Abstract
BACKGROUND:
Circulating miRNAs are promising biomarkers for detection of various cancers. As a “developmental” disorder, cancer showed great similarities with embryos.
OBJECTIVE:
A comprehensive analysis of circulating miRNAs in umbilical cord blood (UCB) and pan-cancers was conducted to identify circulating miRNAs with potential for cancer detection.
METHODS:
A total of 3831 cancer samples (2050 serum samples from 15 types of cancers and 1781 plasma samples from 13 types of cancers) and 248 UCB samples (120 serum and 128 plasma samples) with corresponding NCs from Chinese populations were analyzed via consistent experiment workflow with Exiqon panel followed by multiple-stage validation with qRT-PCR.
RESULTS:
Thirty-four serum and 32 plasma miRNAs were dysregulated in at least one type of cancer. Eighteen serum and 16 plasma miRNAs were related with embryos. Among them, 9 serum and 8 plasma miRNAs with consistent expression patterns between pan-cancers and UCB were identified as circulating oncofetal miRNAs. Retrospective analysis confirmed the diagnostic ability of circulating oncofetal miRNAs for specific cancers. And the oncofetal miRNAs were mainly up-regulated in tissues of pan-cancers.
CONCLUSIONS:
Our study might serve as bases for the potential application of the non-invasive biomarkers in the future clinical.
Introduction
Cancer is one of leading causes of death worldwide. Despite increased understanding of the molecular and clinical characteristics of cancers as well as improvements in treatment strategies, the prognosis of cancer patients is still poor [1]. Due to lack of obvious symptoms and insufficiently sensitive biomarkers, many patients are diagnosed with advanced stages and do not have a choice of surgery which still remains to be the only treatment strategy to cure the disease. Thus, early detection and intervention is the key strategy for improving the long-term survivals of cancer patients. Nowadays, pathological examination of biopsy is still the gold standard for cancer diagnosis. But the invasiveness, inconvenience and potentially subjective discrepancy caused by different operators limit its effectiveness in screening the diseases in large-scale populations. Existing tumor diagnostic biomarkers, such as carcinoembryonic antigen (CEA), carbohydrate antigen 19-9 (CA19-9), squamous cell carcinoma antigen (SCC) and CYFRA21-1, did not show sufficient sensitivity and specificity in diagnosis of cancers [2, 3, 4]. Hence, more novel and reliable biomarkers to detect the diseases for early intervention are urgently needed.
MicroRNAs (miRNAs) are small non-coding RNAs (18–25 nucleotides in length) which could regulate gene expression by inhibiting translation or encouraging transcript degradation. Hundreds of miRNAs are dysregulated in various cancers and could regulate processes related to all major hallmarks of cancers by targeting various mRNAs [5]. Since Mitchell et al. firstly reported the significance of serum miR-141 in diagnosis of prostate cancer, many studies have demonstrated that miRNAs could be stably detected in circulating plasma/serum, and could act as non-invasive biomarkers for early detection of many types of cancers [6]. But the results of identified circulating miRNAs for the same type of cancers were always heterogeneous among the published studies. Differences in research methods and populations might partly explain the inconformity. Meanwhile, expression profiles of circulating miRNAs in serum and plasma samples were also distinctive [7]. Therefore, we have committed to identifying circulating serum and plasma miRNAs as candidate diagnostic biomarkers for various types of cancers with Chinese populations in the past few years. With the almost identical workflow and appropriate normalization methods, serum miRNAs in esophageal squamous cell carcinoma (ESCC) [8], gastric cardia adenocarcinoma (GCA) [9], gastric non-cardia adenocarcinoma (GNCA) [10], colon adenocarcinoma (COAD), rectum adenocarcinoma (READ) [11], pancreatic adenocarcinoma (PAAD) [12], lung squamous cell carcinoma (LUSC) [13] and ovarian cancer (OV) [14], and plasma miRNAs in ESCC [15], GNCA [16], PAAD [17], LUSC [18], lung adenocarcinoma (LUAD) [19], thyroid cancer (THCA) [20], COAD, READ [21] and OV [22] were confirmed and could aid in cancer detection. As cancer is a disease with great heterogeneity as well as similarity, a pan-cancer analysis to show a global view of circulating miRNAs in various cancers is critical and conducive for us to better apply the potential diagnostic tools. To date, no study has conducted a pan-cancer analysis of circulating miRNAs in cancers. To address this issue, we additionally analyzed circulating miRNAs with serum samples in nasopharyngeal carcinoma (NPC), THCA, LUAD, breast cancer (BRCA), uterine corpus endometrial carcinoma (UCEC), prostate adenocarcinoma (PRAD) and cervical squamous cell carcinoma (CESC), and plasma samples in NPC, BRCA, CESC and UCEC according to the consistent criteria. Thus, with the data we previously reported (re-analyzed with the consistent criteria), the pan-cancer analysis of circulating miRNAs was presented in the study.
More and more studies indicated that embryos and tumors possess many similarities in pathophysiological processes, such as immune escape, angiogenesis, gene expression, invasion characteristic between embryo implantation and neoplasm invasion and metastasis [23, 24]. In addition, some sets of genes, proteins or metabolites shared between embryogenesis and tumorigenesis were applied as biomarkers for diagnosis of cancer, such as oncofetal proteins alpha-fetoprotein (AFP) and carcino-embryonic antigen (CEA). Therefore, it enlightens us to study cancers from the view of developmental biology, which may provide a new clue for cancer detection. Umbilical cord blood (UCB) is a newborn’s blood obtained after delivery from umbilical cord and represents the most accessible source of neonatal blood. In the present study, we comprehensively analyzed circulating miRNAs in UCB and pan-cancers and identified oncofetal circulating miRNAs which showed similar expression patterns in UCB and various types of cancers. The study could promote our deeper understanding of circulating miRNAs in cancers and further identification of novel cancer biomarkers.
Materials and methods
Sample collection and preparation
Peripheral blood samples of 15 types of cancer patients as well as age and gender matched normal control (NC) blood samples, and UCB samples were collected during 2010 and 2018 at First Affiliated Hospital of Nanjing Medical University. Blood samples of cancer patients analyzed in the study were all obtained before any clinical treatment and intervention. NC blood samples were collected from adult healthy donors who conducted routine health checkup. UCB samples were obtained immediately after the fetus was expelled.
The blood samples were put in an ethylene diamine tetra acetic acid (EDTA)-containing tubes (Becton, Dickinson and Company) or SST Advance tubes (Becton, Dickinson and Company) for separation of plasma and serum samples within 12 hours. Cell-free plasma samples were isolated using a two-step centrifugal process of 350 reactive centrifugal force (RCF) for 10 min and 20,000 RCF for 10 min (Beckman Coulter, USA), while serum samples were obtained by centrifugation at 1,500 RCF for 10 min and 12,000 RCF for 2 min after allowing blood to clot for 30 min. All the samples were then stored at
The study was approved and conducted in accordance with the guidelines by medical ethics committee of the First Affiliated Hospital of Nanjing Medical University. Each participant received written informed consent.
Study design
To identify differential expression of circulating miRNAs in different types of cancer patients and UCB samples, plasma/serum samples of each type of cancer and UCB as well as their own age and gender matched NC samples were analyzed independently in according to the experiment workflow (Fig. 1; GCA serum, GNCA serum, OV serum and OV plasma samples were evaluated through training and testing stages after the screening phase). In brief, each independent experiment according to different sample types was designed into four stages: screening phase, training, testing and external validation stage. In the screening phase, 2 to 4 case pool samples VS. 1 NC pool sample (every 10 samples were pooled as 1 pool sample) was applied to initially screen potential differential expressed circulating miRNAs using Exiqon miRCURY-Ready-to-Use PCR-Human-panel-I+II-V1.M (Exiqon miRNA qPCR panel, Vedbaek, Denmark; 168 miRNAs). The process of arrays and analyses was conducted consistent with the previous study [16]. Then, those miRNAs fulfilled the criteria (miRNAs with Ct
In total, circulating samples including NPC (serum: 208 cases VS. 238 NCs; plasma: 200 cases VS. 189 NCs), THCA (serum: 100 cases VS. 96 NCs; plasma: 120 cases VS. 114 NCs), ESCC (serum: 200 cases VS. 188 NCs; plasma: 178 cases VS. 205 NCs), GCA (serum: 102 cases VS. 84 NCs), GNCA (serum: 203 cases VS. 167 NCs; plasma: 133 cases VS. 109 NCs), COAD (serum: 84 cases VS. 138 NCs; plasma: 65 cases VS. 132 NCs), READ (serum: 112 cases VS. 138 NCs; plasma: 74 cases VS. 132 NCs), PAAD (serum: 159 cases VS. 137 NCs; plasma: 216 cases VS. 220 NCs), LUSC (serum: 102 cases VS. 108 NCs; plasma: 102 cases VS. 101 NCs), LUAD (serum: 170 cases VS. 170 NCs; plasma: 141 cases VS. 124 NCs), BRCA (serum: 216 cases VS. 214 NCs; plasma: 257 cases VS. 257 NCs), UCEC (serum: 92 cases VS. 102 NCs; plasma: 93 cases VS. 79 NCs), PRAD (serum: 86 cases VS. 86 NCs), CESC (serum: 108 cases VS. 108 NCs; plasma: 101 cases VS. 91 NCs), OV (serum: 110 cases VS. 116 NCs; plasma: 101 cases VS. 100 NCs) and UCB (serum: 120 cases VS. 120 NCs; plasma: 128 cases VS. 127 NCs) were analyzed in the study. Additionally, the signature miRNAs showed consistent expression patterns between cancer patients and UCB were identified as oncofetal miRNAs.
RNA extraction
Total RNA from 200
Quantitative reverse transcription polymerase chain reaction (qRT-PCR)
Amplification of miRNAs was accomplished using Bulge-Loop
Expression profiles of oncofetal miRNAs in tumor tissues
The expression levels of the identified oncofetal miRNAs were evaluated using miRNA isoform data from the TCGA database (
Statistical analysis
The differential expression levels of circulating miRNAs between cases and NCs were assessed by Mann-Whitney test. Multiple logistic regression analysis was used to set up the miRNA panel. ROC curves and the area under the ROC curve were applied to estimate the diagnostic value of the oncofetal miRNAs in detecting specific cancers. Binary logistic regression analyses with ROC curves were performed to evaluate the combination of oncofetal miRNAs in predicting specific types of cancers. SPSS16.0 software (SPSS Inc., Chicago, IL, USA) was used for statistical analyses. A two-sided
Functional enrichment analysis of the oncofetal miRNAs
MicroRNA Data Integration Portal (mirDIP) version 2 (
Results
Subjects analyzed in the study
According to different sample types, each study (except GCA serum, GNCA serum, OV serum and OV plasma samples) analyzed was divided into four stages to identify dysregulated circulating miRNAs in various cancer patients and UCB compared to NCs (Fig. 1). After the initial screening phase with Exiqon panel, the cases and NCs were divided into three stages: the training stage, the testing stage and the external validation stage. The number of samples in each stage was listed in the Table S1.
The workflow of each study by different cases and sample types analyzed in the study (except GCA serum, GNCA serum, OV serum and OV plasma samples which were evaluated through training and testing stages after the screening phase). UCB: umbilical cord blood; NC: normal control; NTC: no template control.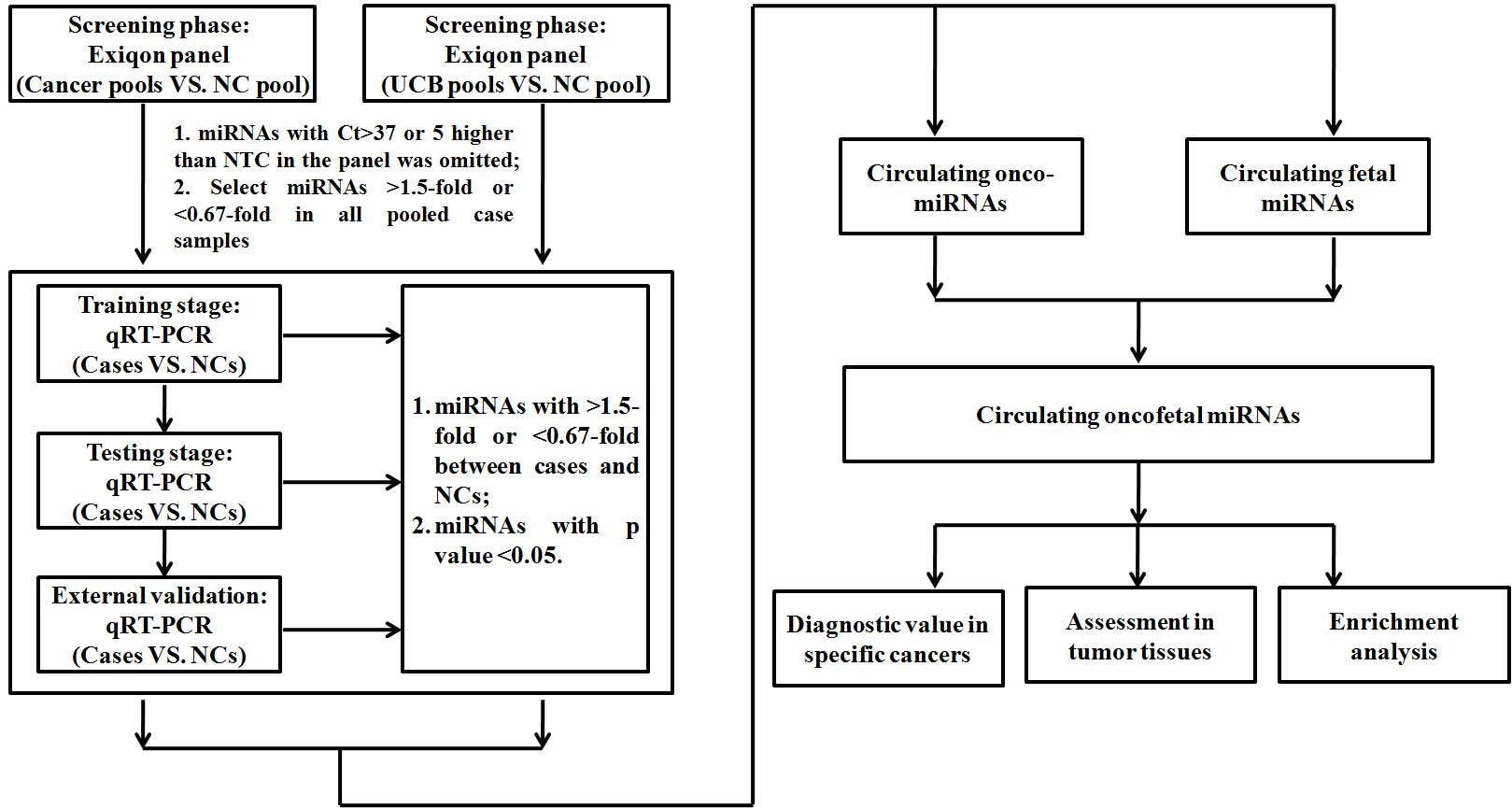
To identify miRNAs with significantly different expression in UCB compared with adult NC, 120 serum and 128 plasma samples isolated from UCB samples were explored respectively in accordance to the experimental workflow. With Exiqon miRNA qPCR panels used in the screening phase, 53 serum miRNAs (42 up-regulated and 11 down-regulated miRNAs; Table S2) and 65 plasma miRNAs (52 up-regulated and 13 down-regulated miRNAs; Table S2) was identified separately in line with the criteria and were subjected to further verification. Consequently, 18 miRNAs including miR-140-5p, miR-144-3p, miR-185-5p, miR-197-3p, miR-19a-3p, miR-19b-3p, miR-20a-5p, miR-20b-5p, miR-210-3p, miR-223-3p, miR-25-3p, miR-382-5p, miR-409-3p, miR-425-5p, miR-483-5p, miR-584-5p, miR-92a-3p and miR-93-5p were confirmed consistently up-regulated in serum samples of UCB compared with adult NC through the training, testing and the external validation stages (Table S3 & Fig. 2; the other miRNAs were shown in Table S4). Meanwhile, sixteen highly expressed plasma miRNAs (miR-106a-5p, miR-130a-3p, miR-140-5p, miR-143-3p, miR-146a-5p, miR-18a, miR-197-3p, miR-20b-5p, miR-210-3p, miR-21-5p, miR-320a, miR-382-5p, miR-409-3p, miR-483-5p, miR-543 and miR-584-5p) were identified in UCB (Table S5 & Fig. 3; the other miRNAs were shown in Table S6).
Expression levels of the eighteen miRNAs in serum of 120 UCB samples and 120 adult NCs. UCB: umbilical cord blood. 
Expression levels of the sixteen miRNAs in plasma of 128 UCB samples and 127 adult NCs. UCB: umbilical cord blood. 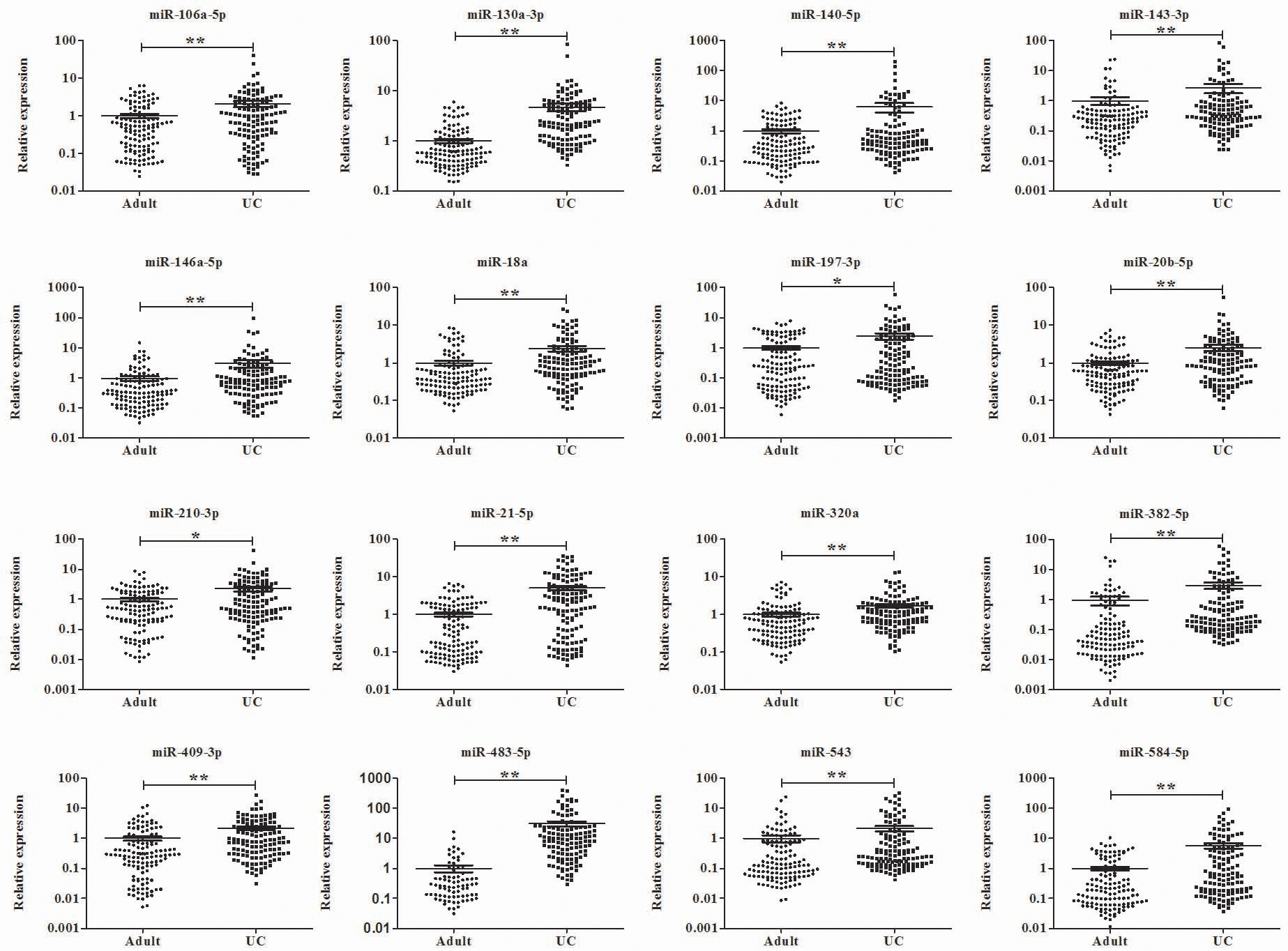
In our previous studies, we demonstrated dysregulated serum miRNAs in ESCC [8], GCA [9], GNCA [10], COAD, READ [11], PAAD [12] LUSC [13] and OV [14], and plasma miRNAs in ESCC [15], GNCA [16], PAAD [17], LUSC [18], LUAD [19], THCA [20], COAD, READ [21] and OV [22] with normalization to different reference genes. To identify circulating miRNAs in pan-cancers, we re-analyzed the data of the serum and plasma miRNAs with the consistent control genes (the spiked in cel-miR-39). Additionally, serum samples of NPC, THCA, LUAD, BRCA, UCEC, PRAD and CESC, and plasma samples of NPC, BRCA, CESC and UCEC were analyzed in accordance with the criteria. In total, serum samples from 15 types of cancers including 2050 patients and plasma samples from 13 types of cancers including 1781 cases with their own age and gender matched NCs were explored to identify circulating miRNAs in pan-cancers.
According to the workflow (GCA serum, GNCA serum, OV serum and OV plasma samples were evaluated through training and testing stages), circulating miRNAs confirmed through the training, testing and the external validation stages by cancer and sample types were determined (Table 1). As shown in Fig. 4A, dysregulated circulating miRNAs identified from serum and plasma samples in the same type of cancers were not identical. However, some miRNAs showed consistent dysregulation in both serum and plasma samples, such as let-7b-5p and miR-140-3p for NPC, miR-106a-5p and miR-93-5p for LUSC, miR-192-5p for PAAD and let-7b-5p for BRCA (“bold” in Fig. 4A). Interestingly, miR-221-3p was down-regulated in serum samples but up-regulated in plasma samples in LUAD. Meanwhile, some miRNAs were dysregulated consistently among different types of pan-cancers. For serum samples, high expression of let-7b-5p, miR-106a-5p, miR-192-5p, miR-195-5p, miR-19a-3p, miR-19b-3p, miR-20a-5p, miR-20b-5p, miR-21-5p, miR-223-3p, miR-25-3p, miR-296-5p, miR-425-5p, miR-92a-3p and miR-93-5p were found in at least two types of pan-cancers. Up-regulation of plasma let-7b-5p, miR-20a-5p, miR-20b-5p and miR-21-5p was confirmed in at least two types of pan-cancers.
Overview of signature miRNAs with potential for cancer detection. (A): Signature miRNAs presented by different human organs. S: serum; P: plasma; Marked red: oncofetal miRNAs; bold: miRNAs with consistent expression pattern in serum and plasma samples. NPC: nasopharyngeal carcinoma; THCA: thyroid cancer; ESCC: esophageal squamous cell carcinoma; GCA: gastric cardia adenocarcinoma; GNCA: gastric non-cardia adenocarcinoma; COAD: colon adenocarcinoma; READ: rectum adenocarcinoma; PAAD: pancreatic adenocarcinoma; LUSC: lung squamous cell carcinoma; LUAD: lung adenocarcinoma; BRCA: breast cancer; UCEC: uterine corpus endometrial carcinoma; PRAD: prostate adenocarcinoma; CESC: cervical squamous cell carcinoma; OV: ovarian cancer; NA: not available. (B): Circos plots of serum (left) and plasma (right) miRNAs in pan-cancers and UCB. Left circos plot: Eight tracks from outer to inner: 1. locations of serum miRNAs on chromosome; 2. serum miRNAs in UCB (red: oncofetal miRNAs); 3. serum miRNAs in ESCC (green), BRCA (blue) and LUAD (violet); 4. serum miRNAs in UCEC (green) and COAD (blue); 5. serum miRNAs in GNCA (green) and PAAD (blue); 6. serum miRNAs in LUSC (green) and NPC (blue); 7. serum miRNAs in PRAD (green) and THCA (blue); 8. serum miRNAs in READ (green) and GCA (blue). Right circos plot: Six tracks from outer to inner: 1. locations of plasma miRNAs on chromosome; 2. plasma miRNAs in UCB (red: oncofetal miRNAs); 3. plasma miRNAs in LUSC (green) and NPC (blue); 4. plasma miRNAs in LUAD (green), UCEC (blue) and BRCA (violet); 5. plasma miRNAs in PAAD (green) and CESC (blue).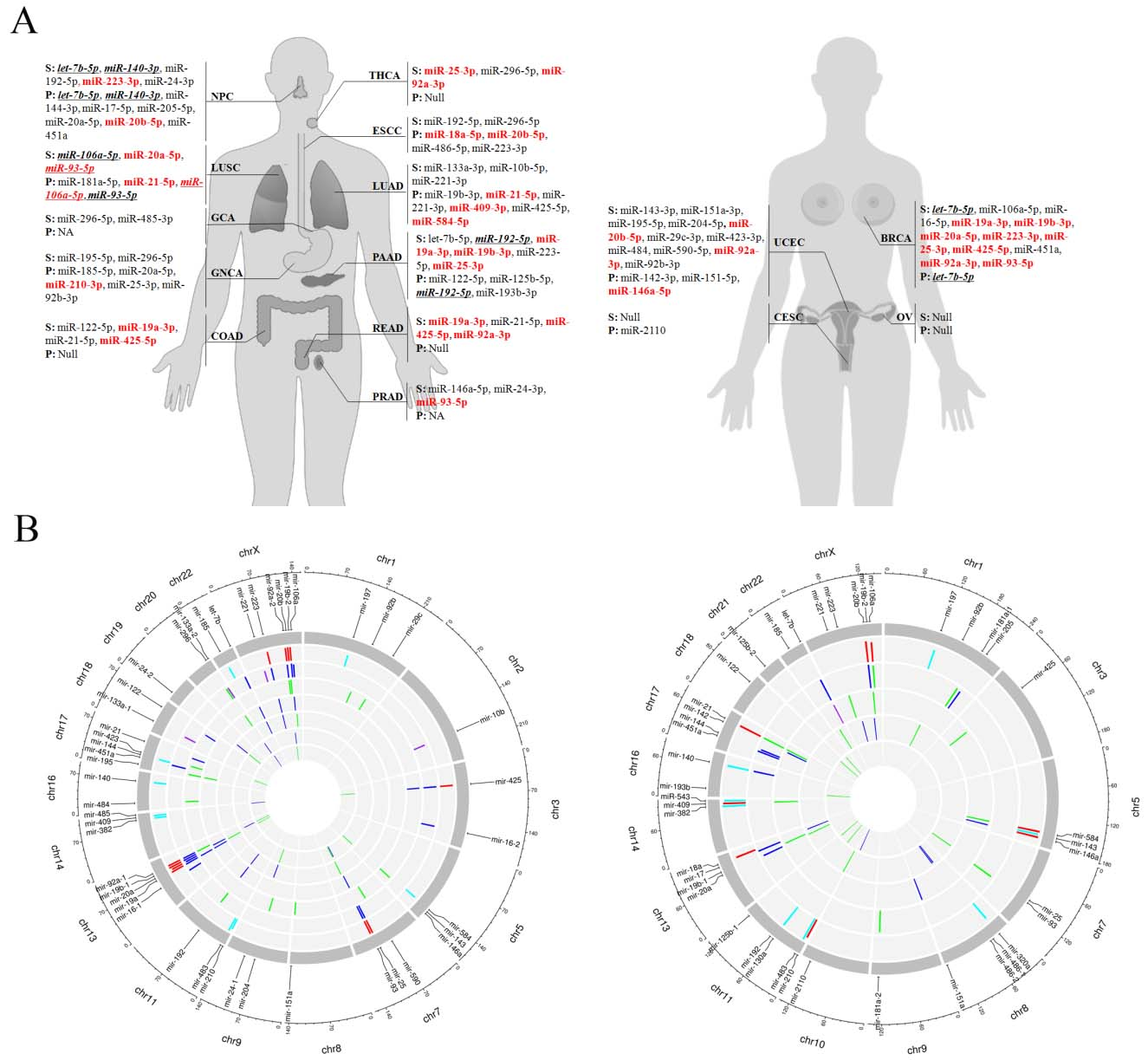
Among 34 dysregulated miRNAs in pan-cancers and 18 miRNAs in UCB from serum samples, up-regulated miR-19b-3p, miR-20a-5p, miR-223-3p, miR-93-5p, miR-25-3p, miR-425-5p, miR-19a-3p, miR-92a-3p and miR-20b-5p showed consistent trend in expression levels were identified as serum oncofetal miRNAs (marked red in Fig. 4A). Similarly, up-regulated miR-106a-5p, miR-146a-5p, miR-18a-5p, miR-20b-5p, miR-210-3p, miR-21-5p, miR-409-3p and miR-584-5p were demonstrated as plasma oncofetal miRNAs among 32 dysregulated miRNAs in pan-cancers and 16 miRNAs in UCB from plasma samples (marked red in Fig. 4A).
All the miRNAs evaluated in serum and plasma samples of pan-cancers and UCB were visualized in circos plots. As shown in Fig. 4B, dysregulated serum miRNAs including oncofetal miRNAs (red bars) mainly located in chromosome 7, 13 and X. And eight of nine serum oncofetal miRNAs (except miR-223-3p) were members of four miRNA clusters (miR-17-92 cluster: miR-19a-3p, miR-20a-5p, miR-19b-3p and miR-92a-3p; miR-191/425 cluster: miR-425-5p; miR-106a-363 cluster: miR-20b-5p, miR-19b-3p and miR-92a-3p; miR-106b-25 cluster: miR-93-5p and miR-25-3p). On the other hand, mainly dysregulated plasma miRNAs was found on chromosome 11, 13, 17 and X (Fig. 4B). And plasma oncofetal miR-409-3p belongs to miR-379-410 cluster, miR-18a-5p to miR-17-92 cluster, and miR-106a-5p and miR-20b-5p to miR-106a-363 cluster.
Diagnostic performance of circulating oncofetal miRNAs in cancer patients
To evaluate the diagnostic capacity of circulating oncofetal miRNAs in cancer patients, ROC analyses were conducted with the specific oncofetal miRNA/miRNA panel (more than one oncofetal miRNA in a type of cancer) in the specific cancers. For serum samples, oncofetal miR-223-3p for NPC, panel of miR-92a-3p and miR-25-3p for THCA, signature of miR-19a-3p and miR-425-5p for COAD, combination of miR-19a-3p, miR-425-5p and miR-92a-3p for READ, panel of miR-19a-3p, miR-19b-3p and miR-25-3p for PAAD, signature of miR-20a-5p and miR-93-5p for LUSC, combination of miR-19b-3p, miR-20a-5p, miR-223-3p, miR-93-5p, miR-25-3p, miR-425-5p, miR-19a-3p and miR-92a-3p for BRCA, and panel of miR-92a-3p and miR-20b-5p for UCEC were explored. And the AUCs for the eight specific types of cancers ranged from 0.668–0.987 (Fig. 5A–H). For plasma samples, AUCs of oncofetal miR-20b-5p for NPC, panel of miR-18a-5p and miR-20b-5p for ESCC, miR-210-3p for GNCA, signature of miR-106a-5p and miR-21-5p for LUSC, combination of miR-21-5p, miR-409-3p and miR-584-5p for LUAD, and miR-146a-5p for UCEC ranged from 0.606-0.76 (Fig. 5I–N).
Receiver-operating characteristic (ROC) curve analyses of the specific serum oncofetal miRNAs (A-H) and plasma oncofetal miRNAs (I-N) to discriminate patients with specific cancers from normal controls. A: miR-223-3p for NPC; B: combination of miR-92a-3p and miR-25-3p for THCA; C: combination of miR-19a-3p and miR-425-5p for COAD; D: combination of miR-19a-3p, miR-425-5p and miR-92a-3p for READ; E: combination of miR-19a-3p, miR-19b-3p and miR-25-3p for PAAD; F: combination of miR-20a-5p and miR-93-5p for LUSC; G: combination of miR-19b-3p, miR-20a-5p, miR-223-3p, miR-93-5p, miR-25-3p, miR-425-5p, miR-19a-3p and miR-92a-3p for BRCA; H: combination of miR-92a-3p and miR-20b-5p for UCEC. I: miR-20b-5p for NPC; J: combination of miR-18a-5p and miR-20b-5p for ESCC; K: miR-210-3p for GNCA; L: combination of miR-106a-5p and miR-21-5p for LUSC; M: combination of miR-21-5p, miR-409-3p and miR-584-5p for LUAD; N: miR-146a-5p for UCEC.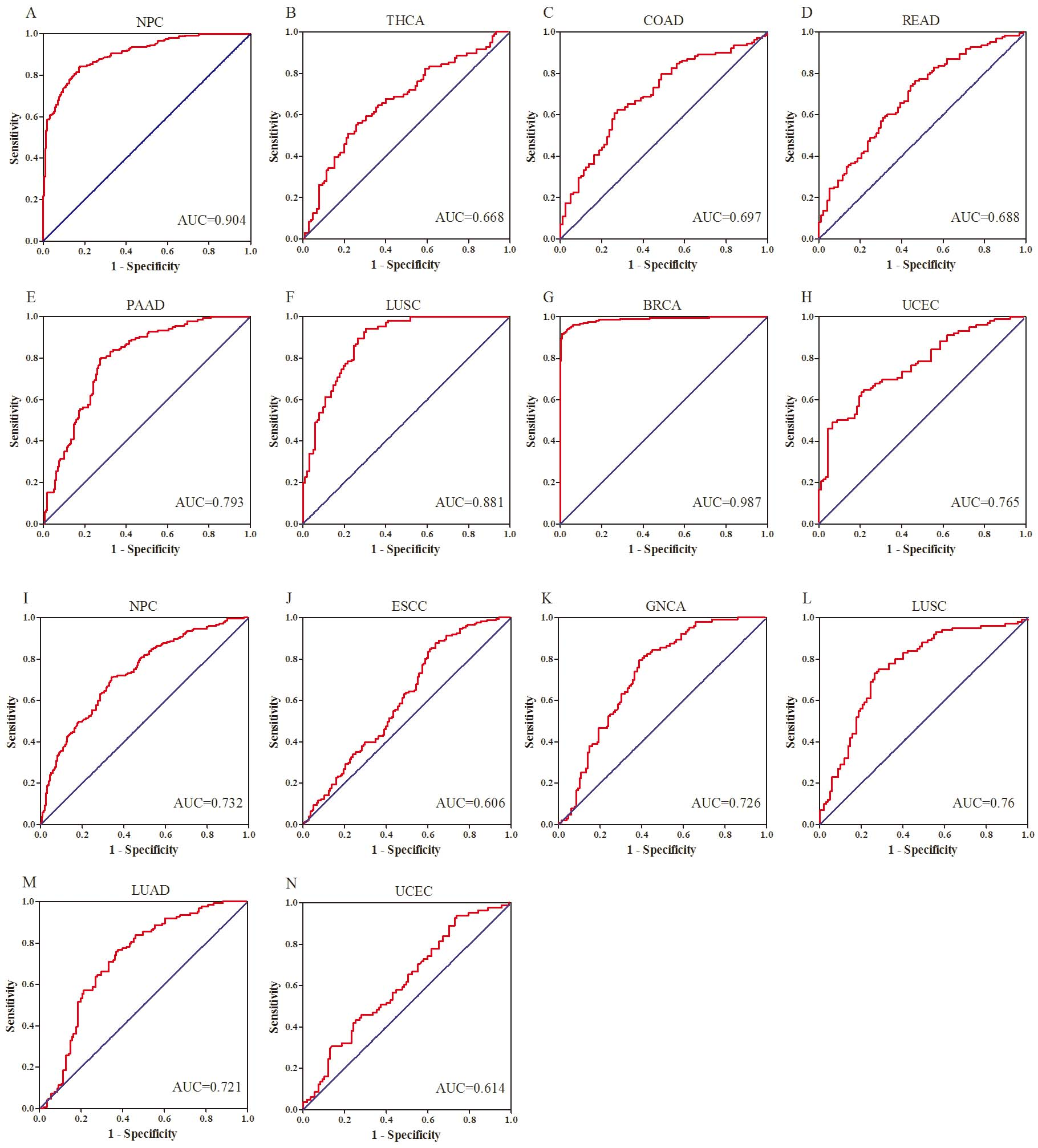
Expression levels of the identified serum and plasma oncofetal miRNAs in 13 types of tumor tissues comparing with normal tissues using data from TCGA. Presented as log(fold change). HNSC: head neck squamous cell carcinoma; THCA: thyroid cancer; ESCA: esophageal cancer; STAD: stomach adenocarcinoma; COAD: colon adenocarcinoma; READ: rectum adenocarcinoma; PAAD: pancreatic adenocarcinoma; LUSC: lung squamous cell carcinoma; LUAD: lung adenocarcinoma; BRCA: breast cancer; CESC: cervical squamous cell carcinoma; UCEC: uterine corpus endometrial carcinoma; PRAD: prostate adenocarcinoma.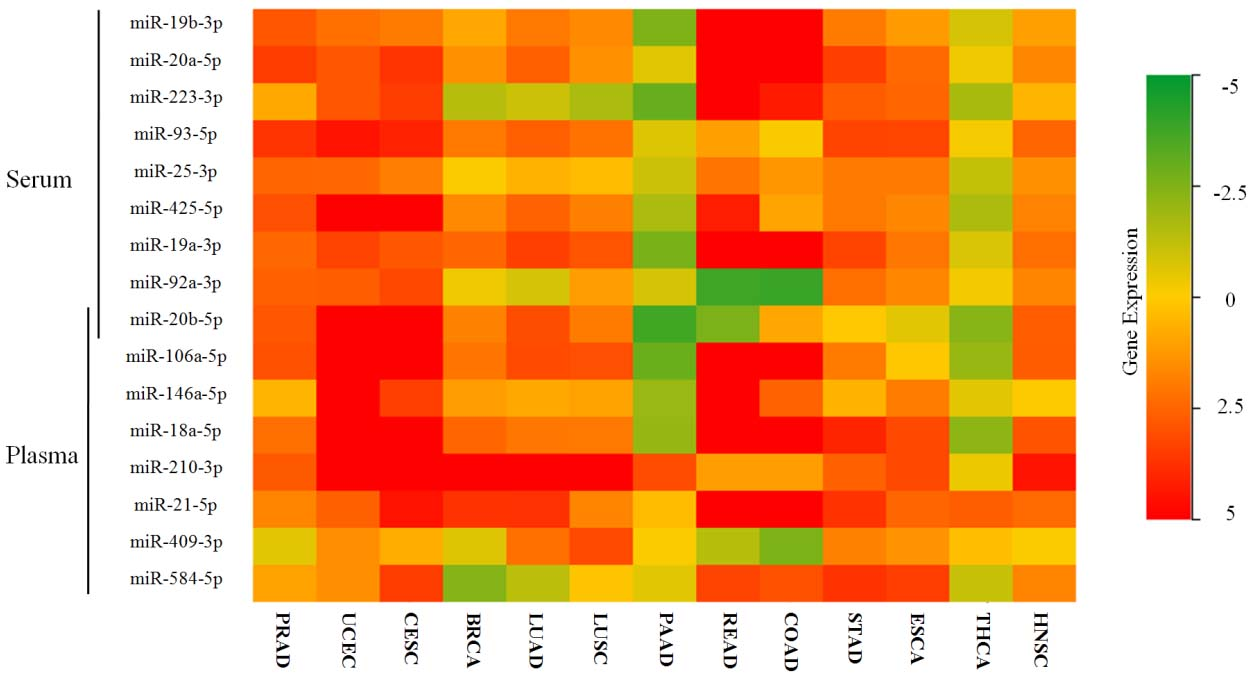
To evaluate the expression levels of the oncofetal miRNAs in tumor tissues, miRNA isoform data of 13 types of cancers and matched normal controls were retrieved from the TCGA database. As shown in Fig. 6, except in PAAD and THCA, the majority of the oncofetal miRNAs showed higher expression in tissues of pan-cancers than those in normal tissues.
Enrichment analysis of the oncofetal miRNAs
To identify genes potentially targeted by the oncofetal miRNAs, mirDIP was assessed. A total of 73 genes were predicted to be common target genes of at least 8 of the 9 serum oncofetal miRNAs. Among them, CPEB3, LCOR, PAPD5, PGM2L1, PTEN and SMG1 could be targeted by all the nine serum oncofetal miRNAs. Meanwhile, among the 28 common target genes (at least 7 of the 8 plasma oncofetal miRNAs), FBXO28, KLF12 and NFAT5 could be targeted by any one plasma oncofetal miRNA (Fig. S1A).
In addition, the web server MISIM v2.0 was used to explore functional enrichment and similarity analysis of the identified oncofetal miRNAs. The associated function, diseases and transcription factors of miRNAs were predicted (Tables S7 and S8; The top 10 terms of function were shown in Fig. S1B). Regardless serum or plasma oncofetal miRNAs could act as onco-miRNAs, play important roles in cancer associated processes such as cell proliferation, apoptosis, and epithelial-to-mesenchymal transition, and could be regulated by well known oncogene MYC, Stat5 and so on. Similarity analyzes showed that miRNAs, especially the serum oncofetal miRNAs, exhibited well functional similarity (Fig. S1C).
Discussion
Most of previous studies have made efforts to identified circulating miRNAs with potential in detecting various types of cancers so as to early diagnose the disease and prolong the survival of patients. However, the investigators always focused on single or few types of cancers with different methods in diverse populations and yielded inconsistent findings among studies. Cancer is a category of diseases shared great similarity (e.g. hallmarks of cancer) and heterogeneity. Pan-cancer analyzes could promote us better understand the complex relationship and interactions of the diseases. Nowadays, a global view of circulating miRNAs in pan-cancers with consistent methods in the same population to show the similarity and heterogeneity of circulating miRNAs in the diseases is lacking. In addition, the similarity and interaction between embryo development and tumorigenesis also encouraged us to explore the cancer related circulating miRNAs from the view of embryo. Thus, we performed the comprehensive analysis of circulating miRNAs in UCB and pan-cancers to identify the potential broad-spectrum or/and specific biomarkers for cancer detection.
In the current study, according to different cancer categories as well as sample types, independent exploration of circulating miRNAs was performed in consistent multiple-stages and analytic methods with 2050 serum samples from 15 types of cancers and 1781
Expression levels of the signature miRNAs in pan-cancers (presented as mean
NPC: nasopharyngeal carcinoma; THCA: thyroid cancer; ESCC: esophageal squamous cell carcinoma; GCA: gastric cardia adenocarcinoma; GNCA: gastric non-cardia adenocarcinoma; COAD: colon adenocarcinoma; READ: rectum adenocarcinoma; PAAD: pancreatic adenocarcinoma; LUSC: lung squamous cell carcinoma; LUAD: lung adenocarcinoma; BRCA: breast cancer; UCEC: uterine corpus endometrial carcinoma; PRAD: prostate adenocarcinoma; CESC: cervical squamous cell carcinoma; OV: ovarian cancer.
plasma samples from 13 types of cancers. In total, 34 serum miRNAs and 32 plasma miRNAs were dysregulated in at least one type of cancer and showed potential in aiding of cancer detection. Analyzes of UCB also demonstrated that 18 serum miRNAs and 16 plasma miRNAs were closely related with embryos. A qualified biomarker should be sensitive and specific to the disease with reliable concentrations to be detected easily. Reportedly, more than 2500 mature human miRNAs have been identified. Concerning sample type, only a small part of miRNome showed relatively high abundance could be stably detected in circulating samples. Thus, we applied Exiqon miRNA qPCR panels to screen potential miRNAs from 168 miRNAs which were evaluated to be stable in serum/plasma with relatively high abundance by the producer (Vedbaek, Denmark) in the initial screening phase of each independent study.
Serum and plasma are two of the most commonly used sample types in clinical diagnosis and investigations. Due to different isolation methods, serum and plasma contain different components including miRNA expression profiles [7, 25]. No study has demonstrated the superiority of application of serum or plasma sample in cancer detection with circulating miRNAs. In our study, except GCA and PRAD, we analyzed the other 13 types of cancers with serum and plasma samples (not matched sample from the same individual) independently. Overall, 15 miRNAs including let-7b-5p, miR-106a-5p, miR-122-5p, miR-140-3p, miR-146a-5p, miR-192-5p, miR-19b-3p, miR-20a-5p, miR-20b-5p, miR-21-5p, miR-25-3p, miR-425-5p, miR-451a, miR-92b-3p and miR-93-5p (miR-221-3p and miR-223-3p showed different expression pattern) were consistently dysregulated in circulation of cancer patients among the 34 serum and 32 plasma miRNAs identified from pan-cancer analyzes (the venn diagram was shown in Fig. S2). For the same sample type, 15 serum miRNAs (let-7b-5p, miR-106a-5p, miR-192-5p, miR-195-5p, miR-19a-3p, miR-19b-3p, miR-20a-5p, miR-20b-5p, miR-21-5p, miR-223-3p, miR-25-3p, miR-296-5p, miR-425-5p, miR-92a-3p and miR-93-5p) and 4 plasma miRNAs (let-7b-5p, miR-20a-5p, miR-20b-5p and miR-21-5p) were respectively found in at least two types of cancers. Among the 13 types of cancers with both serum and plasma samples, NPC (let-7b-5p and miR-140-3p), LUSC (miR-106a-5p and miR-93-5p), PAAD (miR-192-5p) and BRCA (let-7b-5p) possessed consistently dysregulated circulating miRNAs in both types of samples. These miRNAs might show relative commonality among different types of samples and cancers, and might act as broad-spectrum biomarkers to some extent. The other circulating miRNAs might be specific to the type of cancer or/and sample and could be identified as specific biomarkers. The appropriate combination of the broad-spectrum and specific biomarkers could improve the sensitivity and specifity of cancer detection. However, it was noted that except miR-106a-5p in LUSC, let-7b-5p and miR-140-3p in NPC, miR-93-5p in LUSC, miR-192-5p in PAAD and let-7b-5p in BRCA all showed higher fold changes in serum samples than those in plasma samples. Meanwhile, compared to NCs, the mean fold change of 34 serum miRNAs (2.96
Compared to adult NCs, 18 and 16 miRNAs were identified to be up-regulated in serum and plasma samples of UCB, respectively. Among them, miR-140-5p, miR-197-3p, miR-20b-5p, miR-210-3p, miR-382-5p, miR-409-3p, miR-483-5p and miR-584-5p were consistently up-regulated in both sample types of UCB. Similarity of embryos and tumors prompted us to identify the miRNAs which showed consistent expression pattern between fetus and cancer samples as oncofetal miRNAs. Thus, a total of 16 oncofetal circulating miRNAs including 9 serum oncofetal miRNAs (miR-19b-3p, miR-20a-5p, miR-223-3p, miR-93-5p, miR-25-3p, miR-425-5p, miR-19a-3p, miR-92a-3p and miR-20b-5p) and 8 plasma oncofetal miRNAs (miR-106a-5p, miR-146a-5p, miR-18a-5p, miR-20b-5p, miR-210-3p, miR-21-5p, miR-409-3p and miR-584-5p) were confirmed in the study. It seemed that big divergence existed between the expression profiles of oncofetal miRNAs from serum and plasma samples. Only oncofetal miR-20b-5p was overlapped in the expression profiles from the two types of samples. The diagnostic capacity of oncofetal miRNAs was retrospectively evaluated in specific cancers. The results showed that the specific circulating oncofetal miRNA(s) could effectively distinguish specific types of cancer patients from NCs. However, with larger AUCs, serum oncofetal miRNAs might demonstrate better diagnostic performance than plasma oncofetal miRNAs for cancer patients (e.g. AUCs for NPC: serum: 0.904, plasma: 0.732; LUSC: serum: 0.881, plasma: 0.76; UCEC: serum: 0.765, plasma: 0.614). These results showed difference in application of serum and plasma oncofetal miRNAs in diagnosis of cancer patients.
Among the nine serum oncofetal miRNAs, eight miRNAs were identified as members of four miRNA clusters (miR-17-92, miR-191/425, miR-106a-363 and miR-106b-25 cluster), while 4 of 8 plasma oncofetal miRNAs were confirmed from miR-379-410 cluster, miR-17-92 cluster and miR-106a-363 cluster. As the three paralogous clusters, miR-17-92, miR-106a-363 and miR-106b-25 were highly conserved across species, played indispensable roles in processes of embryonic development and were demonstrated to be prooncogenic in a wide range of malignancies [26, 27, 28]. Application of circulating miRNAs from the three clusters as diagnostic biomarkers was also assessed in several types of cancers [29, 30]. Originated from miR-191/425 cluster, miR-425-5p could act as an oncogene in colorectal cancer and gastric cancer, and might serve as non-invasive indicators for detection of colorectal cancer, gastric cancer and so on [31, 32]. As a member of miR-379-410 cluster and a miRNA expressed by embryonic stem cells, miR-409-3p seemed to play controversial roles in tumorigenesis. Oncogenic roles of miR-409-3p were reported in most types of cancers except in prostate cancer [33, 34, 35]. But high expression of circulating miR-409-3p was common in cancer patients such as individuals with breast cancer or colorectal cancer [36, 37]. These findings showed inconsistency between gene ontology and expression levels of circulating miRNAs which should be paid attention to. The other one serum oncofetal miRNAs, miR-223-3p, also showed discrepancy of gene function in various cancers and even displayed the opposite expression pattern in ESCC plasma samples in our study [38, 39, 40]. Among the other 4 plasma oncofetal miRNAs (miR-146a-5p, miR-210-3p, miR-21-5p and miR-584-5p), miR-21-5p was a well known onco-miRNA and was studied thoroughly in various cancers including its capacity as a cancer biomarker [41]. And miR-21-5p was also essential for embryo implantation and development [42]. MiR-210-3p, a typical hypoxia-related miRNA, might function as an oncogene or tumor suppressor during tumorigenesis. But most evidence showed that circulating miR-210 was up-regulated in many solid tumors including breast cancer, pancreatic cancer, lung cancer and esophageal cancer [43]. Similarly, contradictory results of mechanistic function and expression level of miR-146a-5p in cancers were also described by previous studies [44]. Few studies have explored the roles of miR-584-5p in cancer, thus need to be further investigated. Though controversial roles of some miRNAs in cancer were inconclusive, the oncofetal miRNAs confirmed in our study could broad the view of application of circulating miRNAs in cancer detection.
In addition, we also performed bioinformatic analyzes to evaluate the roles of the serum and plasma oncofetal miRNAs might play. Both the serum and plasma oncofetal miRNAs identified in the study could target many targets. Target predication showed that common targets (at least targeted by n-1 miRNAs) of serum oncofetal miRNAs were much more than those of plasma oncofetal miRNAs. Among the six genes which were targeted by all the nine serum oncofetal miRNAs, PTEN, SMG1 and CPEB3 were confirmed as tumor suppressors in various cancers [45, 46, 47]. As for the three genes commonly targeted by eight plasma oncofetal miRNAs, only KLF12 could act as a tumor suppressor in lung cancer [48]. Majority of serum oncofetal miRNAs from the three paralogous clusters, which could regulate a similar set of genes and have overlapping functions, might partly explain the difference of common target genes between the serum and plasma oncofetal miRNAs. Similarity analysis presented similar findings that functional similarity was found among oncofetal miRNAs, yet more closely among the serum oncofetal miRNAs than those in plasma. However, most of serum and plasma oncofetal miRNAs colud be classified as onco-miRNAs in perspective of functional mechanism. These miRNAs were also closely related with immune system, involved in processes of angiogenesis and cell death, regulated vital pathways (e.g. Akt and NF-kB pathways) both in tumorigenesis and embryo development. Predicted associated diseases for the oncofetal miRNAs also covered most of solid tumors. It might show the universality of the oncofetal miRNAs for pan-cancers. Some well known prooncogenic transcription factors, such as MYC, Stat5 and E2F1, were predicted to be upstream regulators of the oncofetal miRNAs. These results showed the close relationships of oncofetal miRNAs and cancer in mechanism, and could promote us better use the potential diagnostic biomarkers.
Currently, it is not completely clear about the precise mechanism of the release of miRNAs into the extracellular environment. Circulating miRNAs were considered as passive leakage or active secretion with protection of protein complexes or membrane-bound vesicles (e.g. exosomes) by various cells. Once absorbed by recipient cells, circulating miRNAs could elicit corresponding regulatory functions and act as messengers among cells [49]. As cancer is a complex systemic disease, dysregulated circulating miRNAs in cancer were not only derived from tumor cells but also from other cells involved in tumorigenesis and development of cancer. Thus, circulating miRNAs could not only reflect the status of tumor itself but also systemic responses (such as local immune response and systemic inflammation) of cancer patients. Understanding relationship between circulating miRNAs and tumor tissue specific miRNAs is important for us to delineate their roles in cancer pathobiology and cancer prevention. In our previous studies, we have analyzed the expression levels of some identified serum/plasma miRNAs in tumor tissues and demonstrated that some miRNAs were not consistently dysregulated in circulation and tissues. To investigate the expression levels of oncofetal miRNAs in pan-cancers, we retrieved and analyzed miRNA isoform data of 13 types of cancers from the TCGA database. The results showed that except in PAAD and THCA, most of oncofetal miRNAs displayed consistent expression pattern between circulation and tissue in pan-cancers. The findings indicated that the oncofetal miRNAs in circulation might be closely related to tumor. Future efforts should be paid to focus on the mechanism of the circulating miRNAs in cancers.
Over the past decade, few studies have reported oncofetal miRNAs in some types of cancers. Yamamoto et al. identified miR-500 as an oncofetal miRNA in hepatocellular carcinoma, due to its consistently high expression in tumor tissues and fetal liver compared to adult normal liver [50]. Becker-Santos et al. determined 37 oncofetal miRNAs which shared similar expression patterns in fetal lung and NSCLC tumors, versus adult non-malignant lung tissues. And the predicted target genes of oncofetal miRNAs were involved in lung development and carcinogensis [51]. However, the two studies only assessed oncofetal miRNAs from tissue samples in single type of cancer. In the present study, we performed pan-cancer analyzes of circulating miRNAs and identified a set of oncofetal miRNAs in serum/plasma samples. The findings might serve as bases in the field of oncofetal miRNAs and contribute to the potential application of the potential non-invasive biomarkers in the future clinical.
Some issues should be addressed and settled before the former establishment of circulating miRNAs as non-invasive biomarkers for cancer detection in clinical. Firstly, it is critical to choose appropriate sample type for examining circulating miRNAs. Wang et al. performed a meta-analysis to assess the diagnostic performance of circulating miRNAs in detection of gastrointestinal cancer and found that plasma-based miRNA assay reached a higher accuracy than serum-based one in gastric cancer, while opposite conclusion was drawn for colorectal cancer [52]. In our study, regardless of the specific type of cancer, we speculated that serum samples might be superior to plasma samples for miRNAs detection when comprehensively take account in expression levels, AUCs in diagnosis of cancer patients and predicated functional mechanism. Secondly, current methodologies for circulating miRNA quantification were varied among studies. Normalization of miRNA concentrations, in order to minimize the impact of biological and technical variabilities, is important for the reliable quantification of circulating miRNAs in serum and plasma samples. The input sample volume, endogenous controls (e.g. miR-16 and miR-1228) and spike-in controls (e.g. ce-miR-39) were the most frequently used controls for normalization. In our previous studies, we have applied endogenous controls (according to other investigators or evaluated by geNorm software), spike-in controls or combination of the two controls as references for identification of circulating miRNAs. However, it is not comparable for the expression profiles using different normalization genes. It was recommended that the simultaneous processing of all samples collected from identical input volumes with the same spiked-in miRNA concentrations for correcting the variability among circulating samples [6]. Thus, we analyzed the expression profiles of circulating miRNAs with consistent volume and the same spiked-in miRNA cel-miR-39 in the study and compared the circulating miRNAs between pan-cancers and UCB. Meanwhile, each study with different types of cancer/UCB and sample types was performed independently which further make our findings more credible. Thirdly, the specificity and sensitivity of circulating miRNAs for diagnosing patients with different types of cancers and different populations are worth to be further explored. In the study, we identified some miRNAs (especially oncofetal miRNAs) might be broad-spectrum, while the other miRNAs specific for some specific types of cancers in Chinese populations. The appropriate combination of these miRNAs might further improve the diagnostic performance of cancer.
Several limitations of the study should be considered. Firstly, most of serum samples and plasma samples of the same type of cancer/UCB were not from the same individuals. Secondly, the diagnostic capacity of the identified circulating miRNAs especially oncofetal miRNAs was not additionally evaluated in all cancer patients to identify the appropriate combination of the miRNAs and cutoff values for discriminating patients with different types of cancers as well as NCs. Thirdly, the existing form of the identified miRNAs in circulation should be further evaluated. Finally, multicenter studies including more patients with different population backgrounds were needed to be further conducted.
In conclusion, we performed a comprehensive analysis of circulating miRNAs in pan-cancers and UCB, and identified circulating miRNAs (especially oncofetal miRNAs) which could act as potential broad-spectrum or/and specific biomarkers for cancer detection. More efforts should be made before the application of circulating miRNAs for the detection of various cancers in the clinical.
Author contributions
Conception: P.L., W.F.C., and W.Z.
Interpretation or analysis of data: X.Z., C.L., Y.Y.
Preparation of the manuscript: C.Z., X.Z., T.S.X., and X.N.G.
Revision for important intellectual content: X.Z., P.L., W.F.C, and W.Z.
Supervision: W.Z.
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-203085.
sj-doc-1-cbm-10.3233_CBM-203085.doc - Supplemental material
Supplemental material, sj-doc-1-cbm-10.3233_CBM-203085.doc
Footnotes
Acknowledgments
This work was supported by the National Natural Science Foundation of China [Grant number: 81672400; 81672788; 81702364] and the Natural Science Foundation of Jiangsu Province [Grant number: BK20171085].
