Abstract
Background:
Carbonic anhydrase 9 (CA9) plays a crucial role in pH regulation and adaptation under hypoxic conditions in the tumor microenvironment. Despite its known involvement in the progression of specific cancers, a comprehensive pan-cancer examination of the prognostic value and biological implications of CA9 has not been performed. This study systematically explored the diverse roles of CA9 across multiple cancer types.
Methods:
Bioinformatics methods were applied via extensive datasets from TCGA, GTEx, CPTAC, CancerSEA, and the public literature. We systematically analyzed the associations between CA9 expression profiles and various clinical parameters, prognosis, immune infiltration, immune-related genes, TMB, MSI, and tumor stemness scores. Additionally, a single-cell functional analysis was conducted.
Results:
CA9 was significantly upregulated in 29 out of 33 cancer types, indicating high discriminatory ability between tumor and normal tissues. Elevated CA9 expression correlated with poor OS and PFIs in multiple cancers, such as GBMLGG, CESC, LUAD, KIPAN, GBM, THYM, LIHC, THCA, PAAD, and KICH. In 39 cancers, CA9 expression was predominantly negatively correlated with the infiltration of 22 immune cell infiltrations. It was also associated with TMB in 12 tumors and with MSI in 9. Single-cell analysis revealed positive links between CA9 and essential processes such as hypoxia, metastasis, angiogenesis, and stemness.
Conclusion:
This study provides compelling evidence that CA9 is a potential pan-cancer prognostic marker and diagnostic tool. The associations of CA9 with immune components and determinants of immunotherapy response indicate the importance of CA9 in advancing cancer research and personalized treatment strategies.
Introduction
Tumor occurrence, progression, metastasis, and recurrence are closely related to the tumor microenvironment (TME). The TME, a complex milieu encompassing various cellular and non-cellular components, plays a pivotal role in influencing tumor progression and therapeutic responses. One critical aspect of the tumor microenvironment is its acidity, which often arises from metabolic alterations. As a driver of cancer, this acidic microenvironment not only promotes tumor cell survival and invasiveness but also poses significant obstacles to conventional therapeutic interventions. 1 Understanding the intricate interplay between tumors and the acidic microenvironment is paramount for developing targeted and effective therapeutic strategies. In this context, exploring the molecular and cellular mechanisms underlying acidification of the tumor microenvironment holds promise for advancing our knowledge and improving clinical outcomes in cancer management.
Carbonic anhydrases (CAs) are a large family of zinc metalloenzymes that catalyze the reversible hydration of carbon dioxide. They catalyze CO2/HCO3− interconversions in different macroscopic and microscopic environments of the body and are thus involved in physiological and pathological processes such as respiration and CO2 excretion, pH homeostasis, electrolyte secretion, gluconeogenesis, adipogenesis and ureagenesis, bone resorption and calcification, and oncogenicity. As a membrane-bound isozyme, CA9 is strongly upregulated in many hypoxic tumors, transferring H+ from the cytoplasm to the external environment via reversible HCO3- dehydration/CO2 hydration, and protecting against pH regulation within the tumor cell, which in turn affects tumor cell survival, invasion and metastasis.2,3 CA9 expression is upregulated and often associated with poor prognosis in many malignancies, including breast,4 -6 colon, 7 head and neck,8,9 kidney,10 -12 lung,13,14 ovary, 15 and pancreatic16,17 cancers. High CA9 expression is mediated by hypoxia-inducible factor (HIF)-1a under hypoxia, which is a common feature of solid tumors. 18 The ability of CA9 to neutralize the intracellular pH by facilitating bicarbonate transport across the plasma membrane provides cancer cells with a survival advantage when exposed to hypoxic and acidic microenvironments. 19
In addition to directly regulating pH in hypoxic and acidic environments, CA9 also participates in cancer growth by providing nutrients and interacting with amino acid transfer proteins. The transport of glutamine into cells is fundamental and is mediated by several amino acid transporters (AATs), such as alanine serine cysteine transporter 2 (ASCT 2), sodium coupled neutral amino acid transporter 2 (SNAT 2), and L-type amino acid transporter 1 (LAT 1), all of which may be affected by the pH gradient produced by CA9. 20 In addition, the activity of CA9 in tumor cells is closely related to key aspects of cancer transmission such as adhesion, migration, and invasion. CA9 can interact with β-catenin, disrupt the calcium binding protein actin junction, weaken cell to cell adhesion, and promote cell migration. 21 In adition, CA9 can regulate the stemness of breast cancer cells through the mTORC1 axis. This CA9 mediated activity is necessary for supporting the transcriptional activity of EMT and “stemness drivers,” such as Notch1 and Jagged1. Accordingly, the pharmacological inhibition of CA9 leads to the depletion of cancer stem cells, delayed tumor growth and the inhibition of lung metastasis in breast cancer.22,23 Gradually, methods targeting CA9 have been developed to inhibit its activity in tumor tissues, and thus far, there has been considerable acceleration in treatment selection in this field. Small molecule inhibitors,24,25 mixed/dual targeted drugs,26 -28 targeted antibodies,29 -31 and adoptive cell therapies (CAR-T cells)32 -34 have been developed at the preclinical level, whereas sulfonamide CA9 inhibitors and antibodies have entered phase Ib/II clinical trials for the treatment and imaging of different solid tumors.
However, the majority of prior investigations have focused primarily on elucidating the role of CA9 within a single specific cancer. Comprehensive pan-cancer analyses examining the prognostic relevance and biological implications of CA9 are scarce in the literature. In our research, we conducted a systematic exploration of the correlations between CA9 expression and immune infiltration levels, DNA methylation patterns, tumor mutational burden (TMB), and microsatellite instability (MSI) across diverse cancer types. Additionally, we investigated the associations between CA9 and immune-related genes, as well as their influence on the TME. Our findings indicate that CA9 can serve as a prognostic indicator across various tumors, exerting a pivotal role in tumor immunity by influencing the composition of tumor-infiltrating immune cells, MSI, and TMB. This research contributes valuable insights into the functional role of CA9 in the context of tumor immunotherapy.
Methods
Data Acquisition and Processing
We retrieved comprehensive datasets for our study from the UCSC (https://xenabrowser.net/) database, specifically obtaining gene expression, clinical information, and associated mutation data spanning various cancers within The Cancer Genome Atlas (TCGA) pan-cancer initiative. The TCGA tumor abbreviations and the full names are provided in Table S1. Concurrently, CA9 expression data in normal tissues were obtained from the Genotype Tissue Expression (GTEx) project through the UCSC platform. To ensure uniformity in our analyses, we applied a log2(TPM + 0.001) transformation to each expression value. Furthermore, for a more nuanced understanding of CA9 protein expression in diverse normal tissues and tumors, we accessed protein expression data from the National Institutes of Health (NIH) Proteomic Data Commons (CPTAC) database (https://pdc.cancer.gov/pdc/). The reporting of this study conforms to the STREGA (Strengthening the Reporting of Genetic Association Studies: An Extension of the STROBE Statement) statement of equator network reporting guidelines. 35
Prognostic Analyses
We employed the “coxph” function from the R package “survival” (version 3.5) 34 to construct Cox proportional hazards regression models for analyzing the associations between gene expression and survival indicators, including overall survival (OS), progression-free interval (PFI), disease-specific survival (DSS), and disease-free interval (DFI), for each tumor. As a complement, we obtained a high-quality prognostic dataset for TCGA from a previous TCGA prognostic study published in Cell. 36 Finally, we excluded patients whose follow-up time was shorter than 30 days. Prognostic significance was determined through statistical tests, specifically the log-rank test.
Immunological Infiltration Assessment
For the assessment of immunological infiltration, we utilized the “deconvo_CIBERSOR” method from the R package “IOBR” (version 0.99), 37 which employs a robust approach to enumerate cell subsets from tissue expression profiles. The composition of immune cell infiltration in each tumor was subsequently reassessed on the basis of gene expression.
Additionally, we employed the R package “ConsensusTME” (version 0.0.1.900) 38 to calculate gene expression-based stromal, immune, and ESTIMATE scores for each tumor. In addition, 150 marker genes for 5 types of immune pathways including the chemokine (41), receptor (18), MHC (21), immunoinhibitor (24), and immunostimulatory (46) pathways, were identified from a previous study conducted by Hu et al. 39 This comprehensive approach provides a detailed understanding of the tumor microenvironment by evaluating both immune cell composition and overall stromal and immune components, enhancing the depth of our analysis.
Gene Mutation Analysis of CA9
cBioPortal (https://www.cbioportal.org/) 40 provides a web-based platform for analyzing CA9 gene tumor genomic profiles. We used this database to analyze the pan-cancer mutation frequency of CA9, whose mutational alterations include mutation, amplification, and deep deletion.
Single-Cell Function Analysis
CancerSEA (http://biocc.hrbmu.edu.cn/CancerSEA/) 41 provides 14 functional statuses at the single-cell level, including angiogenesis, apoptosis, cell cycle, differentiation, DNA damage, DNA repair, EMT, hypoxia, inflammation, invasion, metastasis, proliferation, quiescence, and stemness. Thus, we used the CancerSEA database to identify CA9-related functional states.
Immunofluorescence (IF) and Immunohistochemistry (IHC)
To ascertain the cellular localization of CA9 and discern disparities in protein expression levels, IF and IHC images were acquired from the Human Protein Atlas (HPA) database (http://www.proteinatlas.org/). The downloaded images were meticulously analyzed to gain insights into the subcellular distribution of CA9 and variations in its protein expression across various tissues.
Additionally, 16 immunohistochemistry sections of 5 different tumor tissues (cervical cancer, endometrial cancer, renal clear cell carcinoma, hepatocellular carcinoma, and colorectal cancer) were obtained from the Department of Pathology at the Second Affiliated Hospital of Chongqing Medical University. In-house immunohistochemical staining for CA9 was systematically performed on these sections. CA9 polyclonal antibody (cat no: 1107-1-AP) was purchased from proreintech.co (Wuhan China). This experiment was reviewed and approved by the Ethics Committee of the Second Affiliated Hospital of Chongqing Medical University (No. 2025KL-34). Image-pro plus 6.0 (Media Cybernetics, Inc., Rockville, MD, USA) was used to measure the mean density (Integrated Optiondensity (IOD) /area) of CA9.
Statistical Analysis
The Wilcoxon rank sum test and Kruskal-Wallis test were used to compare gene expression differences. Spearman or Pearson correlation analysis was used to calculate the correlation between 2 groups. Kaplan-Meier and Cox regression analysis were used to compare the survival characteristics of the 2 groups of patients. The above statistical analysis and visualization were performed via R software (version 4.3.1) and GraphPad Prism 8.
Results
Expression of CA9 Between Tumor and Normal Tissues
In TCGA pan-cancer database, we detected significant CA9 expression upregulation in 16 tumor tissues compared with normal tissues—GBM, CESC, LUAD, COAD, COADREAD, BRCA, KIRP, KIPAN, UCEC, HNSC, KIRC, LUSC, READ, PAAD, BLCA, and CHOL—and was accompanied by downregulation in 4 tumor tissues—PRAD, LIHC, THCA, and KICH (Figure 1A). Owing to the small number of normal tissues in TCGA, we included GTEx and TCGA for co-analysis. In total, we observed significant upregulation in 26 out of 34 tumors (Figure 1B). Moreover, we compared CA9 protein abundance in different CTPAC tumors. Compared with that in normal tissue, increased CA9 expression was observed in breast cancer, colon cancer, head and neck cancer, kidney cancer, lung squamous cell cancer, pancreatic ductal adenocarcinoma, and uterine cancer but decreased only in stomach cancer (Figure 1C-L).

Expression of CA9 in pan-cancer from different databases: (A) CA9 expression in TCGA database, (B) CA9 expression in TCGA + GTEx database, and (C-L) CA9 protein abundance in CTPAC.
The Prognostic Value of CA9
We constructed Cox proportional hazard models to investigate the associations between CA9 expression levels and patient prognosis, including OS, PFI, DSS, and DFI, across diverse tumor types. Elevated CA9 expression was associated with unfavorable OS in GBMLGG, CESC, LUAD, KIPAN, GBM, THYM, LIHC, THCA, PAAD, and KICH but was associated with favorable outcomes in DLBC. Moreover, high CA9 expression was associated with a poor PFI in GBMLGG, BRCA, CESC, KIRP, KIPAN, THYM, LIHC, and CHOL (Figure 2A and B).

The univariate regression for OS: (A), PFI (B), DSS (C), and DFI (D) in pan-cancer.
For DSS, GBMLGG, BRCA, CESC, LUAD, KIPAN, GBM, THYM, LIHC, THCA, and PAAD patients had poor prognoses (Figure 2C). However, regarding the DFI, no specific correlation with gene expression was detected (Figure 2D). In summary, elevated CA9 expression has emerged as a variable but generally adverse prognostic factor across multiple tumor types, highlighting its potential significance in predicting patient outcomes.
CA9 is Differentially Expressed in Different Tumor Stages
We assessed the association between CA9 expression and tumor stage and observed significant differences in 11 tumors such as COAD, COADREAD, BRCA, KIRP, KIPAN, THYM, LIHC, PAAD, BLCA, ACC, and CHOL. The observed trend indicated a notable increase in CA9 expression levels with advancing tumor stages, as depicted in Figure 3A. For example, in BRCA, KIPAN and LIHC, as the tumor stage increases from stage I to stage II, III, or even stage IV, the expression of CA9 also increases significantly.
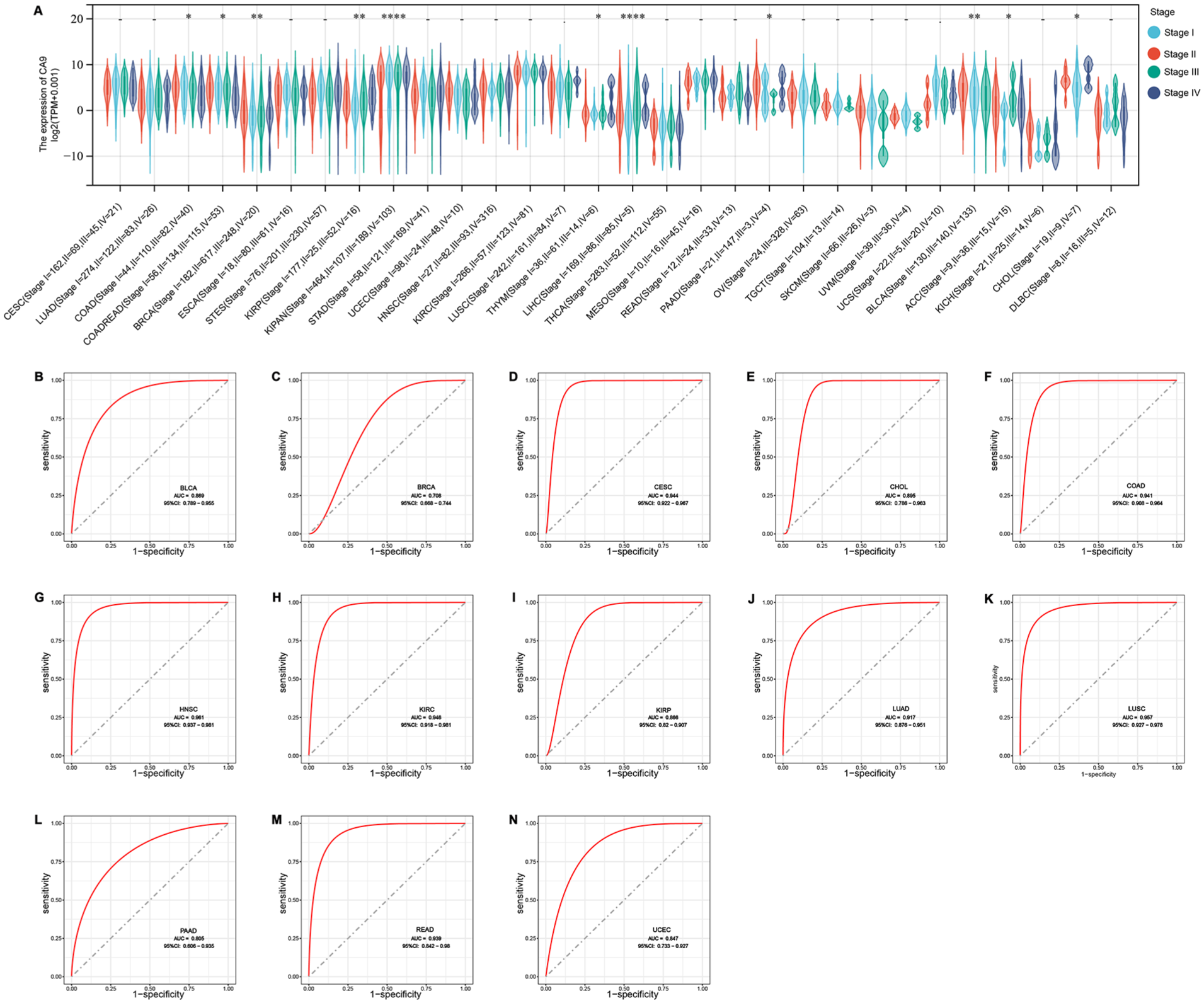
CA9 differentially expressed in tumor stages and can distinguish tumors from normal tissue: (A) the relationship between the expression of CA9 and tumor stages and (B-N) ROC analysis of CA9 expression between tumor and paired normal samples.
CA9 Showed Excellent Ability to Distinguish Tumors from Normal Tissue
We conducted a receiver operating characteristic (ROC) analysis to assess the discriminatory capacity of CA9 between tumor and paired normal samples across various cancers. The results revealed that CA9 exhibited an excellent ability to differentiate between tumor and normal tissues. The area under the curve (AUC) values were notably high, predominantly within the range of 0.8 to 0.9 for several cancers, including BLCA, CESC, CHOL, COAD, HNSC, KIRC, KIRP, LUAD, LUSC, PAAD, READ, and UCEC. Notably, a slightly lower AUC of 0.708 was observed for BRCA (Figure 3B-N). This finding suggested that CA9 expression levels hold promising diagnostic potential, particularly in specific cancers where CA9 expression demonstrates robust discriminatory performance between tumor and normal tissues.
Association Between CA9 Expression and Tumor-infiltrating Lymphocytes
Utilizing the CIBERSORT algorithm, we obtained immune cell infiltration scores spanning 22 categories in a total of 39 tumors. We subsequently calculated Pearson’s correlation coefficients to assess the relationship between CA9 expression and immune cell infiltration scores. Notably, we observed significant correlations between CA9 expression and immune infiltration in 37 out of the 39 cancer types investigated. Across most cancers, CA9 demonstrated a predominantly negative correlation with the infiltration levels of the 22 immune cell categories, as illustrated in Figure 4A.

Correlation of CA9 expression and tumor microenvironment. (A) Relationship between CA9 expression and tumor-infiltrating lymphocytes. The shadow depth of the lower triangle represents the size of the P-value for statistical analysis, while the shadow depth of the upper triangle represents the size of the correlation coefficient (*P < .05). (B-J) Correlation between CA9 expression and ESTIMATE score in GBMLGG, KIPAN, and KIRC.
Moreover, upon calculating stromal, immune, and ESTIMATE scores for each patient using the R package ESTIMATE, we discovered that CA9 expression was significantly positively correlated with stromal, immune, and overall ESTIMATE scores in GBMLGG, KIPAN, and KIRC, as depicted in Figure 4B to J. These findings underscore the intricate association between CA9 expression and the TME, emphasizing its potential impact on both immune cell infiltration and overall stromal and immune components in specific cancer types.
CA9 Expression and Immune Gene Correlations
Furthermore, gene correlation analyses were conducted to explore the associations between CA9 expression and immune pathway marker genes, including chemokines, receptors, major histocompatibility complex (MHC) genes, immunoinhibitors, and immunostimulators. As depicted in Figure 5, a significant correlation was observed between CA9 and nearly all of these immune-related genes. Notably, in CHOL, MESO, SARC, STAD, STES, THCA, HNSC, and CESC, the majority of immune-related genes were negatively correlated with CA9. In contrast, significant positive correlations were identified in KIPAN, KIRC, GBMLGG, LIHC, PRAD, LUAD, OV, BRCA, and COAD. These findings underscore the intricate and context-dependent relationships between CA9 expression and immune-related pathways across cancers.
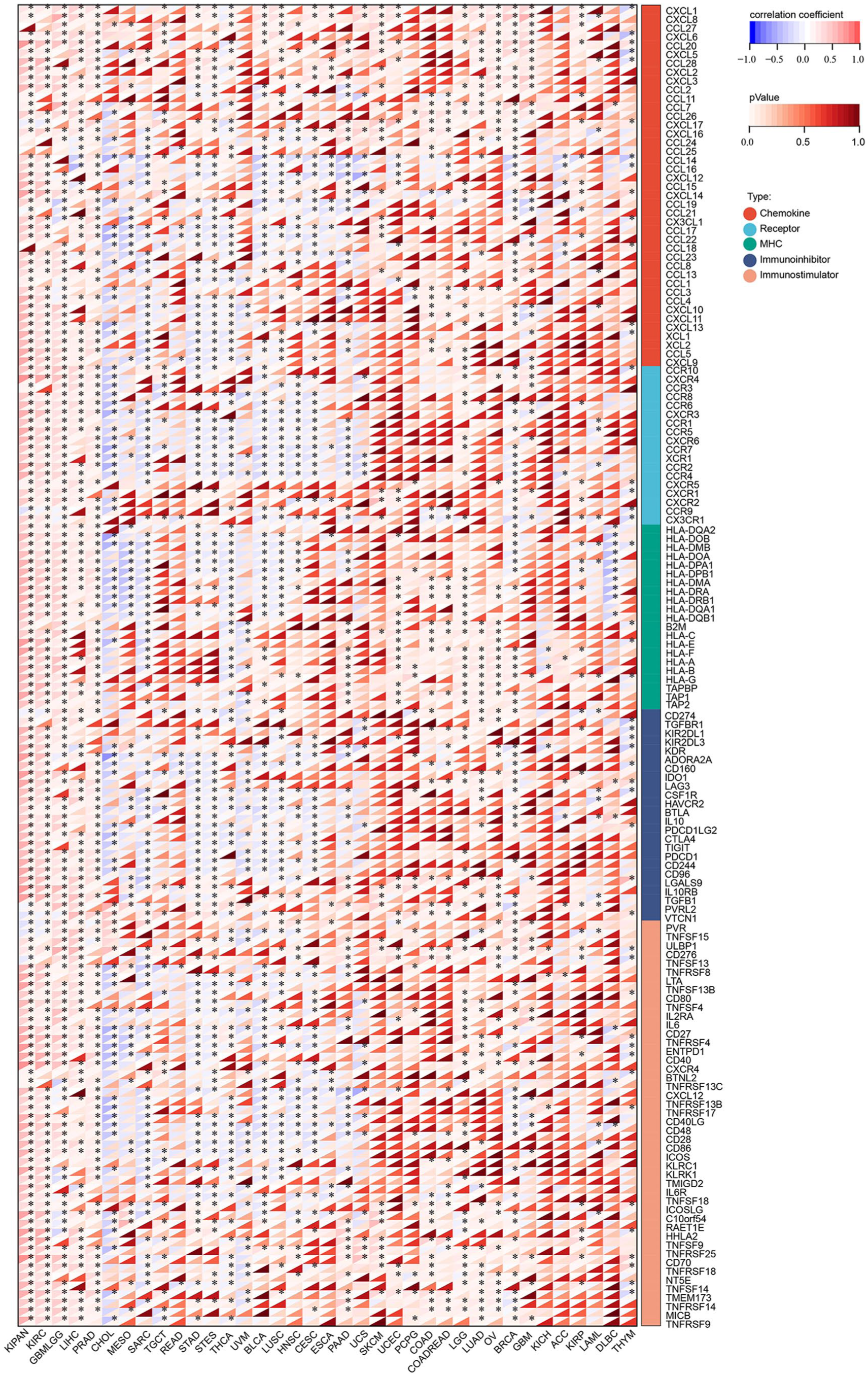
CA9 expression and immune pathways marker genes correlation. The shadow depth of the lower triangle represents the size of the P-value for statistical analysis, while the shadow depth of the upper triangle represents the size of the correlation coefficient.
CA9 and Immunotherapy
TMB, MSI, and DNA methylation are pivotal factors influencing immunotherapy outcomes, including responses to immune checkpoint therapy. Consequently, we investigated the associations between CA9 expression and these crucial factors. First, we explored CA9 genetic alterations across cancers via cBioPortal. The frequency of genetic variation in CA9 was highest in PAAD (6.52%) and was predominantly amplified. The second and third highest frequencies of CA9 were found in HNSC (5.35%) and UCEC (4.61%), respectively, and were predominantly amplified and mutated (Figure 6C).

CA9 mutation landscape (C) and its correlation with TMB (A), MSI (B), DNAss (D).
We subsequently examined the correlation between CA9 expression and TMB, MSI, and DNA methylation. CA9 expression was significantly positively correlated with TMB in 12 tumor types, including GBMLGG (R = .11, P = .0041), LGG (R = .10, P = .0318), COAADREAD (R = .14, P = .0066), KIRP (R = .1928, P = .0012), KIPAN (R = .13, P = .0009), THYM (R = .3268, P = .0003), PCPG (R = .16, P = .0310), UCS (R = .30, P = .0222), KICH (R = .374124197166968, P = .0020), and DLBC (R = .34), P = .0369). LUAD (R = −.14, P = .0015), and UVM (R = −.24, P = .0318) were negatively correlated with TMB (Figure 6A). A significant correlation with MSI was detected in 9 tumors, with a positive correlation in 7 tumors, such as COAD, COADREAD, STES, SARC, STAD, KICH, and THYM, and a negative correlation in GBMLGG and KIPAN (Figure 6B). Furthermore, when DNA-based tumor stemness scores were calculated via methylation profiling from a previous study, 42 significant correlations were observed in 15 tumors. A positive correlation was noted in 11 tumors, including GBMLGG, LGG, BRCA, STES, SARC, STAD, PRAD, HNSC, LUSC, BLCA, and KICH, and a negative correlation was noted in CESC, LIHC, and THCA (Figure 6D).
Single-cell Gene Function Analysis
To elucidate the intricate mechanisms governed by CA9, we performed gene function analyses at the single-cell level via CancerSEA. The results revealed a positive correlation between CA9 expression and key cellular processes, including hypoxia, metastasis, angiogenesis, and stemness (Figure 7A). Specifically, CA9 was positively correlated with hypoxia in BRCA, GBM, NSCLC, and RCC.

Single cell gene function analysis of CA9 across Pan-cancer (A) and in individual cancer types (B-H).
In contrast, a negative correlation was observed between CA9 and processes associated with DNA repair, apoptosis, DNA damage, epithelial-mesenchymal transition (EMT), and metastasis in UM. Additionally, negative correlations with DNA repair and inflammation were noted in BRCA, whereas negative correlations with DNA repair, the cell cycle, and DNA damage were observed in RB.
Cellular Localization and Tissue Expression of CA9
Understanding its cellular localization is crucial for comprehending its functional importance, particularly in the context of cancer biology and the development of targeted therapies. Through IF staining of A431 cells, we determined the subcellular localization of CA9. As a membrane protein, CA9 strongly fluoresces on the surface of the cell membrane (Figure 8A). This distinct membrane localization aligns with its role in mediating pH regulation and adaptation to hypoxic conditions within the cellular microenvironment. According to the immunohistochemistry results, CA9 was more highly expressed in kidney, liver, urinary bladder and colon cancer tissues than in normal tissues (Figure 8B-F). In our validation cohort, CA9 was highly expressed on the cell membrane in pathological samples of CRC, LIHC, KIRC, CESC, and UCEC, with particularly strong expression (mean density) in KIRC and UCEC (Figure 8G). This heightened expression profile in various cancer types underscores the potential significance of CA9 as a biomarker and therapeutic target.

Immunofluorescence (IF) of CA9 in A431 cell (A) and immunohistochemistry (IHC) in HPA tumor/normal tissue (B-F) and validation cohort (G and H).
Discussion
Owing to lifestyle changes and advances in screening and treatments, overall cancer mortality has been declining over the years. 43 However, there are still many tumors with increasing incidence rates and little progress in treatment. The search for effective diagnostic and prognostic markers and targets is still a relentless pursuit for clinicians. According to our study, CA9 may be a candidate gene.
As a transmembrane isoenzyme that mediates the regulation of acidity within and outside the cell membrane, the normal expression of CA9 is limited to the gastric epithelium, bile ducts, choledochal ducts, pancreatic ducts, rapidly proliferating normal cells of the small intestine, and, to a lesser extent, the central nervous system. 44 Consistent with various studies7,16,45 showing that CA9 is highly expressed in certain tumors, our study revealed that CA9 expression was widely upregulated in various cancers, highlighting its potential as a robust diagnostic marker across a wide spectrum of cancers. The prognostic value of CA9 is evident, as elevated CA9 expression is correlated with unfavorable outcomes in several cancers, including GBMLGG, CESC, and LUAD. However, a favorable prognosis was noted in DLBC, emphasizing the complexity of the role of CA9 in different malignancies. The differential expression in various tumor stages further underlines the potential utility of CA9 as an indicator of disease progression, with increased expression associated with advanced stages in multiple cancers.
The tumor microenvironment consists of complex tumor cells, tumor-infiltrating immune cells, the extracellular matrix, and the vasculature and is generally hypoxic and acidic. 46 Tumor-infiltrating immune cells play a role in either tumor suppression or tumor promotion, 47 whereas hypoxia and an acidic environment are considered 2 important factors in suppressing the immune response to tumors. 48 CA9 is strongly upregulated in many hypoxic tumors and, in turn, acts through reversible HCO3− dehydration/CO2 hydration to transfer protons from the cytoplasm to the external environment,2,49 which further exacerbates acidic alterations in the tumor microenvironment. The correlation between CA9 expression and immune cell infiltration, which was negatively correlated with CD8+ T, Treg, macrophage, and NK cell infiltration, highlights its involvement in shaping the TME. The predominantly negative correlation with immune cell infiltration suggested a potential immunosuppressive role of CA9.
Owing tumor heterogeneity, the correlation between CA9 and immune-related genes varies from tumor to tumor. High CA9 expression corresponds to low expression of chemokines, receptors, MHC, immunoinhibitors, and immunostimulators in CHOL, MESO, SARC, STAD, STES, THCA, HNSC, and CESC, but the reverse is true in other tumors. The use of immunotherapy has increased owing to the discovery of immune checkpoints (programed death (PD)-1, PD ligand 1 (PD-L1), and cytotoxic T lymphocyte antigen (CTLA)-4) and the use of immune checkpoint inhibitors.50,51 However, there is also a need for a comprehensive assessment of the population for which ICIs are indicated and the therapeutic efficacy of the treatment because of possible life-threatening inflammatory toxicity. 52 The relevance of CA9 to these immune genes suggests that it may serve as a novel immunotherapeutic target or as a reference for the response to ICIs in various cancers.
TMB and MSI are also reliable biomarkers of prognosis in various cancers and predictors of the efficacy of immunotherapy in many cancers, such as breast cancer, colorectal cancer, and head and neck cancer.53 -55 Our study revealed that CA9 was positively correlated with TMB in 10 tumors and was positively correlated with MSI in 7 tumors, which is consistent with previous research, further supporting the role of CA9 as a potential biomarker linked to genomic instability and responsiveness to immunotherapy. The concordance between CA9 expression and TMB/MSI reinforces the potential utility of CA9 as a comprehensive prognostic and predictive marker in the context of cancer management and immunotherapeutic interventions.
Single-cell functional analysis revealed that CA9 is significantly associated with hypoxia and angiogenesis in BRCA, CRC, GBM, RB, and RCC. HIF-1a is highly expressed under hypoxic conditions, which in turn promotes the upregulation of CA9 expression, 18 thus there is no doubt that a significant correlation between the 2 is observed at the pancancer level. Studies have shown that combining CA9 inhibitors and antiangiogenic agents in TNBC significantly inhibits tumor growth and distant metastasis, 56 whereas antiangiogenic inhibitors alone increase the incidence of distant metastasis. 57 A major result of that research was that blocking CA9 not only reduced the number of vessels in the primary tumor but also reduced the permeability of the remaining vessels. These results highlight the potential therapeutic implications of targeting CA9, particularly in the context of modulating angiogenesis and addressing the challenges associated with antiangiogenic treatment strategies.
In summary, this study revealed the expression patterns of CA9 in various cancers and its association with prognosis. While initial bias assessment was performed using our laboratory’s constrained dataset, the statistical power necessitates expanded validation through independent cohorts and orthogonal experimental approaches. We found that certain variants of CA9 are significantly associated with poorer survival and a higher risk of disease recurrence in certain types of cancer. This discovery not only emphasizes the importance of CA9 as a potential biomarker, but also provides a theoretical basis for personalized treatment strategies based on CA9. Inhibitors targeting CA9 (SLC-0111) are currently in early clinical trials and have shown good safety and certain anti-tumor activity, paving the way for further exploration of their potential as therapeutic targets. 58 In the future, with the deepening development of precision medicine concepts and advances in technological means, CA9 is expected to become an important component of personalized medicine. By integrating multiple omics data, we can construct more accurate disease models and achieve personalized prediction and intervention for each patient. Especially for patients with complex genetic backgrounds or refractory cancers, customized treatment based on CA9 may lead to unprecedented treatment opportunities. However, further fundamental research is needed to elucidate the mechanism of action of CA9, as well as large-scale clinical validation to ensure its applicability and effectiveness in different populations.
Conclusion
In conclusion, this comprehensive pan-cancer study of CA9 highlights its diverse roles and potential clinical significance. The differential expression of CA9 across multiple cancers underscores its utility as a potent marker for both diagnostic and prognostic purposes. In addition, the correlations between CA9 expression and the TME, TMB, MSI, DNAss, etc., show that CA9 is a valuable tool for evaluating the efficacy of immunotherapies. The role of CA9 in tumorigenesis and development needs to be further investigated. Considering its broad impact across different cancer types, CA9 has emerged as an excellent candidate for future individualized targeted therapies.
Supplemental Material
sj-doc-1-cix-10.1177_11769351251380520 – Supplemental material for Systematic Analysis of CA9 as a Pan-Cancer Marker for Prognosis and Immunity
Supplemental material, sj-doc-1-cix-10.1177_11769351251380520 for Systematic Analysis of CA9 as a Pan-Cancer Marker for Prognosis and Immunity by Qiang Yi, Yaoyao Mei, Zhu Yang and Yi Liu in Cancer Informatics
Supplemental Material
sj-doc-2-cix-10.1177_11769351251380520 – Supplemental material for Systematic Analysis of CA9 as a Pan-Cancer Marker for Prognosis and Immunity
Supplemental material, sj-doc-2-cix-10.1177_11769351251380520 for Systematic Analysis of CA9 as a Pan-Cancer Marker for Prognosis and Immunity by Qiang Yi, Yaoyao Mei, Zhu Yang and Yi Liu in Cancer Informatics
Footnotes
Acknowledgements
We extend our sincere gratitude to SangerBox for their instrumental support in generating graphical illustrations during this research.
Ethical Considerations
This experiment was reviewed and approved by the Ethics Committee of the Second Affiliated Hospital of Chongqing Medical University (No. 2025KL-34).
Consent to Participate
All participants gave voluntary, informed consent for the research use and publication of their IHC slices.
Consent for Publication
All participants gave voluntary, informed consent for publication of their IHC slices.
Author Contributions
LY and YZ conceived the study. MYY and YQ downloaded the public data and performed the data analysis. YQ drafted the manuscript. All authors participated in the revision and approved the publication of the manuscript.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was funded by the “Chongqing Talents Program” of the Chongqing Municipal People’s Government (grant No. cstc2022ycjh-bgzxm0062), Scientific and Technological Research Program of Chongqing Municipal Education Commission (grant NO: KJQN202200442, YL), and Chongqing Municipal Education Commission (grant No. HZ2021006).
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
All datasets for this study are available from public databases or can be obtained by contacting the corresponding author (including the IHC information from the Second Affiliated Hospital of Chongqing Medical University).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
