Abstract
BACKGROUND:
The disorder of LINC01207 has a significant regulatory effect on cancers, nevertheless its role in non-small cell lung cancer (NSCLC) have not been illustrated. This study investigated the regulatory effect of LINC01207 on NSCLC and clarify its molecular mechanism.
METHODS:
Bioinformatics analysis was used to find the target lncRNA, miRNA and mRNA. LncBase and TargetScan databases predicted the relationship between LINC01207, miR-525-5p and ARHGAP11A. Dual-luciferase reporter gene assay and RNA binding protein immunoprecipitation assay were used to verify the binding relationship between genes. Fluorescence in situ hybridization assay was used to localize the expression of LINC01207 in NSCLC tissue. qRT-PCR and Western blot assays were used to measure the expression of LINC01207, miR-525-5p and ARHGAP11A. CCK-8 assay, Transwell assay and flow cytometry assay were used to detect NSCLC cell abilities. Mouse xenograft models further determined the effect of LINC01207 on the growth of NSCLC in vivo.
RESULTS:
LINC01207 was up-regulated in NSCLC tissue and cells, which was mainly localized in the cytoplasm. LINC01207 knockdown could inhibit the proliferation, migration and invasion of cancer cells and induce cell apoptosis. In addition, silencing LINC01207 could suppress tumor growth in vivo. LINC01207 could sponge and inhibit the expression of miR-525-5p in NSCLC cells, and inhibiting LINC01207 and miR-525-5p simultaneously could reverse the effect of miR-525-5p on the progression of NSCLC cells. Further study on downstream target genes showed that miR-525-5p could restrain the expression of ARHGAP11A, and then affect the progression of NSCLC. LINC01207 acting as a competing endogenous RNA (ceRNA) could regulate the expression of ARHGAP11A by competitively binding with miR-525-5p.
CONCLUSION:
LINC01207 regulates the progression of NSCLC by regulating the miR-525-5p/ARHGAP11A axis as a ceRNA and plays a carcinogenic role. In conclusion, our study elucidates the mechanism of LINC01207 regulating the progression of NSCLC, and provides a new idea for the diagnosis and treatment of NSCLC guided by lncRNA.
Introduction
The incidence and mortality of lung cancer are the highest among all cancers, and the number of lung cancer deaths accounts for 18.4% of all cancer deaths in 2018 [1]. In recent decades, despite improvements in surgery and medication, the 5-year survival rate of patients with lung cancer has been still very low, ranging from 4% to 17% in different cancer stages and regions [2]. Lung cancer is divided into small cell lung cancer (SCLC) and non-small cell lung cancer (NSCLC), and more than 85% of patients with lung cancer are diagnosed with NSCLC [3]. Poor prognosis is a staple cause of high mortality of NSCLC [4]. At present, understanding of the biological mechanisms of NSCLC remains limited and thus no more effective measures can be taken to improve the therapeutic effect. Therefore, it is of great significance to find novel prognostic biomarkers in the early stage of NSCLC.
Long non-coding RNAs (lncRNAs) function as oncogenes or tumor suppressor genes in the progression of cancers. Many lncRNAs have been identified as key genes that regulate the proliferation, migration, invasion, apoptosis and epithelial mesenchymal transition (EMT) of NSCLC cells, and may become diagnostic markers [5]. For example, lncRNA PVT1 in NSCLC tissue and cells can act as a molecular sponge of miR-17-5p to influence the expression of BAMBI, thus improving cell growth activity [6]. Hu et al. [7] found that LINC01296 can facilitate the proliferation and migration of NSCLC cells by regulating miR-5095. Yang et al. [8] found that LINC00525 as a ceRNA of miR-338-3p, indirectly regulates IRS2 and hastens the progression of NSCLC, thus it is believed that LINC00525 can inhibit tumor growth in vivo by regulating miR-338-3p/IRS2 axis. In addition, lncRNA HOXD-AS1 is a potential prognostic biomarker of NSCLC and can indirectly regulate the expression of matrix metallopeptidase 9 (MMP9) by binding with miR-133b [9, 10]. Although the functions of some lncRNAs in NSCLC have been reported, the specific role of most lncRNAs in NSCLC remains unclear.
Recently, it has been found that LINC01207 is abnormally expressed in pancreatic cancer and plays a role as an oncogene. LINC01207 has been found to be located in the 4q32 locus of the genome, containing three exons and two introns. It has been reported that LINC01207 can facilitate the proliferation of lung adenocarcinoma cells and is related to tumor node metastasis (TNM) classification [11]. LINC01207 is up-regulated in pancreatic cancer and promotes cell progression. Besides, silencing LINC01207 can promote cell apoptosis and autophagy, and inhibit cell growth [12]. These studies indicate that LINC01207 may be a promising biomarker and target for cancer diagnosis and treatment. However, knowledge about the regulatory mechanism of LINC01207 in NSCLC remains low.
In this study, LINC01207 was chosen as the research object to study the biological function and possible molecular mechanism of it in regulating the progression of NSCLC cells. Our study will help to better understand the oncogenic mechanism of LINC01207 and provide a potential target for NSCLC.
Materials and methods
Bioinformatics analysis
GSE101929 and GSE44077 chips of NSCLC were acquired through GEO database (
Patients and tissue samples
From March 2018 to December 2019, 33 pairs of tumor tissue and matched adjacent tissue samples were obtained from all the patients who underwent surgical tumor resection in the Second Affiliated Hospital of Zhejiang University School of Medicine. All samples were frozen immediately in a liquid nitrogen tank. The registered patients met the following criteria: (a) definite pathological diagnosis of NSCLC, (b) informed consent was obtained before surgery, (c) having not received chemotherapy. The project was approved by the Research Ethics Committee of the Second Affiliated Hospital of Zhejiang University School of Medicine.
Cell culture
Normal human bronchial epithelial cells HBE (ATCC
Plasmid construction and cell transfection
The shRNA and corresponding negative control of LINC01207 were synthesized by Sengong Biotechnology Co., Ltd. (Shanghai, China). MiR-525-5p inhibitor (miR-inhibitor) and corresponding negative control (NC inhibitor), miR-525-5p mimic and blank control (NC mimic) were purchased from RiboBio Co., Ltd. (Guangzhou, China). After transfection for 48 h, the cells were collected. ARHGAP11A overexpression (ARHGAP11A) and pLent vectors (oe-NC) were purchased from Vigene Biosciences (USA). According to the manufacturer’s instructions, the transfection was performed with Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA).
qRT-PCR
Total RNA was isolated from cells or tissue using Trizol reagent (Invitrogen, USA). The content and quality of RNA were evaluated using Nanodrop ND-2000 spectrophotometer (Thermo Scientific™, USA). In order to evaluate the expression of LINC01207 and ARHGAP11A, RevertAid First Strand cDNA Synthesis Kit (Thermo Scientific™, USA) was used to reverse transcribe 2
Fluorescence in situ hybridization (FISH)
FISH was performed using a FISH probe synthesized by RiboBio Co., Ltd, and the FISH Kit (RiboBio Co., Ltd, Guangzhou, China) was used according to the manufacturer’s instructions. The cells on the cover glass were washed with PBS and fixed with 4% paraformaldehyde for 10 min at room temperature, and then infiltrated with PBS containing 0.5% Triton X-100 at 4
Cell counting kit-8 assay (CCK-8)
CCK-8 (Beyotime, Shanghai, China) was used to detect cell proliferation. The cells were inoculated into 96 well plates with a cell concentration of 5
Transwell assay
Transwell inserts (8
Flow cytometry
To analyze cell apoptosis, the cells were digested with trypsin and then resuspended in the dark in a binding buffer containing Annexin V-FITC (BD Biosciences, San Jose, CA, USA) and propidium iodide (PI; BD Biosciences) for 15 min. Flow cytometry (BD Biosciences) was used to analyze the stained cells.
Dual-luciferase reporter gene assay
The 3’-untranslated region (3’-UTR) sequence of LINC01207 or ARHGAP11A was amplified from human genomic DNA. Then these sequences were subcloned into pGL3 luciferase reporter vector (Promega, Madison, WI, USA), respectively. The potential miR-525-5p binding site was mutated by Quick Change Site-directed Mutagenesis Kit (Agilent Technologies, Santa Clara, CA, USA). WT (MUT) of LINC01207 vector or ARHGAP11A vector and NC mimic or miR-525-5p mimic were cotransfected into NSCLC cells, respectively. After transfection for 48 h, both renilla luciferase activity and firefly luciferase activity were measured by the DualLucy Assay Kit (Promega, Madison, WI, USA).
RNA binding protein immunoprecipitation (RIP)
Magna RNA immunoprecipitation (RIP) Kit (Millipore, Billeria, USA) was used to conduct RIP assay. The cells were transfected with LINC01207 and miR-525-5p and resuspended in the lysate buffer. Then 100
Western blot
The cell lysates were extracted by Radio-Immunopre- cipitation Assay (RIPA) lysis buffer (Beyotime, P.R., China), and the protein concentration was determined by BCA Protein Determination Kit (Beyotime, P.R., China). After denatured at high temperature, the proteins were separated from the sample buffer by sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE), transferred to PVDF membrane (Millipore), sealed with 5% albumin from bovine serum (BSA) for 2 h, and then incubated with primary antibodies at 4
Animal assay
In order to confirm the role of LINC01207 in promoting the proliferation of NSCLC cells in vivo, subcutaneous tumor mouse models were established. Twelve male BALB/c nude mice (4 weeks old) were randomly divided into two groups, with 6 in each group. The stable low-expression group of SPC-A1 cells transfected with LINC01207 lentivirus and negative control group were washed with PBS and injected subcutaneously into the axillary region on either side of nude mice. The tumor size was evaluated by the caliper once a week. The following formula was used to estimate the tumor size: Volume
The expression and localization of LINC01207 in NSCLC tissue and cells. A: Volcano plot of differentially expressed genes in GSE101929 microarray, X-axis represents-log10p value, and Y-axis represents logFC value. Red dots represent genes with significantly high expression in tumor and green dots represent genes with significantly low expression in tumor; B: The expression of LINC01207 in NSCLC tumor tissue and matched adjacent tissue; C: The expression of LINC01207 in normal pulmonary bronchial epithelial cell lines and NSCLC cell lines was detected by qRT-PCR; D: The localization of LINC01207 in HCC827 and SPC-A1 cell lines was detected by FISH (400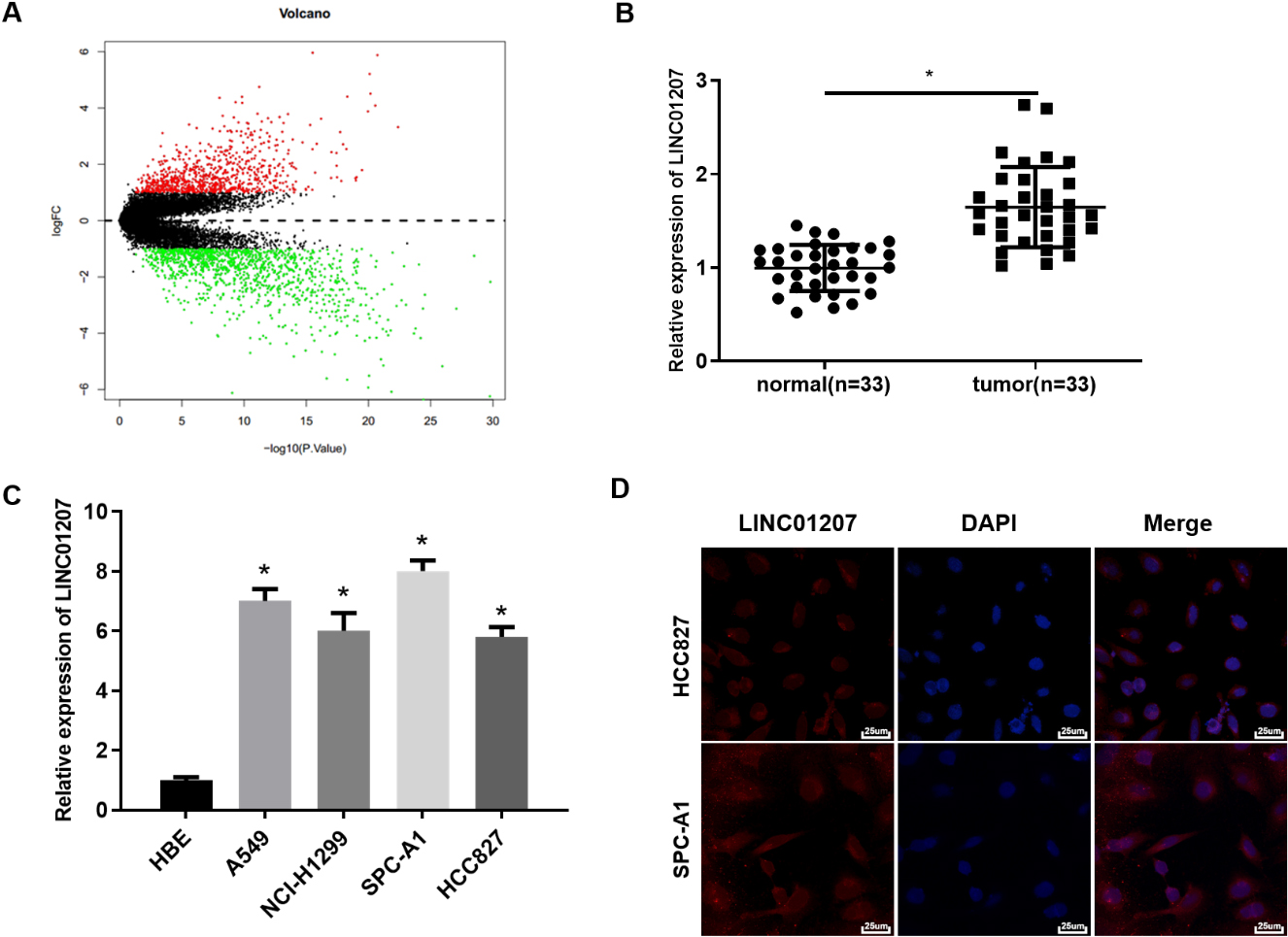
Immunohistochemical staining was conducted on paraffin-embedded tumor tissue of nude mice. By streptavidin-peroxidase coupling method, rabbit anti-ARHGAP11A antibody (Abcam, Cambridge, UK) and Ki67 antibody (1:500, ab15580; Abcam, Cambridge, UK) were used as primary antibodies and goat anti-rabbit IgG H&L (Abcam, Cambridge, UK) was used as secondary antibody for IHC detection. The tissue sections were observed under a microscope at 200
Statistical analysis
SPSS 22.0 (IBM Corp. Armonk, NY, USA) and Graphpad Prism 6.0 Software (Graphpad Inc., San Diego, CA, USA) were used for statistical analysis. All measured data were expressed as mean
Results
LINC01207 is up-regulated in NSCLC tissue and cells
A total of 1967 lncRNAs with significant difference were finally obtained through bioinformatics methods (Fig. 1A). Among these DElncRNAs, 7 lncRNAs were significantly up-regulated in NSCLC(Supplementary Table 3), of which LINC01207 was the most up-regulated. Furthermore, the expression of LINC01207 in 33 pairs of NSCLC tissue and matched adjacent tissue was analyzed by qRT-PCR. It was found that the expression of LINC01207 in NSCLC tissue was significantly higher than that in matched adjacent tissue (Fig. 1B). Moreover, the expression of LINC01207 in NSCLC cell lines was tested as well. The results showed that compared with HBE cells, LINC01207 was significantly up-regulated in NSCLC cell lines A549, NCI-H1299, SPC-A1 and HCC827 (Fig. 1C). Next, SPC-A1 cells and HCC827 cells were used for subsequent assays. Because the subcellular localization of lncRNA is closely related to its biological function and potential molecular mechanism, lncRNA may play the role of ceRNA in cytoplasm [13]. Therefore, the subcellular localization of LINC01207 was detected by RNA-FISH. The result demonstrated that majority of LINC01207-positive cells were mainly located in the cytoplasm and a few in the nucleus (Fig. 1D). These results manifested that LINC01207 was highly expressed in NSCLC tissue and cells, and may exert ceRNA function.
LINC01207 knockdown suppresses the proliferation, migration and invasion of NSCLC cells, and facilitates cell apoptosis. A: The expression of LINC01207 in HCC827 and SPC-A1 cells was tested by qRT-PCR; B: The proliferation of HCC827 and SPC-A1 cells after transfection with sh-LINC01207 was detected by CCK-8 assay; C: The migration and invasion of HCC827 and SPC-A1 cells after transfection with sh-LINC01207 was detected by Transwell assay; D: The cell apoptosis of HCC827 and SPC-A1 cells after sh- transfection with LINC01207 was detected by flow cytometry; * Indicates 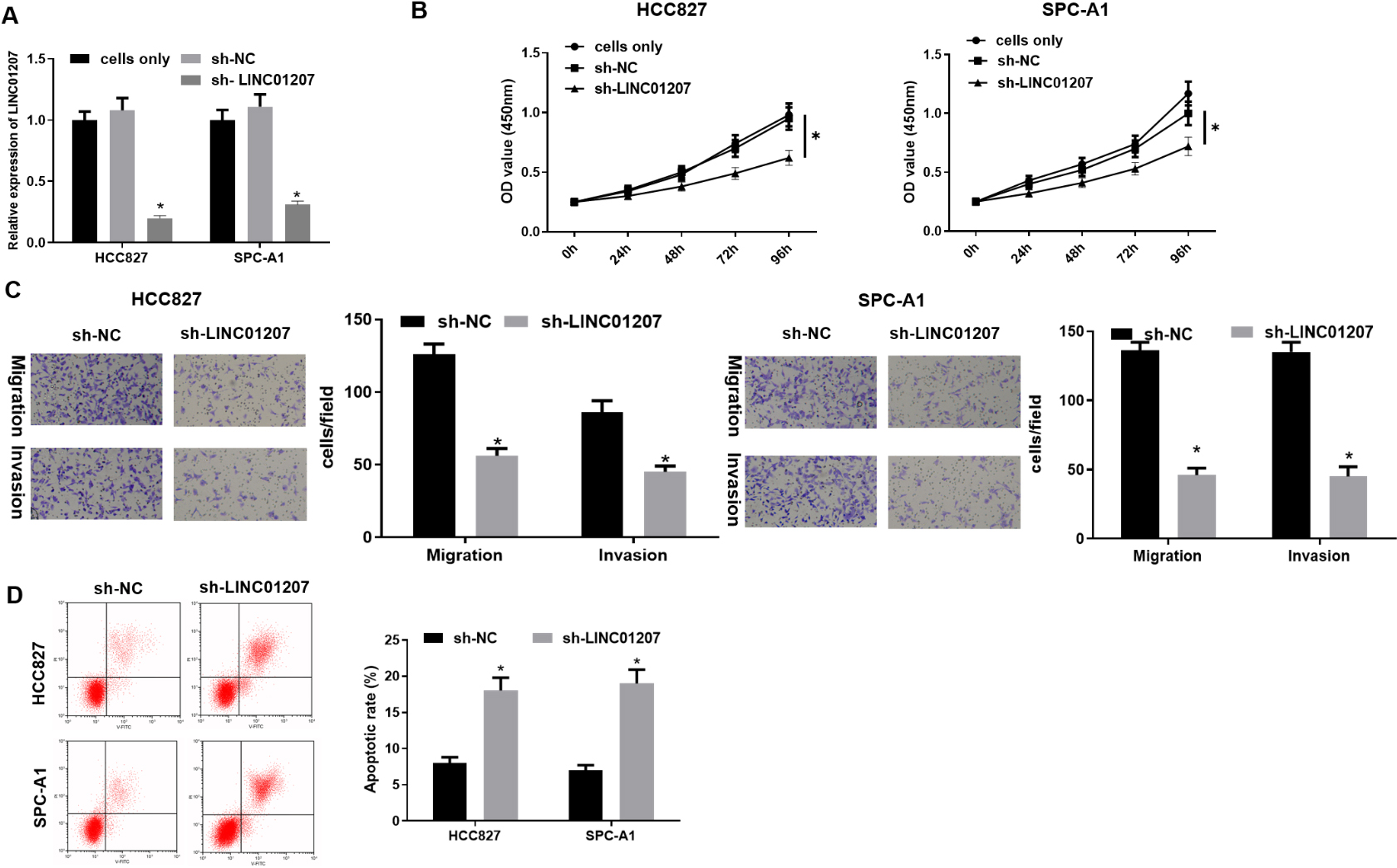
To assess the biologic role of LINC01207 in NSCLC, sh-NC and sh-LINC01207 were transfected into HCC827 cells and SPC-A1 cells. It was found that when sh-NC was transfected, the expression of LINC01207 was no different from the cells only group without any treatment, while the expression of LINC01207 in the sh-LINC01207 group was remarkably down-regulated (Fig. 2A). CCK-8 was used to detect cell proliferation. The result manifested that cell proliferative ability of the sh-NC group was similar to that of the blank group, while LINC01207 knockdown repressed cell proliferation (Fig. 2B). Therefore, the basic biological behaviors of our cells were not affected by transfecting sh-NC. Then, the effect of LINC01207 inhibition on cell migration and invasion was detected by Transwell assay. The result showed that the migration and invasion number of cells after LINC01207 knockdown was significantly reduced (Fig. 2C). What’s more, flow cytometry assay detected the changes in cell apoptosis. Flow cytometry analysis using Annexin V/PI double staining showed that the percentage of apoptotic cells in sh-LINC01207 group was significantly higher than that in control group (Fig. 2D). Therefore, LINC01207 knockdown could hamper the proliferation, migration and invasion of NSCLC cells, and promote cell apoptosis.
LINC01207 inhibits the expression of miR-525-5p in NSCLC cells
Some studies showed that LINC01207 can promote the development of tumor and play a regulatory role through the mechanism of ceRNA [11, 12]. In order to further understand the regulatory mechanism of LINC01207 in NSCLC cells, the downstream miRNAs of LINC01207 were predicted through RNA22 database. Meanwhile, miRNAs with significantly differential expression in GSE29248 and GSE102286 microarrays (Fig. 3A) were included, and miR-525-5p was obtained by intersecting the predicted miRNAs with the DEmiRNAs. Both GSE29248 and GSE102286 microarrays showed that miR-525-5p was significantly lowly expressed in NSCLC (Fig. 3B, C). The expression of miR-525-5p in 33 tumor tissue and matched adjacent tissue, and it was found that miR-525-5p was lowly expressed in tumor tissue (Fig. 3D) and negatively correlated with the expression of LINC01207 (Fig. 3E). By analyzing the binding sites through RNA22 database, it was found that complementary binding sites existed between LINC01207 and miR-525-5p (Fig. 3F).
LINC01207 inhibits the expression of miR-525-5p. A: The intersection between the predicted downstream miRNAs of LINC01207 and the DEmiRNAs in GSE29248 and GSE102286 microarrays, and the middle part represents the intersection of the three groups of data; B, C: The differential expression of miR-525-p in GSE29248 and GSE102286 microarrays, in which the X-axis represents the sample type and the Y-axis represents the expression level, the gray boxplot represents the normal sample, and the red boxplot represents the tumor sample; D: The expression of miR-525-5p in 33 cases of NSCLC tissue and matched adjacent tissue was detected by qRT-PCR assay; E: The expression relationship between miR-525-5p and LINC01207; F: The binding relationship between LINC01207 and miR-525-5p was detected by dual-luciferase reporter gene assay; G: The expression level of miR-525-5p in HCC827 cells and SPC-A1 cells after sh-LINC01207 was transfected was detected by qRT-PCR assay; H: The targeted binding relationship between miR-525-5p and LINC01207 in HCC827 cell line was detected by dual-luciferase reporter gene assay; I: The enrichment of miR-525-5p and LINC01207 in Ago2 immunoprecipitation was detected by RIP; * Indicates 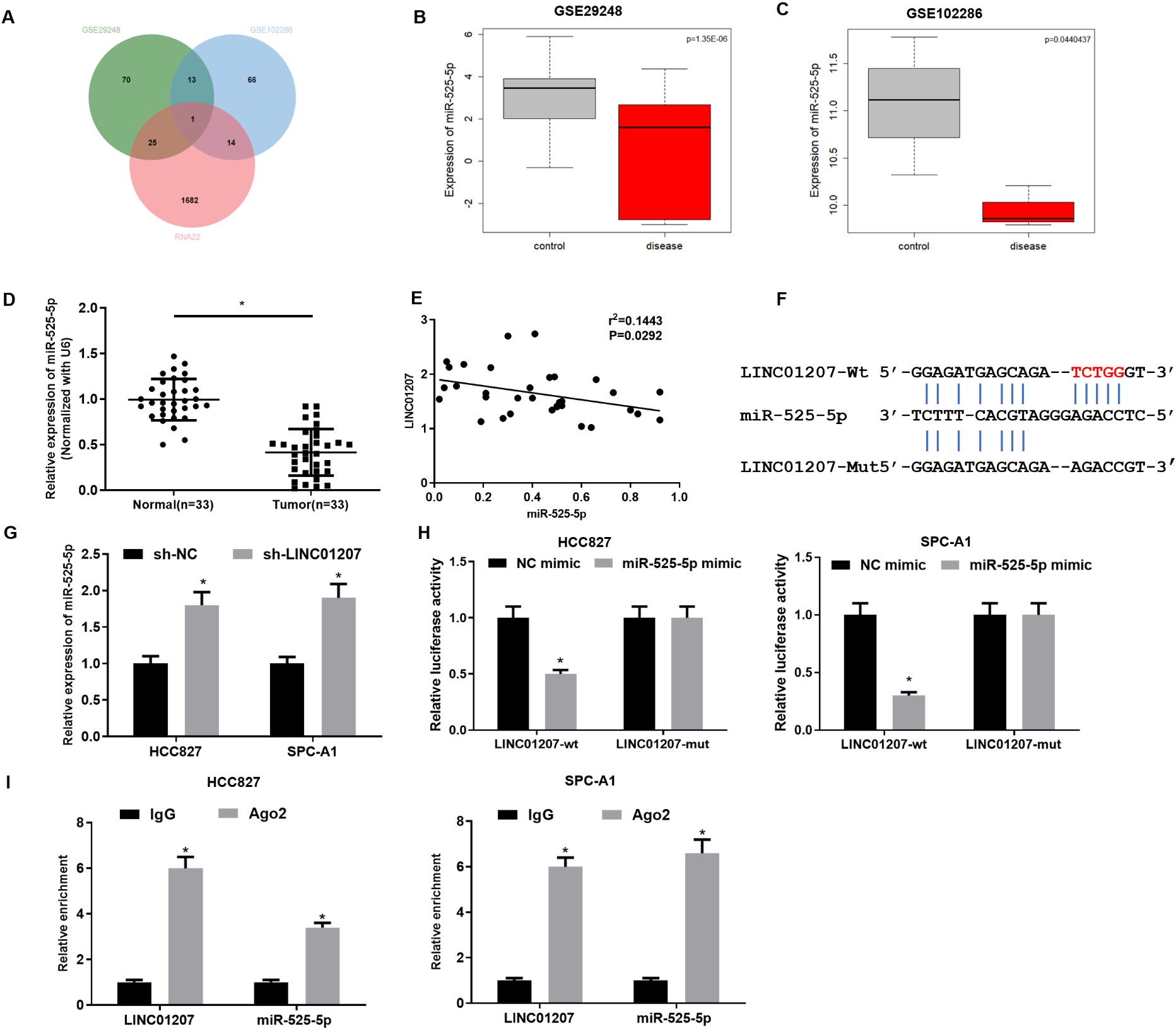
To verify the above prediction, sh-LINC01207 was first transfected into HCC827 and SPC-A1 cells to examine whether the expression of miR-525-5p was affected by sh-LINC01207. qRT-PCR assay showed that the expression of miR-525-5p was significantly up-regulated after sh-LINC01207 was transfected into cells (Fig. 3G). Then, LINC01207-WT and LINC01207-MUT reporter vectors were constructed for dual-luciferase reporter gene detection. The results showed that the luciferase activity of LINC01207-WT was significantly decreased after miR-525-5p was overexpressed, while the luciferase activity of cells containing LINC01207-MUT vector was not affected (Fig. 3H). RIP assay showed that miR-525-5p and LINC01207 were enriched in HCC827 and SPC-A1 cells after immunoprecipitation with Ago2 antibody (Fig. 3I), indicating the binding relationship between miR-525-5p and LINC01207. In conclusion, our assays showed that there was a binding relationship between LINC01207 and miR-525-5p in NSCLC.
MiR-525-5p inhibition reverses the effect of LINC01207 knockdown on NSCLC cells. A: Cell proliferation potential in sh-NC 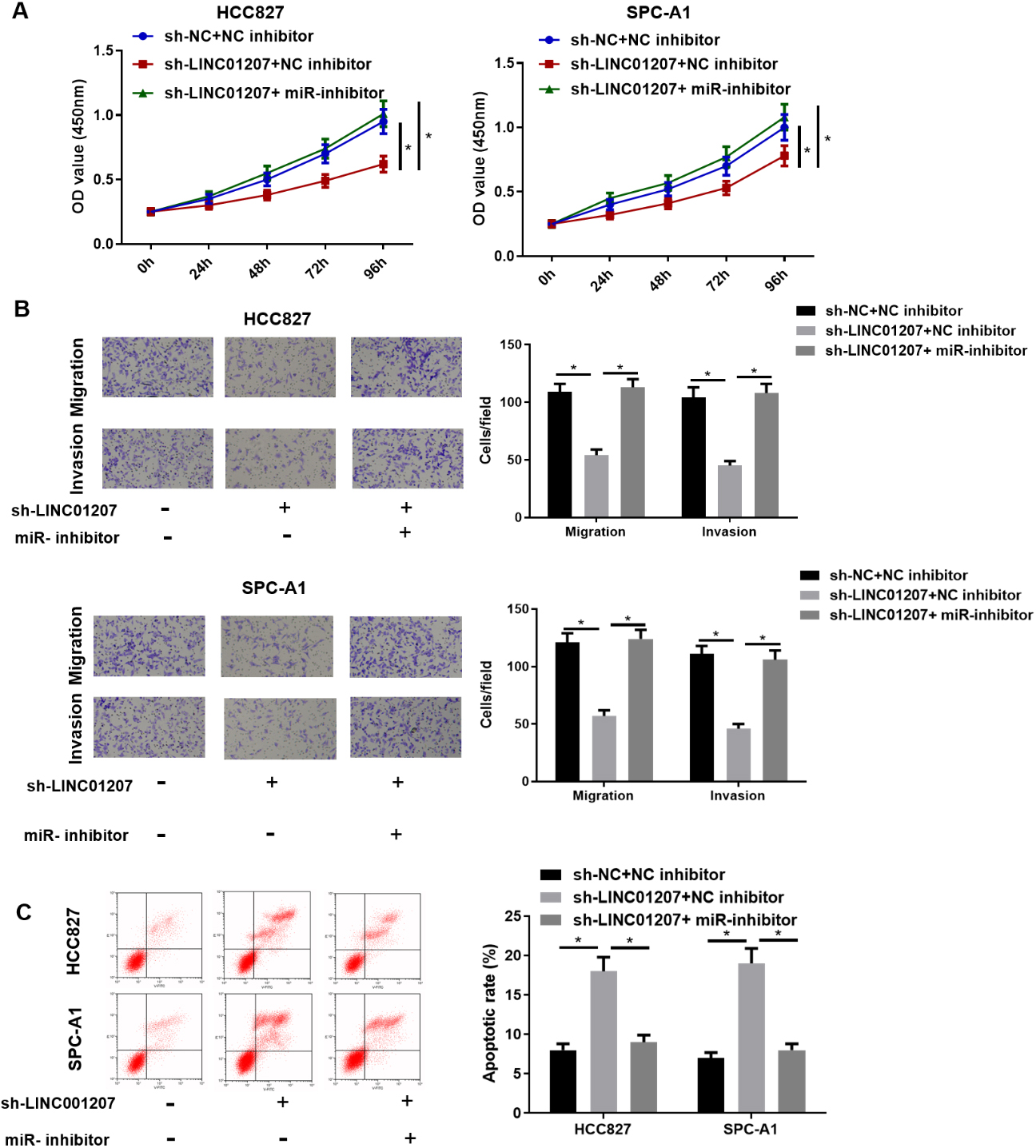
LINC01207 targets ARHGAP11A by sponging miR-525-5p. A: The intersection between the predicted target mRNAs of miR-525-5p and significantly up-regulated mRNAs in the GSE44077 microarray, with the middle part representing the intersection of the data in four groups; B: The differential expression of three candidate genes in GSE44077, green boxplot for normal samples and red boxplot for tumor samples; C: The binding sites of miR-525-5p and ARHGAP11A were predicted by TargetScan; D: The targeted relationship between miR-525-5p and ARHGAP11A was detected by dual-luciferase reporter gene assay; E: The enrichment of ARHGAP11A in Ago2 immunoprecipitation was detected by RIP assay; F: The expression of LINC01207, miR-525-5p and ARHGAP11A in each treatment group was detected by qRT-PCR assay; G: The protein expression of ARHGAP11A in each treatment group was detected by Western blot assay. * Indicates 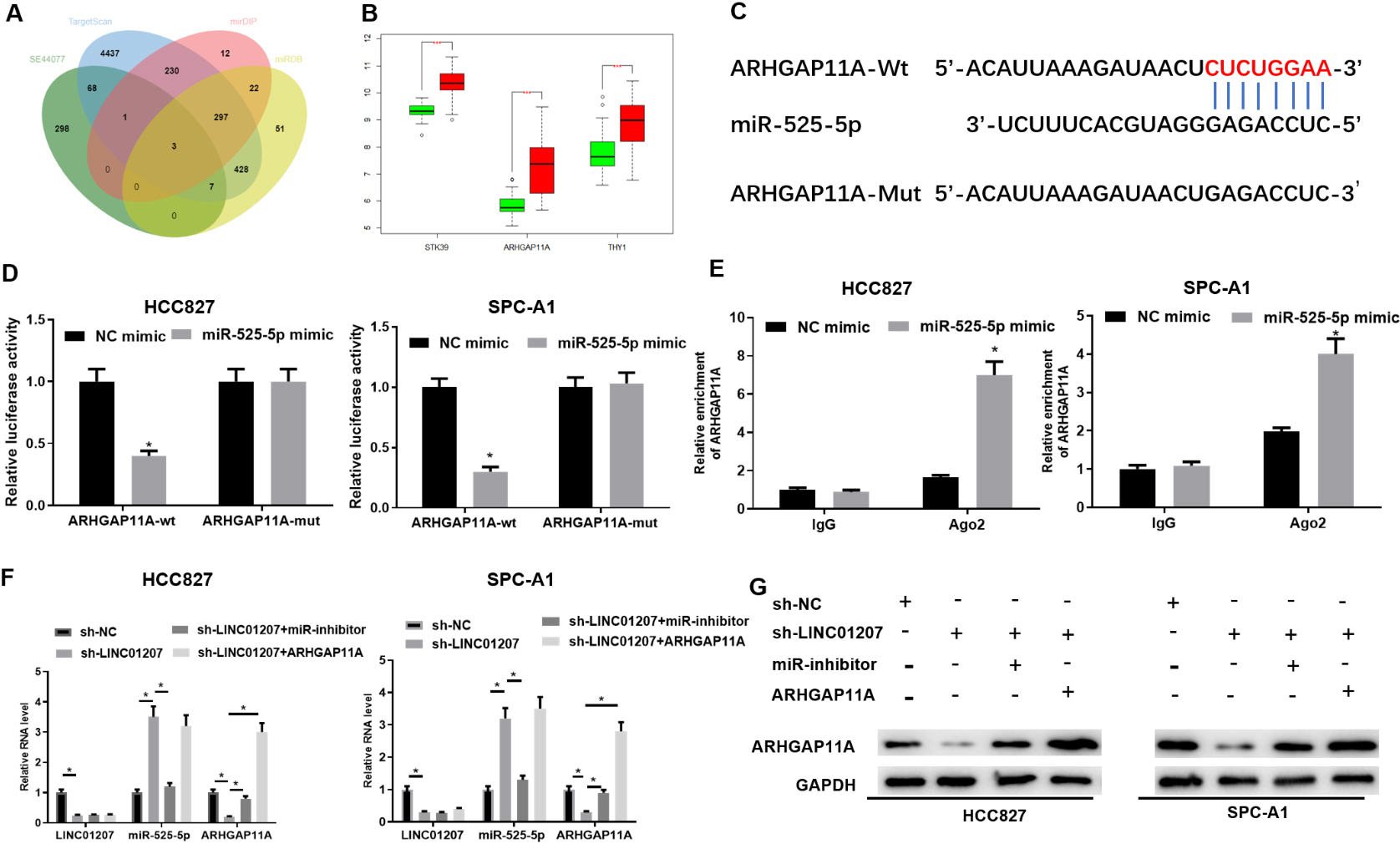
In order to investigate whether LINC01207 regulated the malignant phenotype of NSCLC cells via regulating miR-525-5p, sh-NC
LINC01207 silencing inhibits NSCLC cell growth by regulating miR-525-5p/ARHGAP11A axis. A: The proliferation potential of HCC827 and SPC-A1 cells was detected by CCK-8 assay; B: The migration and invasion of HCC827 and SPC-A1 cells were detected by Transwell assay (100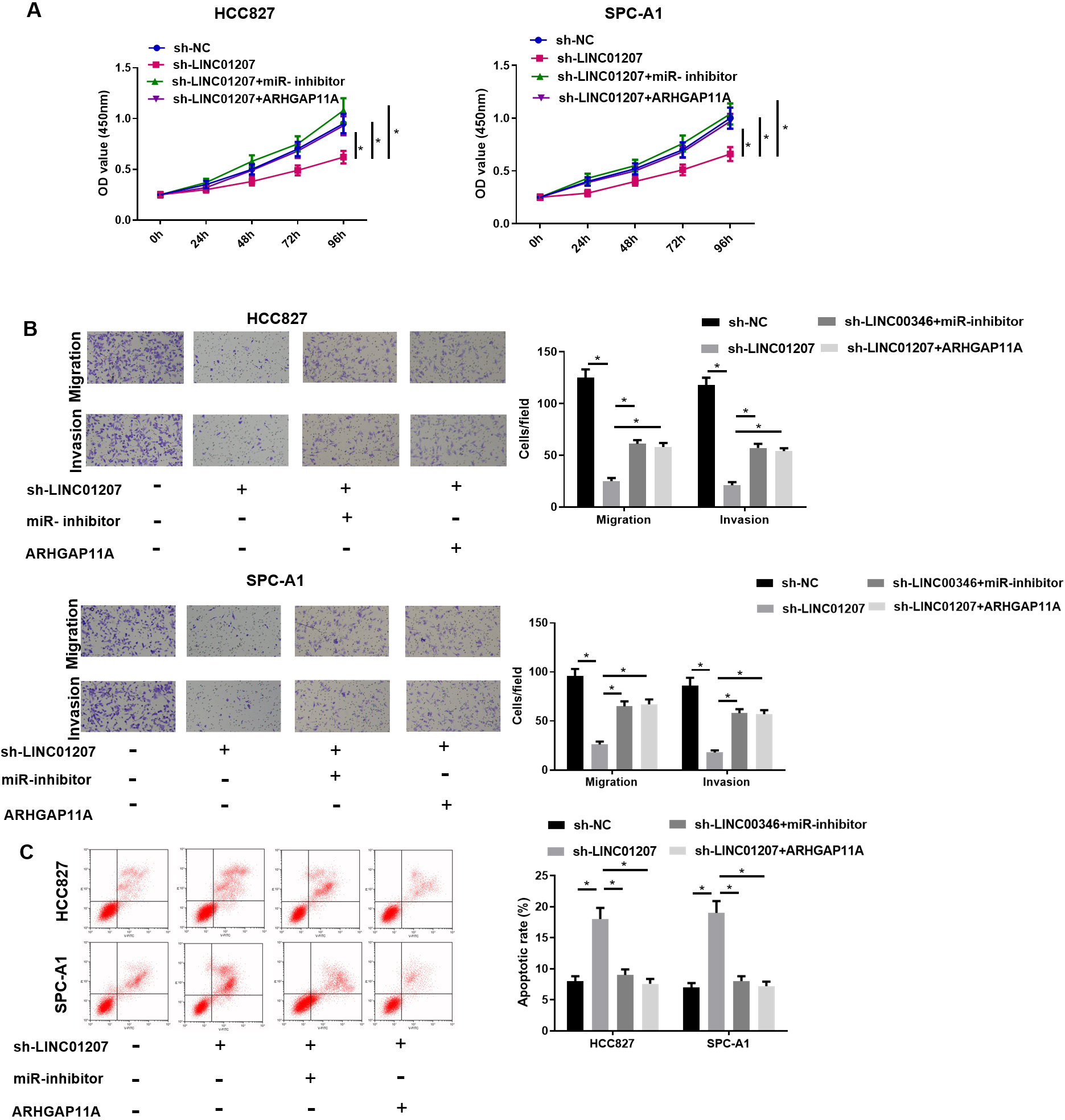
LINC01207 knockdown inhibits the progression of NSCLC A: The tumor. images and tumor weight of nude mice in each treatment group were analyzed after 35 days; B: Tumor growth curves of nude mice in each treatment group; C: The expression of LINC01207, miR-525-5p and ARHGAP11A in tumor tissue in each treatment group was detected by qRT-PCR assay; D: The number of Ki-67-positive cells in xenogeneic tumor tissue was detected by immunohistochemistry (200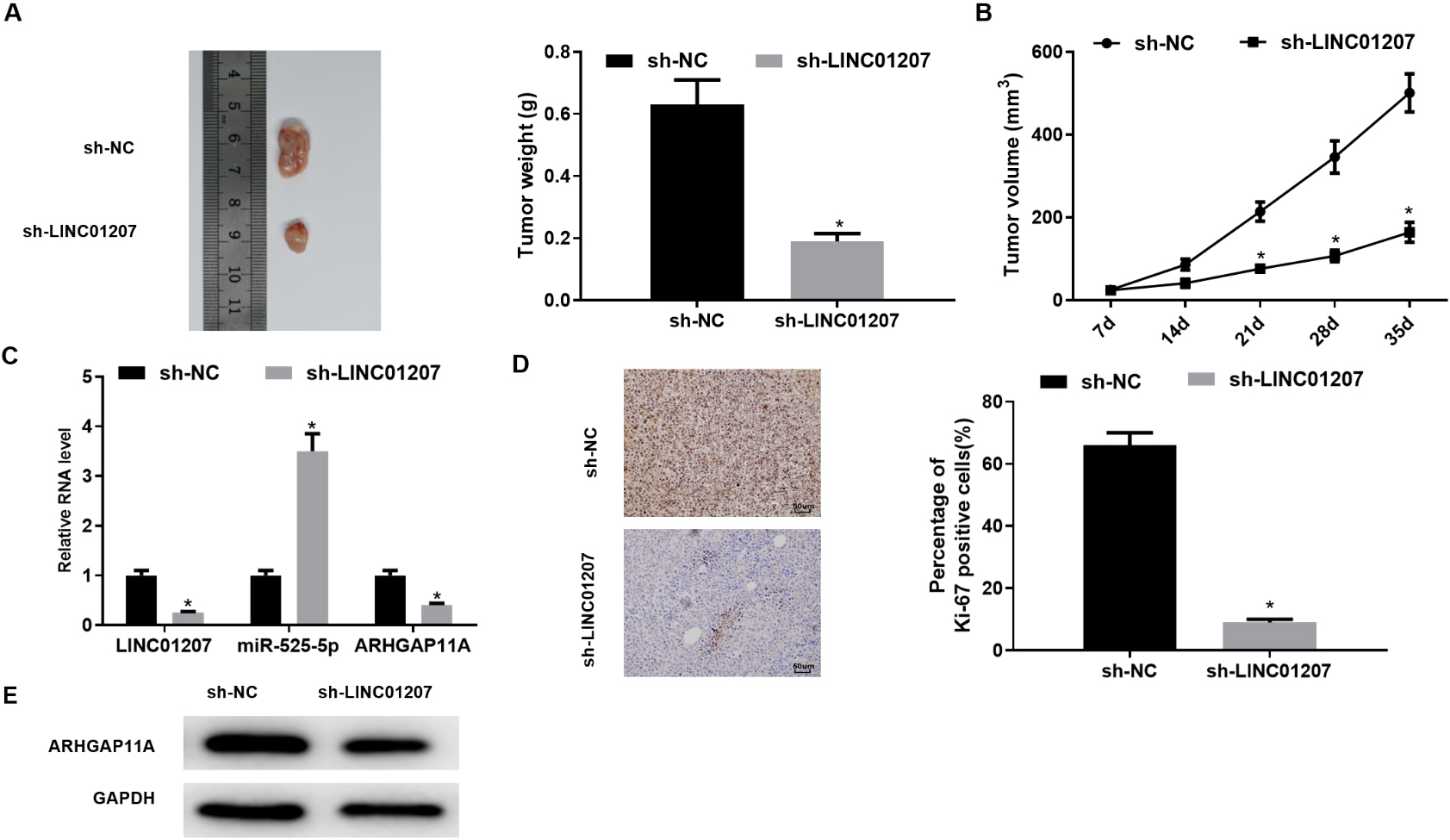
The downstream target mRNAs of miR-525-5p were predicted by TargetScan and other databases, and the highly expressed mRNAs in GSE44077 microarray were intersected with the predicted results to narrow the range of candidate genes (Fig. 5A). Three DEmRNAs that had specific binding sites with miR-525-5p were obtained. The expression of these three candidate genes in GSE44077 was analyzed (Fig. 5B), and it was found that the expression change of ARHGAP11A was the largest, and its expression difference was extremely significant. These results and previous reports suggested that LINC01207 could affect the expression of ARHGAP11A via miR-525-5p through the mechanism of ceRNA, and finally regulated the progression of NSCLC. The targeted sites of ARHGAP11A and miR-525-5p were predicted through bioinformatics website TargetScan (Fig. 5C), and the binding relationship between ARHGAP11A and miR-525-5p was verified by dual-luciferase reporter gene assay and RIP assay. Compared with the control group, the luciferase activity in SPC-A1 cells co-transfected with ARHGAP11A-WT and miR-525-5p-mimic was significantly decreased. However, the luciferase activity of the cells transfected with ARHGAP11A-MUT and miR-525-5p mimic did not change significantly (Fig. 5D). RIP results showed that ARHGAP11A was significantly enriched in the Ago 2 immunoprecipitation of miR-525-5p group (Fig. 5E) after Ago 2 protein was pulled down. This indicated that ARHGAP11A could directly bind to miR-525-5p in HCC827 and SPC-A1 cells. Next, to better validate the regulatory role of LINC01207 and miR-525-5p in ARHGAP11A expression, ARHGAP11A mRNA expression in treated cells was assessed. It was found that transfection of ARHGAP11A overexpression notably up-regulated ARHGAP11A expression(Supplementary Figure 1). By ensuring the good transfection efficiency, we performed the following experiments. It was found that when LINC01207 was knocked down, the expression of miR-525-5p was significantly up-regulated, while the mRNA and protein levels of ARHGAP11A were significantly down-regulated by qRT-PCR and Western blot. Furthermore, LINC01207 knockdown and miR-525-5p inhibition or ARHGAP11A overexpression partially reversed the effect of LINC01207 knockdown on the expression level of ARHGAP11A (Fig. 5F, G), indicating that LINC01207 and ARHGAP11A competitively bound with miR-525-5p. In conclusion, these results indicated that LINC01207 and miR-525-5p regulated the expression of ARHGAP11A in NSCLC cells.
LINC01207 acts as a ceRNA and regulates the growth of NSCLC cells via targeting ARHGAP11A through miR-525-5p
In order to study whether LINC01207 exerted its oncogenic function by targeting ARHGAP11A via miR-525-5p, multiple transfection groups with differential expression were constructed for verification. The results showed that LINC01207 knockdown inhibited the proliferation, migration and invasion of HCC827 and SPC-A1 cells, and facilitated cell apoptosis. On the basis of LINC01207 knockdown, miR-525-5p inhibition or ARHGAP11A overexpression reversed the effects of LINC01207 knockdown on HCC827 and SPC-A1 cells (Fig. 6A–C). These findings suggested that LINC01207 hastened the progression of NSCLC by regulating miR-525-5p/ARHGAP11A axis.
LINC01207 knockdown inhibits the progression of NSCLC in vivo
In order to further analyze the effect of LINC01207 on NSCLC, mice xenograft models in vivo were established. After 35 days, it was found that compared with sh-NC group, the tumor size and weight of sh-LINC01207 group were significantly reduced (Fig. 7A and B). Then the expression of LINC01207, miR-525-5p and ARHGAP11A in tumor tissue was evaluated by qRT-PCR assay. The results confirmed that LINC01207 and ARHGAP11A were significantly lowly expressed in sh-LINC01207 group compared with control group, and miR-525-5p was significantly highly expressed in sh-LINC01207 group (Fig. 7C). Ki-67 immunostaining showed that the percentage of Ki-67-positive cells in the subcutaneous tumors with LINC01207 knockdown was significantly decreased compared with that in the control group (Fig. 7D). Western blot assay showed that LINC01207 silencing reduced the expression of ARHGAP11A in tumor tissue (Fig. 7E). These results suggested that LINC01207 knockdown inhibited the progression of NSCLC in vivo.
Discussion
Lung cancer is one of the most serious malignant tumors with incidence and mortality ranking first in the world. Because of the atypical symptoms and signs of early stage lung cancer, the detection rate is very low [14]. Many patients are not diagnosed as lung cancer in time as the early-stage symptoms are not obvious, leading to the diagnosis in late stage or cancer cell metastasis [15]. Hence, it is vital to find specific targets for NSCLC. Previous studies manifested that lncRNA TRERNA1 is up-regulated in NSCLC tissue and cell lines, and its expression is associated with pathologic stage of NSCLC and dismal outcomes, which fosters the malignant progression of NSCLC [16]. As such, lncRNA-XIST is aberrantly highly expressed in NSCLC tissue and cells, downregulation of which represses NSCLC development by activating miR-335/SOD2/ROS signal pathway mediated pyroptotic cell death [17]. Accordingly, seeking out aberrantly highly-expressed lncRNAs in NSCLC may bring a new turn for NSCLC treatment. In this study, LINC01207 which was significantly up-regulated in NSCLC was identified based on gene expression profile analysis, and the effect of LINC01207 knockdown on the biological functions of NSCLC in vitro and in vivo was investigated. It was found that LINC01207 knockdown inhibited the progression of NSCLC cells. In vivo assays showed that LINC01207 knockdown suppressed the tumor growth of NSCLC, suggesting that LINC01207 may play a role as an oncogene.
LncRNAs regulate the functions of mRNAs by regulating downstream miRNAs in cancer cells [18]. For instance, Yu et al. [19] proved that lncRNA PTAR facilitates cell progression in NSCLC through sponging miR-101. Zhang et al. [20] believed that lncRNA FENDRR regulates TIMP2 expression by binding to miR-761, thereby suppressing the malignant progression of NSCLC. In this study, in order to find the downstream miRNAs of LINC01207 in NSCLC, bioinformatics analysis, dual-luciferase reporter gene assay and RIP analysis were conducted, and it was confirmed that LINC01207 was the molecular sponge of miR-525-5p. miR-525-5p is differentially expressed in many cancers, such as cervical cancer [21] and laryngeal squamous cell carcinoma [22]. It was found that the expression of miR-525-5p was abnormal in NSCLC through assays. In addition, it was also demonstrated that miR-525-5p inhibition could reverse the inhibitory effect of LINC01207 knockdown on the progression of NSCLC cells. It has been reported that miR-525-5p can inhibit the metastasis of cervical cancer by blocking UBE2C/ZEB1/2 axis [21]. MiR-525-5p can also affect the progression of laryngeal squamous cell carcinoma by targeting PIK3R1 [22]. Nevertheless, the downstream targets and the specific regulatory mechanism of miR-525-5p in NSCLC need to be further studied.
In this study, it was noted that ARHGAP11A was a downstream target of miR-525-5p and was highly expressed in NSCLC tissue, which is consistent with the study on basal like breast cancer [23]. Highly-expressed ARHGAP11A in hepatocellular carcinoma facilitates cell proliferation, invasion, migration, and EMT in vitro [24]. Besides, a study [25] illustrated that increased ARHGAP11A in lung adenocarcinoma results in poor patient prognosis. In this investigation, further exploration on molecular mechanism found that LINC01207 affected the post-transcriptional control mechanism of miR-525-5p. LINC01207 as a sponge of miRNA weakened the binding of miR-525-5p and ARHGAP11A to decrease miR-525-5p and increase ARHGAP11A, thereby exerting an oncogenic effect (Fig. 8). It was also verified that miR-525-5p inhibition or ARHGAP11A overexpression could reverse the effects of sh-LINC01207 on proliferation, migration, invasion and apoptosis of HCC827 and SPC-A1 cells. Therefore, it was suggested that LINC01207 regulated ARHGAP11A and promoted the progression of NSCLC via sponging miR-525-5p.
The molecular mechanism of LINC01207 regulates NSCLC through miR-525-5p/ARHGAP11A axis.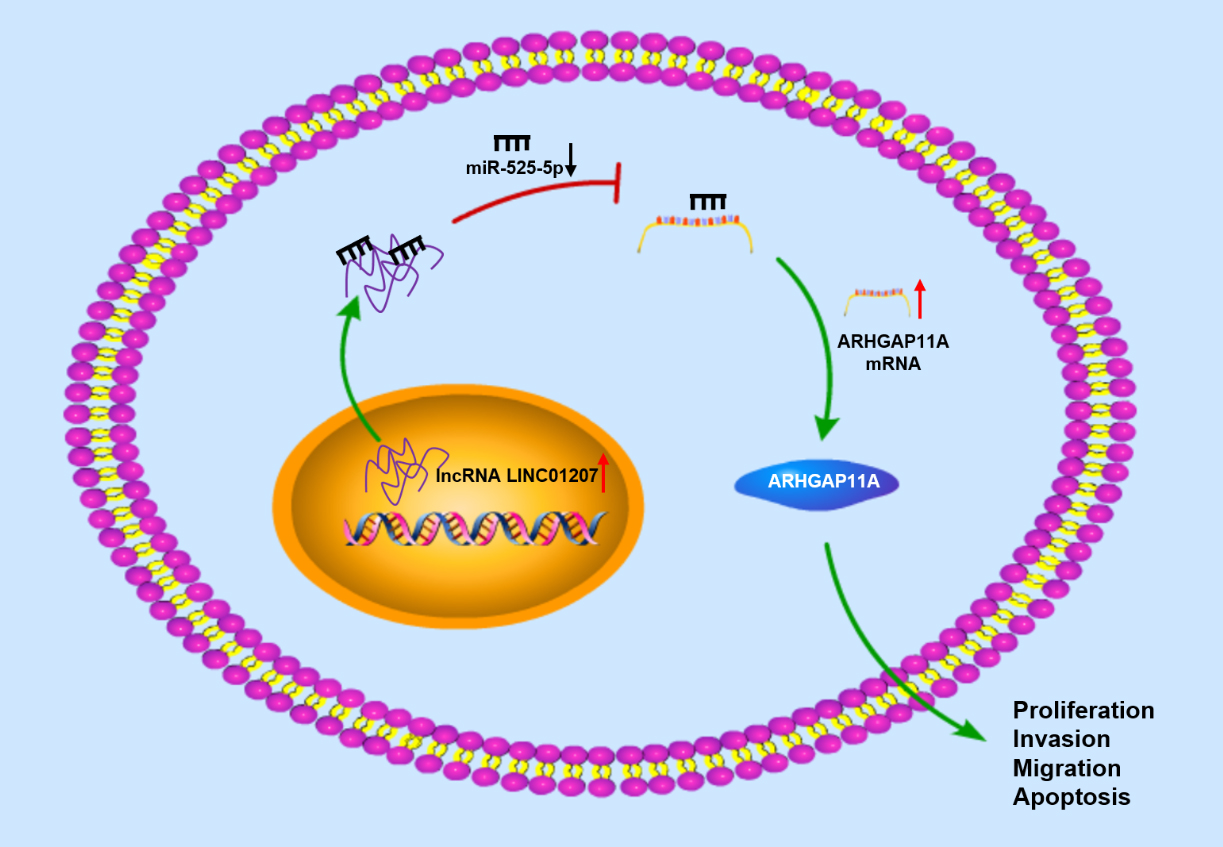
In conclusion, it could be determined that LINC01207 played an important role in NSCLC as an oncogene. Our study verified for the first time that LINC01207/miR-525-5p/ARHGAP11A axis regulated and promoted the progression of NSCLC. This study may offer clues for elucidating the tumorigenesis of NSCLC, and may offer the theoretical basis for developing new diagnosis and treatment methods of NSCLC. However, there are still some limitations in this study. For instance, we did not investigate the downstream signaling pathway that ARHGAP11A may affect. In the future, we will further fulfill the regulatory network of this project to provide a more solid theoretic basis for NSCLC treatment.
Ethics approval and consent to participate
Not applicable.
Consent for publication
All authors consent to submit the manuscript for publication.
Availability of data and materials
The data used to support the findings of this study are included within the article. The data and materials in the current study are available from the corresponding author on reasonable request.
Funding
Not applicable.
Authors’ contributions
Conception: Bin Zhang.
Interpretation or analysis of data: Bin Zhang.
Preparation of the manuscript: Bin Zhang, Zhou Jin.
Revision for important intellectual content: Zhou Jin.
Supervision: Hao Zhang.
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-203197.
sj-docx-1-cbm-10.3233_CBM-203197.docx - Supplemental material
Supplemental material, sj-docx-1-cbm-10.3233_CBM-203197.docx
Footnotes
Acknowledgments
Not applicable.
Conflict of interest
The authors declare that they have no potential conflicts of interest.
