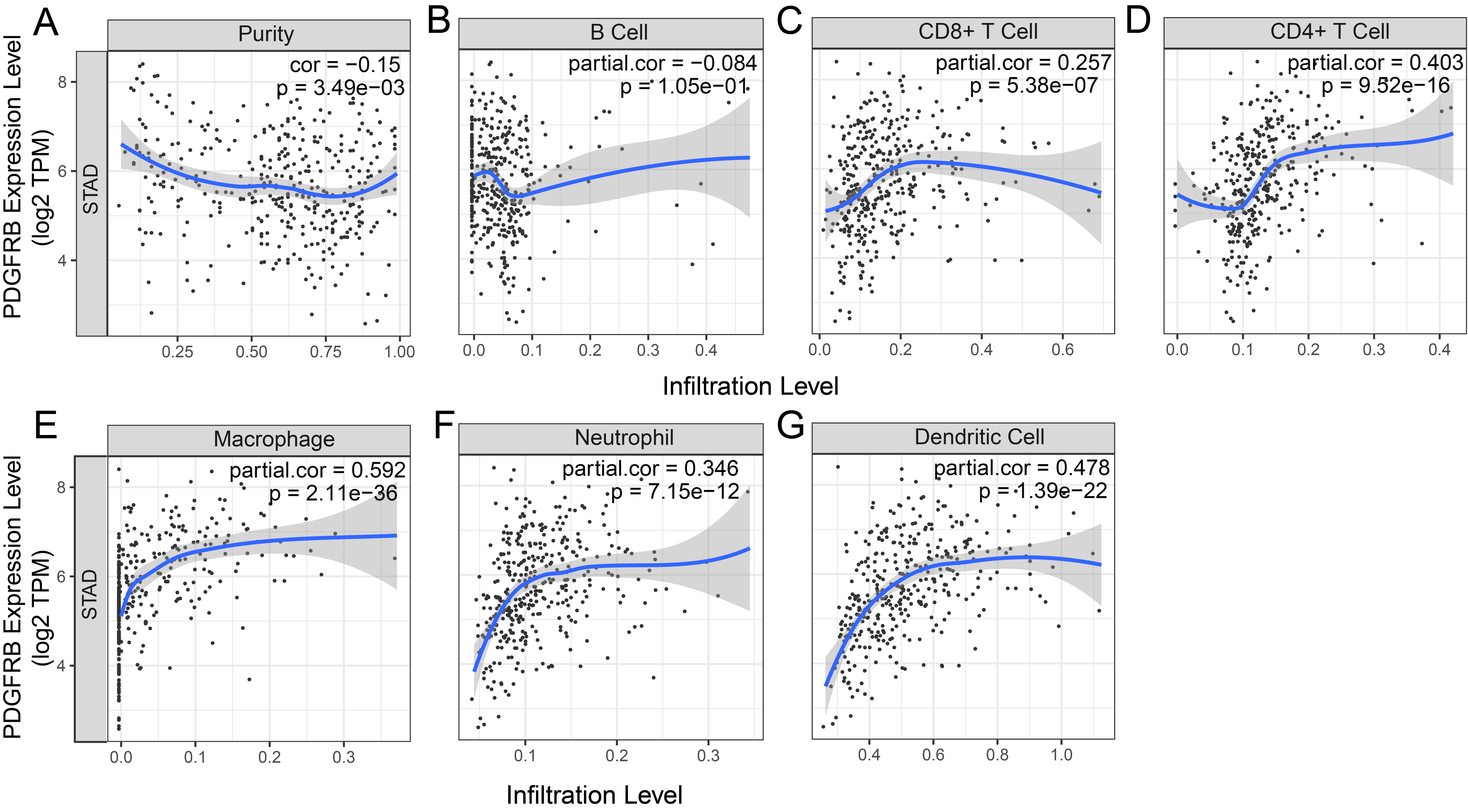
Cumulative survival analysis for (A) B cells, (B) CD8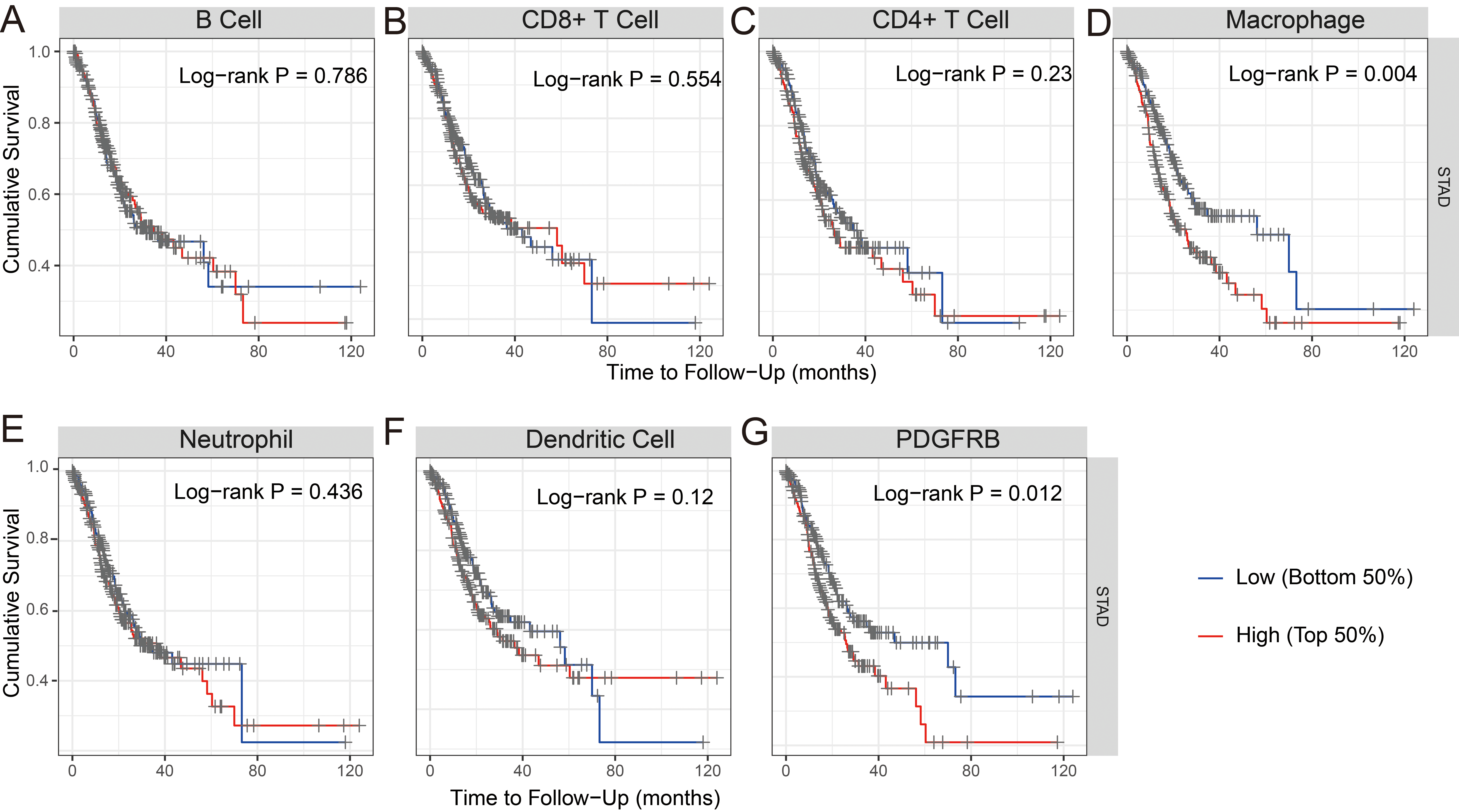
We examined whether the transcription levels of PDGFRB in gastric tumors correlated with immune infiltration. TIMER was used to analyze the correlations between PDGFRB expression and STAD. The results showed that PDGFRB did not correlate with B cells but significantly positively correlated with the levels of immune infiltration of CD8
Correlation analysis of PDGFRB expression and immune marker sets
To investigate the relationships between PDGFRB and tumor-infiltrating immune cells, we calculated the correlations between PDGFRB and the immune marker sets of CD8
Scatterplots of the correlations between PDGFRB expression and gene markers of (A) monocytes, (B) TAMs, (C) M1 macrophages, and (D) M2 macrophages.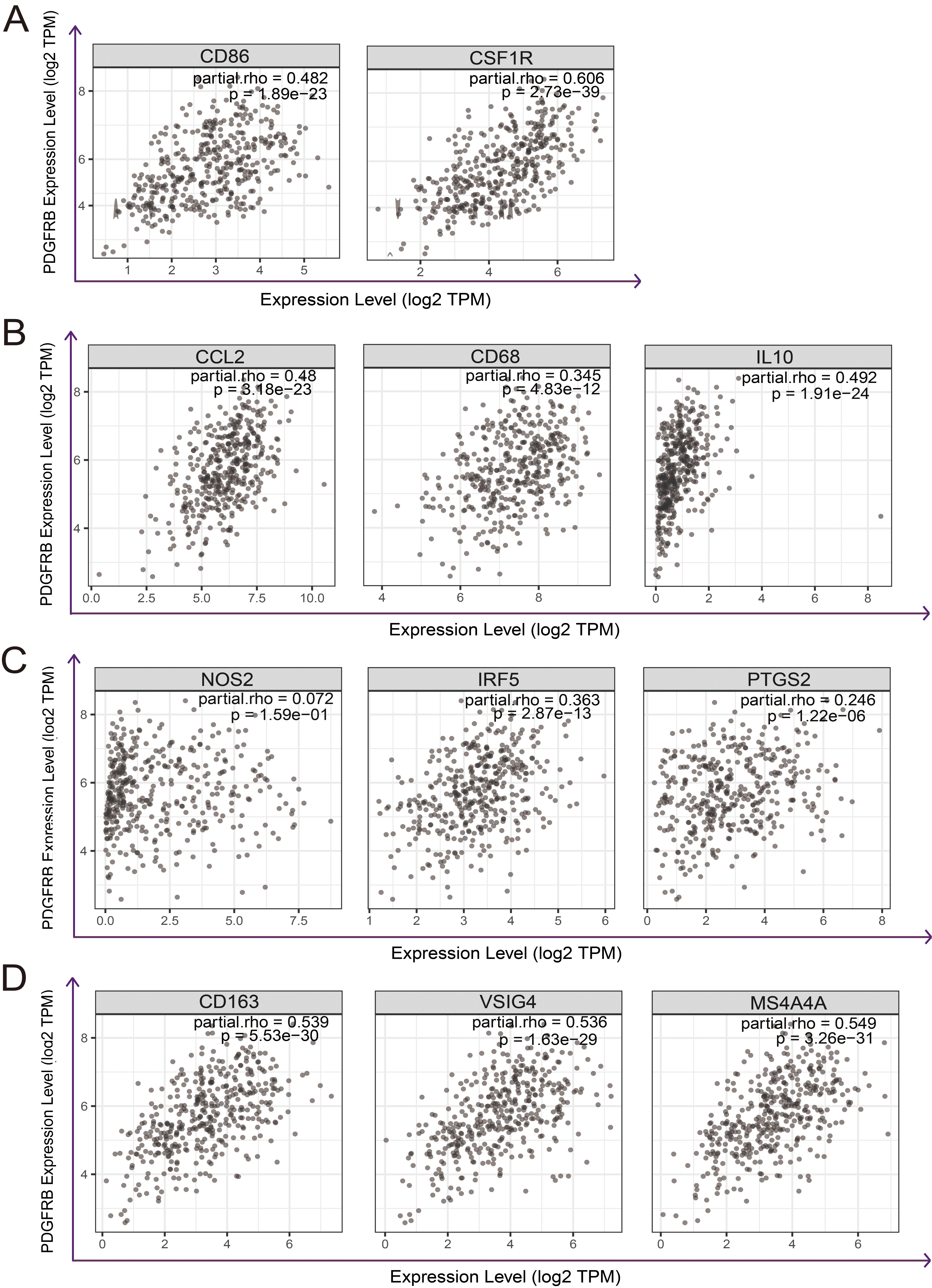
Correlation analysis between PDGFRB and related gene markers of immune cells in TIMER
Bold values indicate
Correlation analysis between PDGFRB and related gene markers of monocyte and macrophages in GEPIA
Using CIBERSORT, the 22 immune cell components for GC samples in TCGA were infered (Fig. 8A). The results showed that macrophages M2 was the most abundant immune cell across the samples (Fig. 8B). PDGFRB expression had correlation with M2 macrophages (
The immune cell components predicted by CIBERSORT. (A) The barplot of the immune cell components. (B) The boxplot of the immune cell components. (C) Macrophages M2 components are plotted according to PDGFRB expression level. (D) The scatterplot of the PDGFRB expression and the macrophages M2 components.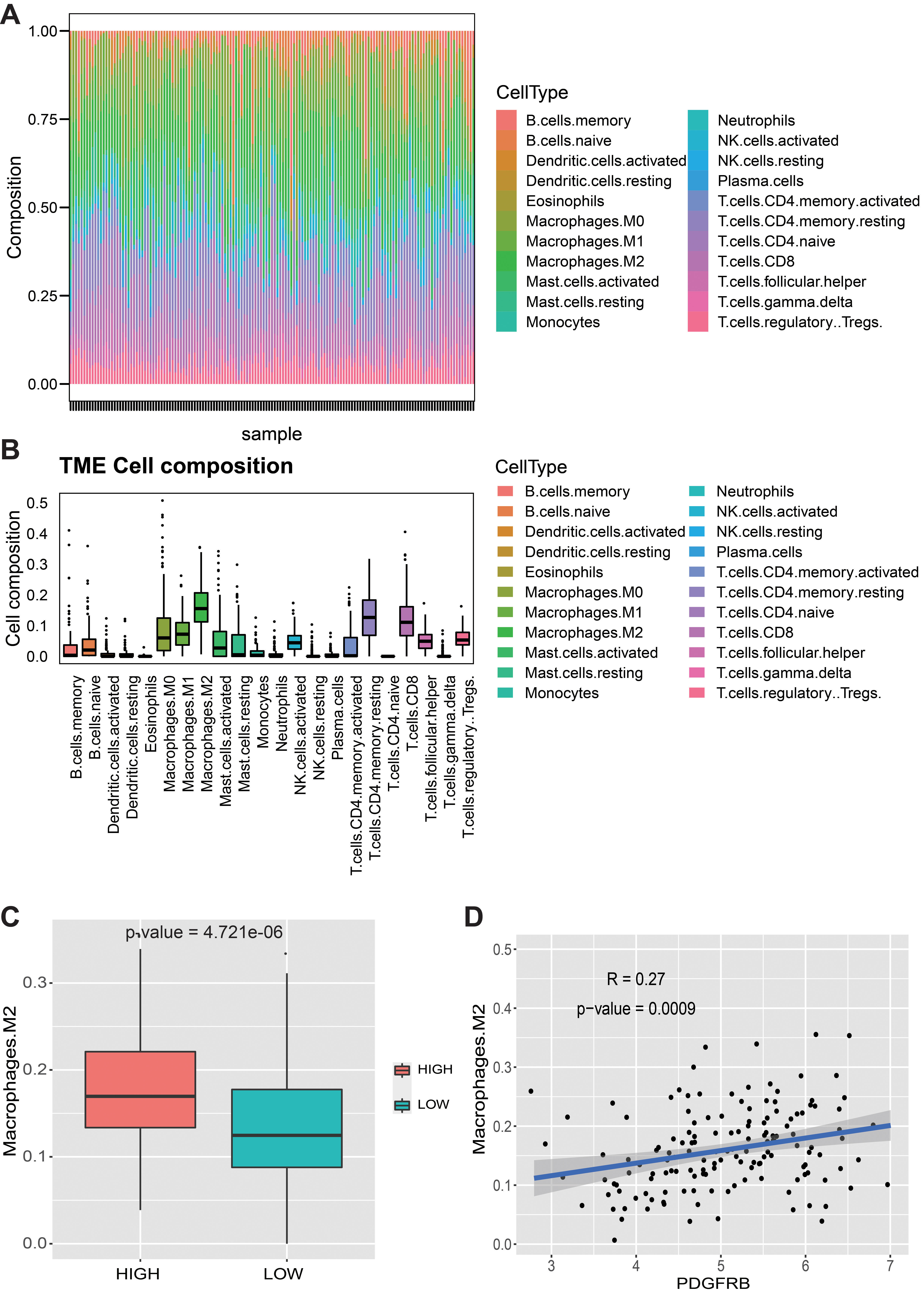
Although Researchers have been trying to explore the biomarkers in GC [39, 40, 41, 42], the prognosis biomarker of GC patients remains poor Platelet-derived growth factor receptors play a critical role in angiogenesis and tumor cell proliferation and metastasis [43]. PDGFRB promotes proliferation and migration of cancer cells and angiogenesis, and has been used as a target in several cancer treatments [19, 44]. In this study, differential expression of PDGFRB between cancer and normal tissues was observed in GC. The results demonstrated that PDGFRB expression was remarkably increased in GC tissues. Abnormal gene expression may be associated with tumorigenesis and prognosis of patients [45].
GEPIA and Kaplan-Meier Plotter analyses revealed that high PDGFRB expression correlates with poor prognosis in GC. Both of them showed that high PDGFRB expression correlates with a high HR for OS and PFS in GC. High PDGFRB expression correlates with poor prognosis in GC stages 2–4, T2–T4, and N1–N3. The highest HRs for OS and PFS were observed when PDGFRB was highly expressed. High PDGFRB expression is most likely to impact the prognosis of GC patients with lymph node metastasis. This finding is in line with a previous study reporting that PDGFRB overexpression correlate with cancer progression and lymphogenous metastasis of gastric carcinoma [46], suggesting that PDGFRB may serve as an oncogene to promote gastric carcinogenesis. To study the molecular functions and underlying mechanisms of PDGFRB in gastric carcinogenesis, we performed coexpression analysis for PDGFRB in GC. Functional analysis of PDGFRB and its positively coexpressed genes showed that they are involved in extracellular matrix organization, cell adhesion, blood vessel development and angiogenesis which play important roles in carcinogenesis and cancer progression [47]. Controlled remodeling of the extracellular matrix is essential for the growth, invasion, and metastasis of malignant tumors [48]. Tumorigenesis-related pathways such as the “PI3K-Akt signaling pathway” was also identified which was widely reported to play a crucial role in cancer occurrence by promoting cell proliferation and inhibiting apoptosis [49].
The analysis of the correlation between immune infiltration levels, various immune marker sets, and PDGFRB expression provides insights into the potential significance of PDGFRB for tumor immunology and its use as a cancer biomarker. Gene markers of M1 macrophages, such as PTGS2 and NOS2, showed weak correlations with PDGFRB expression, whereas M2 markers, such as CD163, VSIG4, and MS4A4A, showed strong correlations (
Our results also indicate that PDGFRB has the potential to activate Tregs and induce T cell exhaustion. PDGFRB upregulation positively correlates with the expression of Treg and T cell exhaustion markers (FOXP3, CCR8, STAT5B, TIM-3, PD-1, CTLA4, and LAG3). TIM-3, also called HAVCR2, a crucial exhausted T cell surface protein, highly correlates with PDGFRB expression in STAD, playing a crucial role in tumor immunological tolerance [58]. Furthermore, we found significant correlations between PDGFRB expression and the regulation of several markers of Th cells (Th1, Th2, Tfh, and Th17). These correlations could be indicative of a potential mechanism whereby PDGFRB regulates T cell functions in STAD. Our study provides insights into possible mechanisms that might explain why PDGFRB expression correlates with immune cell infiltration and poor prognosis in GC. Although this study indicated that PDGFRB plays an important role in the recruitment and regulation of infiltrating immune cells in GC, the immune reaction is a complex and dynamic process. Therefore, further study is necessary to explore the role of PDGFRB involvement in the immune response in GC.
Conclusion
In summary, we demonstrated that overexpression of PDGFRB was associated with clinicalpathological parameters and predicted a poor prognosis. High PDGFRB expression was closely associated with increased immune infiltration levels of CD8
Authors’ contributions
Conception: Baohong Liu, Yongliang Lou and Lingling Zhao.
Interpretation or analysis of data: Baohong Liu, Xingxing Xiao and Ziqin Lin.
Preparation of the manuscript: Baohong Liu.
Revision for important intellectual content: Yongliang Lou and Lingling Zhao.
Supervision: Yongliang Lou and Lingling Zhao.
Availability of data and materials
The analyzed data sets generated during the study are available from the corresponding author on reasonable request.
Funding
This work was supported by the National Natural Science Foundation of China [grant number 82002117]; open fund of Wenzhou Key Laboratory of Sanitary Microbiology [grant number ZD202003KF02].
Declaration of competing interest
The authors declare that they have no competing interests.
