Abstract
BACKGROUND:
Leukocyte infiltration plays an critical role in outcome of various diseases including Lung adenocarcinoma (LUAD).
OBJECTIVES:
To understand the genetic and epigenetic factors affecting leukocyte infiltration and identification and validation of immune based biomarkers.
METHOD:
Correlation analysis was done to get the associations of the factors. CIBERSORT analysis was done for immune cell infiltration. Genetic and epigenetic analysis were performed. Cox regression was carried out for survival.
RESULTS:
We categorized the TCGA-LUAD patients based on Leukocyte fraction (LF) and performed extensive immunogenomic analysis. Interestingly, we showed that LF has a negative correlation with copy number variation (CNV) but not with mutational load. However, several individual genetic mutations, including KRAS and KEAP1, were significantly linked with LF. Also, as expected, patients with high LF showed significantly increased expression of genes involved in leukocyte migration and activation. DNA methylation changes also showed a strong association with LF and regulated a significant proportion of genes associated with LF. We also developed and validated an independent prognostic immune signature using the top six prognostic genes associated with LF.
CONCLUSION:
Together, we have identified clinical, genetic, and epigenetic variations associated with LUAD LF and developed an immune gene-based signature for disease prognostication.
Abbreviations
Introduction
Lung adenocarcinoma (LUAD) is a significant cause of cancer related fatalities globally. Even with improvements in therapy protocols, the 5-year survival rate remains abysmal [1, 2]. Late diagnosis of patients (
Material and methods
Patients sample LF and CIBERSORT analysis
The clinical, expression, DNA methylation, CNV, and mutation data of LUAD patients were downloaded from TCGA (
Expression analysis
To identify the genes differentially expressed between tumor and normal, a
Mutation analysis
MAF file for LUAD samples were obtained from the GDC data portal (
Methylation analysis
The standard method of preprocessing Illumina 450K array DNA methylation data was used [14, 15, 16]. Additionally, probes were filtered based on standard deviation (sd) and those with sd
Copy number analysis
Masked segment mean for all the LUAD samples was obtained from the GDC data portal. To recognize the regions that are significantly amplified or deleted across the samples in low and high LF cohorts, GISTIC 2.0 algorithm (
Survival analysis
Cox regression analysis was performed to identify the genes associated with survival prediction. The genes with at least
Other statistical analysis
For comparison of two groups, the two-sided t-test was performed in GraphPad8.3, and tests with
Results
Characterization of immune nature of LUAD with high and low LF
Infiltration of leukocytes in the tumor environment plays a key role in the development and progression of cancers, including LUAD [12, 19]. While LUAD’s LF is the highest of all cancers, LUAD is also a rather heterogeneous tumor [12]. To understand the immune nature of LUAD, patients were divided into low (LF
LF of LUAD patients: A. LF is plotted against TCGA-LUAD patient numbers. The patients with LF 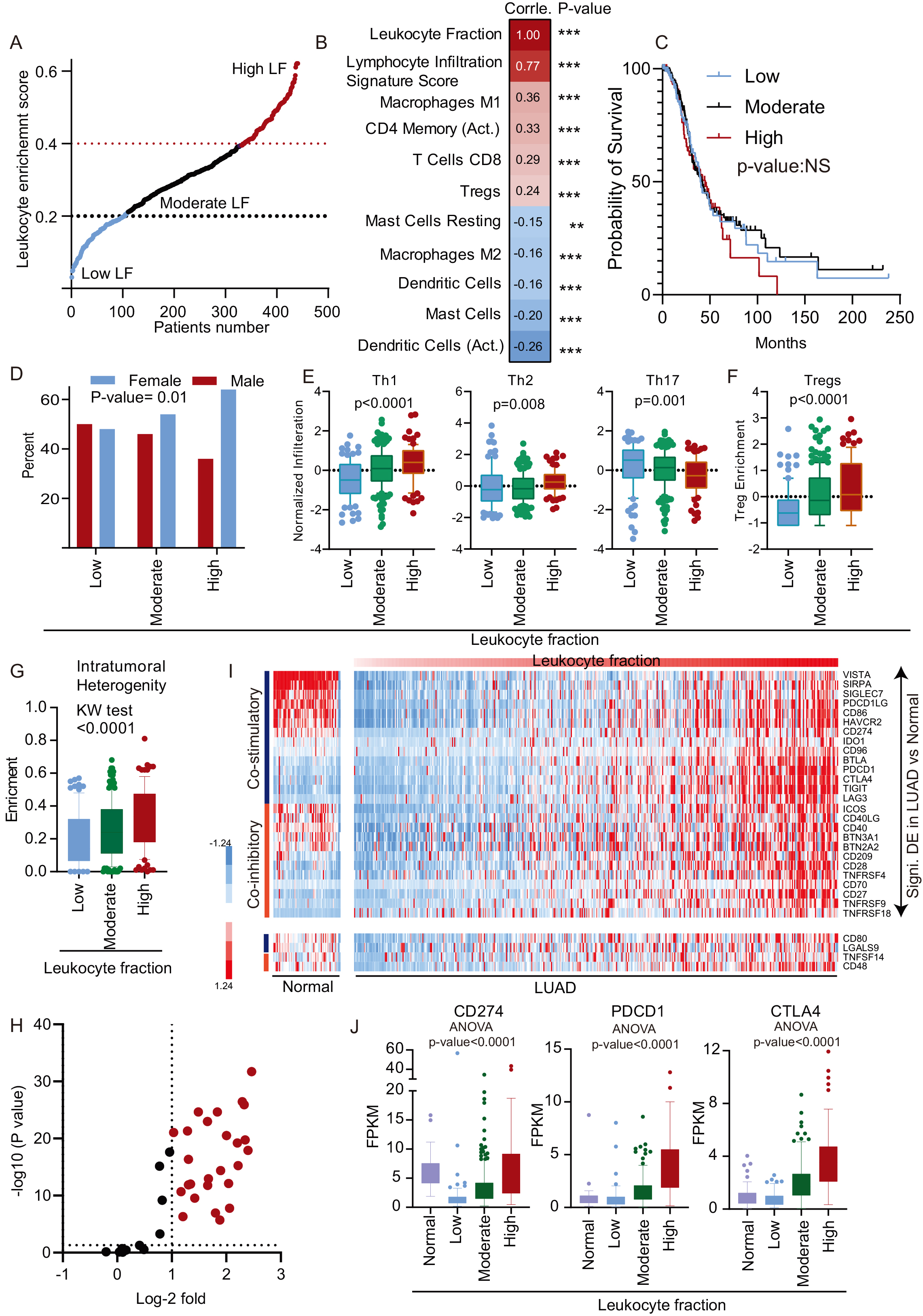
Identification of genetic aberration with LF in LUAD: A. The correlation matrix showing correlation coefficient of mentioned genomic elements. B. The total mutation burden and CNV fraction of LUAD patients was plotted in low, moderate, and high LF patients and shown in the box plot. Kruskal-Wallis test was performed to obtain the 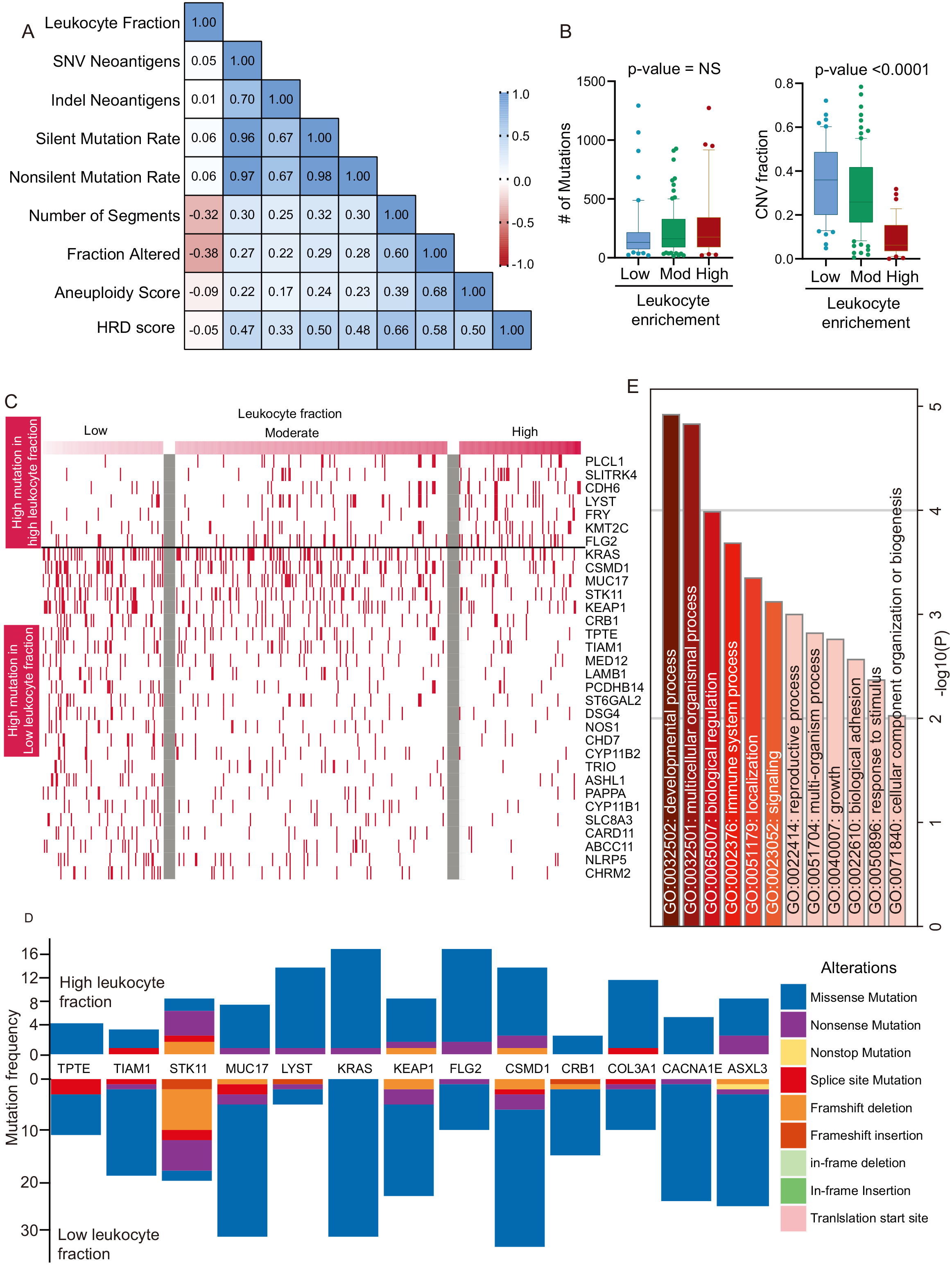
Next, we looked at the effect of different clinicopathological features on the LF of LUAD patients. Interestingly, we found that the gender of the patient has a significant association with the LF. A higher proportion of female patients were associated with a higher leukocytes fraction (Fig. 1D, Supplementary Table 3). However, other clinical parameters, such as age and stage of patients, did not show significant association with LF(Supplementary Table 3). CD4
More recently, immune checkpoint genes have become a crucial determinant of the therapy outcome due to advances in immunotherapy protocols [20]. The association of stimulatory and inhibitory checkpoint molecules with LF of patients was calculated, and the expression of these genes were compared in high vs. low LF using the
Taken together, the LUAD samples are heterogeneous in terms of the LF, and the enrichment of cells varies by cell type. Gender and intertumoral heterogeneity also play an essential role in the LF of LUAD patients.
The genetic makeup of cancer is not only a determining factor in the treatment option and the response to therapy, but it also affects the immune contexture [21]. We performed a correlation analysis to understand the relationship between genetic aberrations and the LF. There was a significant positive correlation between the LF and the number of segments and the modified fraction, suggesting that copy number variation is associated with LUAD immune cell infiltration (Fig. 2A). However, there was no correlation with LF in other genetic aberrations. To validate the finding, we compared the mean mutation and CNV fraction in LUAD patients with low, moderate, and high LFs. The analysis showed that there was no significant difference in the overall mutation burden, although the CNV fraction was significantly lower in the high LF (Fig. 2B).
Genetic aberrations were associated with the presence or absence of immune cells. For example, VHL and STK11 mutations are associated with lower macrophages infiltration [22]. In the same way, PI3KCA, MET mutations are associated with CTL infiltration [22]. We have therefore identified recurrent gene mutations (mutation frequency
Significant number of CNV in low and high LFs have been identified. At chromosome arm level, the low LF group showed significantly higher copy number variations in 1p, 1q, 6p, 7p, 8q and 17q and lower copy number of 3p, 3q, 9p, 19p and 19q (Fig. 3A). Many focal level differences between high LF and low LF of CNVs were also identified (Fig. 3B and C). The common genes associated with cancer with significant focal copy number aberration are given in Fig. 3B and C.
Copy number changes associated with LF: A. Arm level copy number analysis was performed and plotted. 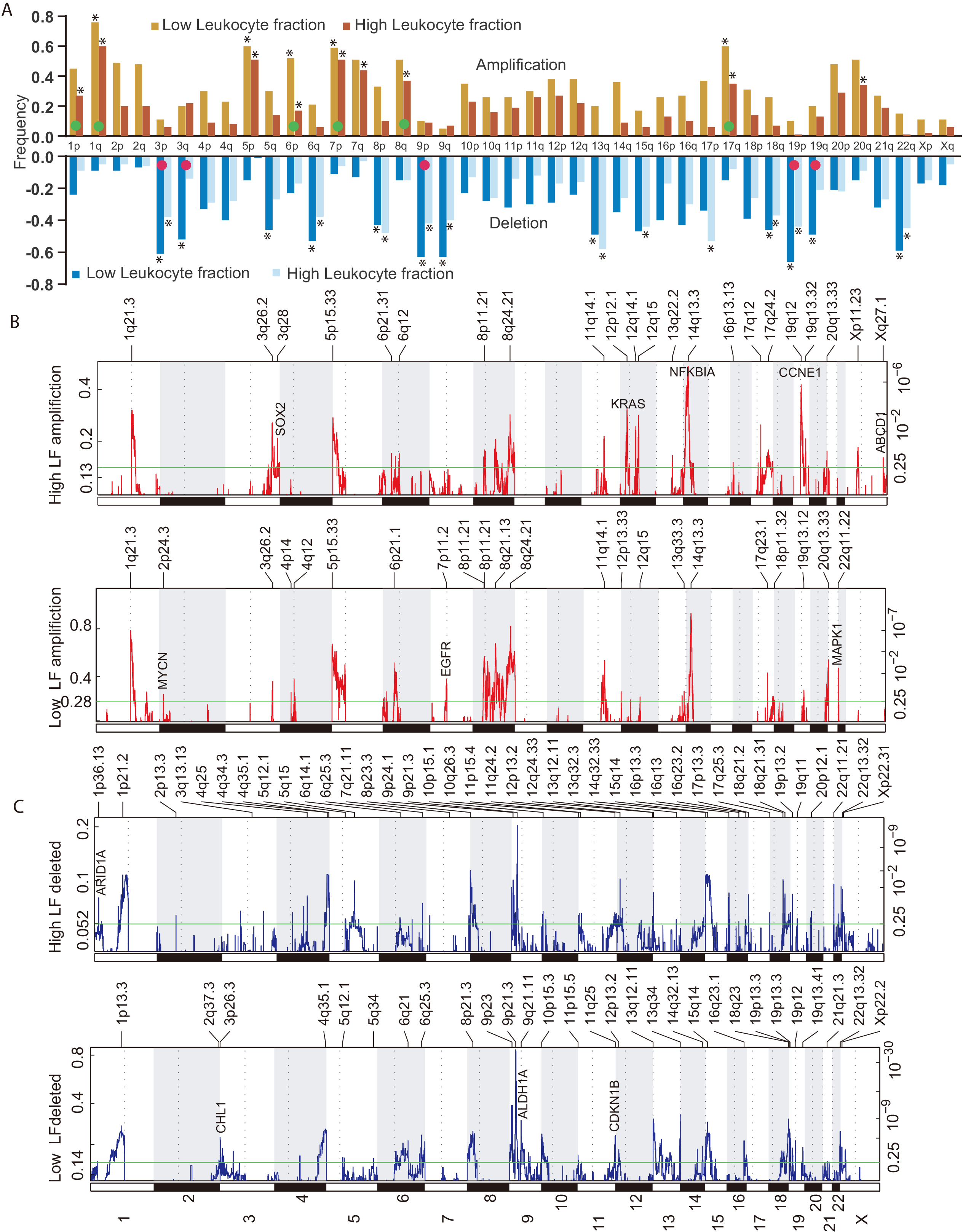
Methylation and expression landscape of LF in LUAD: A. The CpGs methylation was correlated with LF and CpGs with 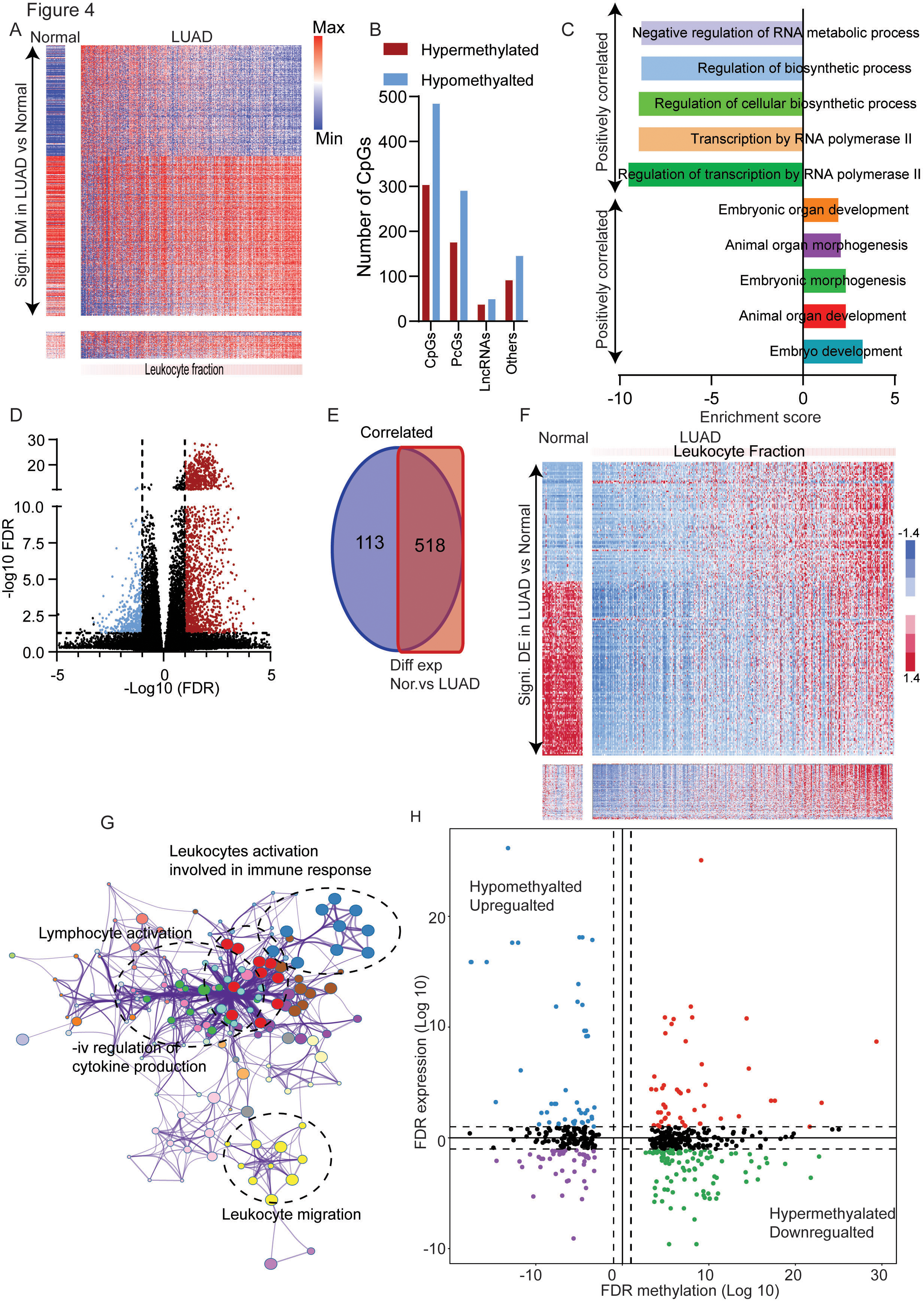
Development of immune risk signature for LUAD: A. Cox regression analysis was performed using the genes associated with LF and plotted as a volcano plot. The Red genes are negatively associated with survival, and blue genes are positively associated with survival. B. The six selected survival genes were used for GO analysis, and enriched terms are shown in the form of a bar plot. C. A heatmap of six prognostic immune genes in normal and LUAD. The LUAD patients are arranged in increasing order of LF. D. The IRS was calculated and plotted. The patients were divided into three groups based on the IRS. E. The LUAD patients were divided into low and high risk at the median, and the Kaplan-Meier plot was made. The survival was compared using log-rank analysis. F. The patients were divided into low, intermediate, and high immune risk groups, as shown in Fig. 4D, and the Kaplan-Meier plot was made. The survival of the groups were compared using the log-rank test. G. The LUAD patients with stage I were taken and divided at the median of the IRS, and the Kaplan-Meier plot was obtained. The survival was compared using the log-rank test. H. The univariate and multivariate Cox regression analysis was performed with the given variable, and a forest plot was prepared to show the results. 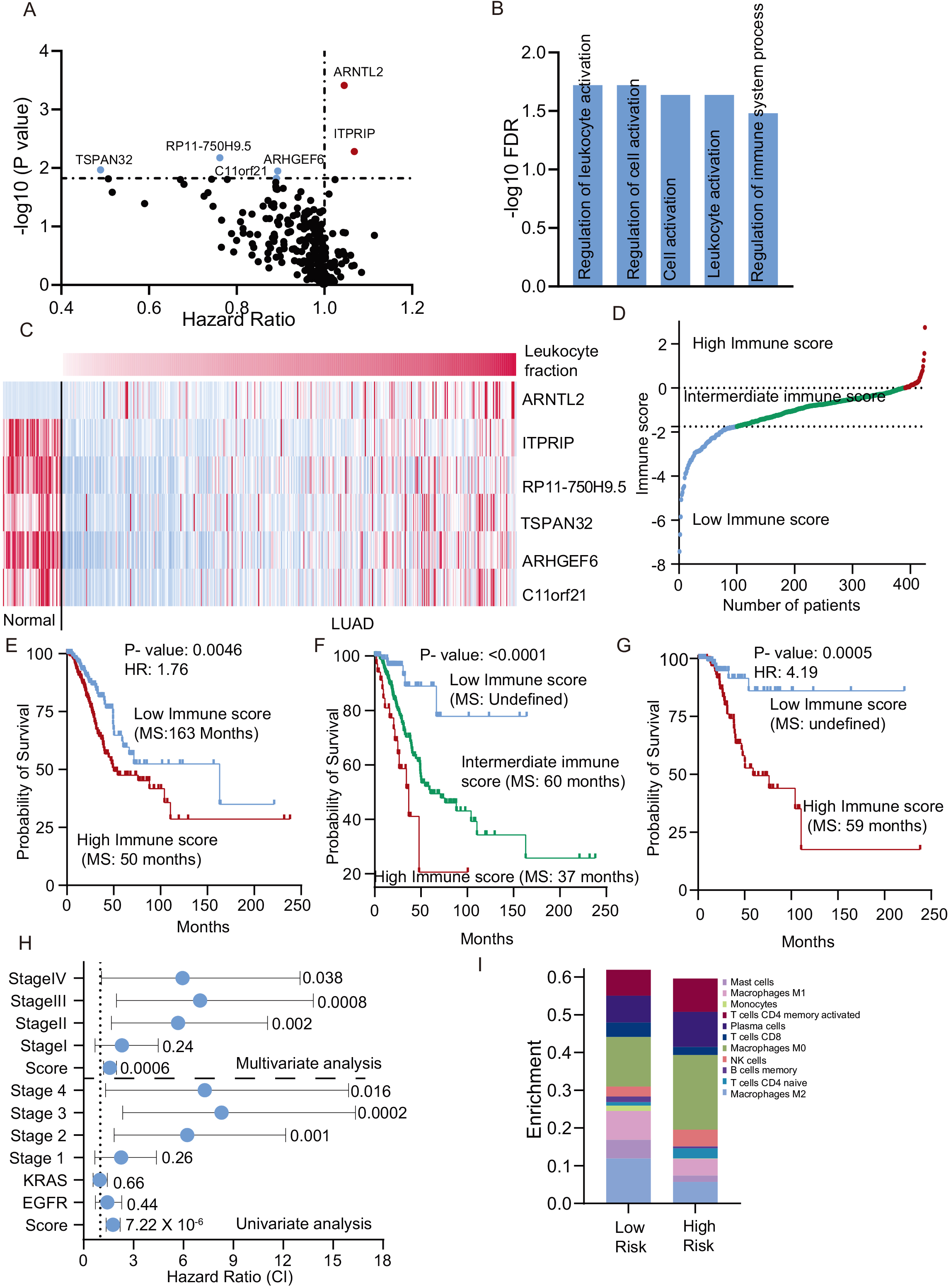
Validation of IRS in both the independent cohorts: A to E for GSE66728 and F to H for EGAD00001004421 cohort. A. Heatmap of six prognostic immune genes in the testing cohort and normal lung. B. The IRS was calculated using the expression of six genes in the testing cohort. The patients were divided into low, intermediate, and high immune risk groups. C. The testing cohort patients were divided into a high and low immune risk group, and the Kaplan-Meier plot was made. A Log-rank test was performed to compare survival. D. The training group patients were divided into three groups, and the Kaplan-Meier plot was made. A log-rank test was done to compare survival. E. Univariate and multivariate Cox regression analysis was performed in the testing cohort, and a forest plot was made to show the HR (CI). 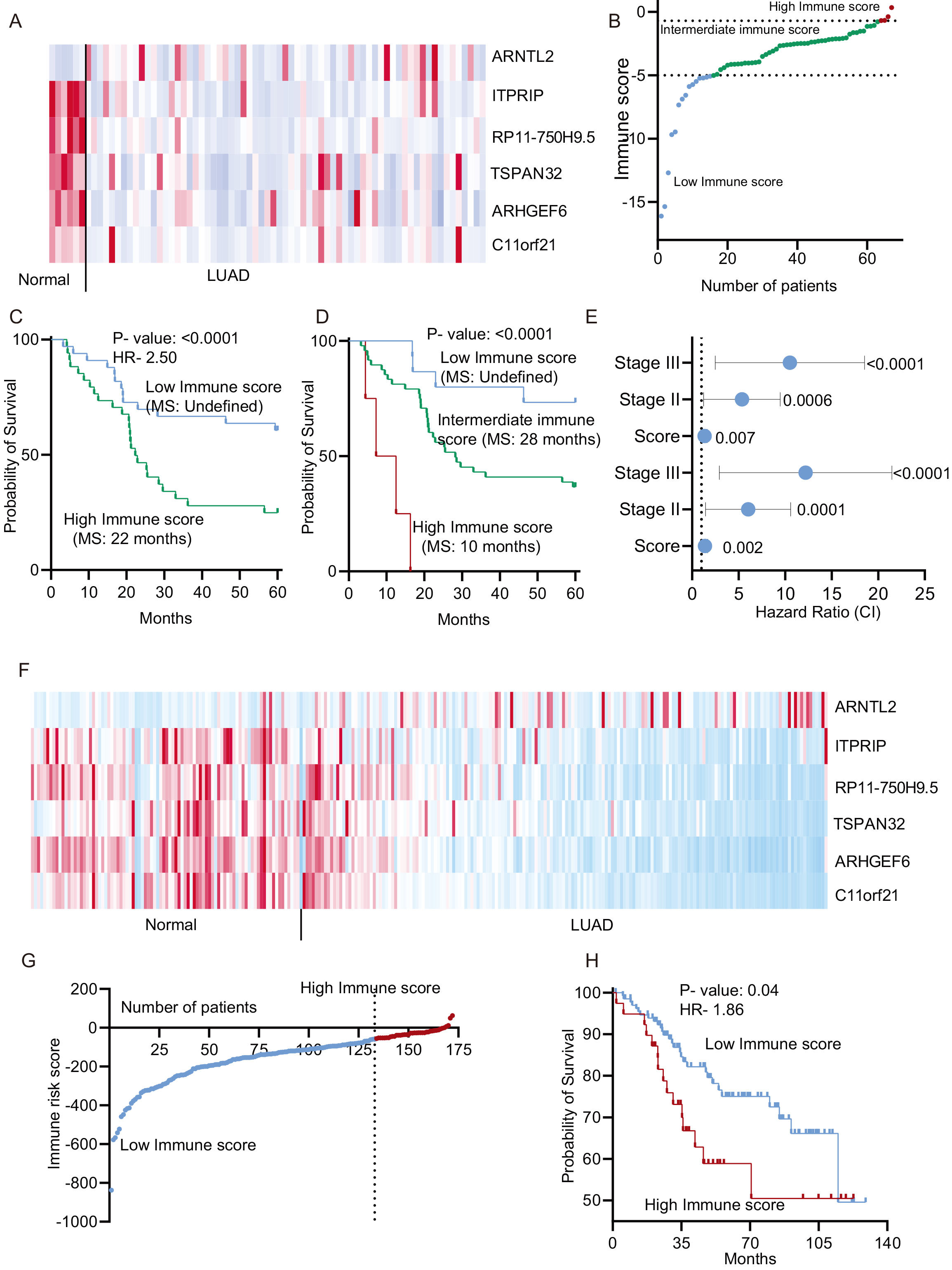
DNA methylation changes have a significant impact on the behaviour of the tumor, including the immune microenvironment [23]. In order to understand the role of DNA Methylation in the LF, the beta-values of CpG from TCGA-LUAD 450K-methylation data were correlated with the LF. A selection criterion (described in the method section) was used to identify 787 CpGs, the methylation level of which was strongly correlated with the LF (303 negative correlation and 484 positive correlation) (Fig. 4A and B). A major fraction (95.42%) of these CpGs have also been significantly differentially methylated in LUAD compared to normal lung samples (Fig. 4A) suggesting that changes in methylation pattern during carcinogenesis is an important factor in determining the immune cell infiltration of tumors. GO analysis showed that genes whose methylation was associated with a higher LF, regulated metabolism pathway associated with transcription, and genes whose methylation was correlated with a lower LF, were involved in the regulation of developmental pathways (Fig. 4C).
Further, we compared the expression pattern of the genes in low and high leukocyte using selection criteria of the correlation coefficient, and differential expression as explained in the method section. A total of 631 genes were identified as associated with LF in LUAD (Fig. 4D). Interestingly, the majority of genes were positively associated with LF, and more than 80% of these genes (
Development of LUAD prognostic model using LF associated genes
In our initial analysis, we have shown that the LF is not associated with survival of the LUAD patients (Fig. 1C). However, further investigation of genes whose expression was correlated with LF, led to the identification of prognostic genes in LUAD. To identify the genes which were strongly correlated with survival, unbiased Cox-regression analysis was performed as detailed in method section. The study identified 6 genes (ARNTL2, ITRIP-risk genes), (RP11-750H9.5, TSPAN32, ARHGEF6, and C11orf21-protective genes) with a strong association with survival (Fig. 5A, Supplementary Table 1). Further analysis showed that these prognostic genes regulate various immune cell regulatory pathways (Fig. 5B). Interestingly, ARNTL2 was downregulated, and all other genes were up regulated in LUAD compared to normal (Fig. 5C). Next, the expression value and Cox-regression correlation coefficient were used to calculate an immune risk score (IRS) (Material and method, and Fig. 5D). The patients were divided at the median, and the Log-rank test was performed. The analysis showed that patients with high IRS are at 1.75 times higher hazard than the patients with low IRS (Fig. 5E). Furthermore, the division of patients into three groups (high IRS
Validation of prognostic markers
It is crucial to validate the performance of prognostic markers in an independent dataset. Hence, we used another set of data set comprising 67 LUAD patients to test the prognostic strength of the IRS. First, we checked the expression of all the genes in normal (
Next, we used the second testing cohort of samples with 172 LUAD East Asian patients for the validation of immune risk score. First, we checked the expression of six genes in normal and LUAD samples and found that like other two cohort, expression of ARNTL2 was higher and other genes was lower in LUAD compared to normal samples (Fig. 6F). The risk score was calculated as described in method section and plotted (Fig. 6G). The Kaplan-Meier analysis was done at multiple cut off and the cut off at which low and high immune risk groups were found most significantly different was used for division (Fig. 6G and H). The analysis showed the significant poor survival of the high IRS patients compared to low IRS patients (Fig. 6H).
Discussion
The immune microenvironment of cancer plays a key role in patients’ behaviour towards therapy and overall outcomes [19, 24]. In our current research, multiple analyses were conducted to identify the genetic and epigenetic markers associated with the LF in LUAD tumors. Although leukocytes consist of all the different adaptive and innate immune response cells, our results have shown that the LF is positively associated with lymphocytes and M1 macrophages and negatively associated with dendritic cells, M2 macrophages, and mast cells. This result suggests that as more cells infiltrate the tumor, lymphocyte and tumor-promoting macrophages infiltrate, preferably. Other cells, such as dendritic cells, infiltrate more in tumors with a lower LF. The LF was not found to be associated with survival in LUAD tumors. Interestingly, we observed that significantly more female patients showed a high LF. This was a unique observation and needs further investigation. CD4
The genetic makeup of the tumor is a major influencer of the immune microenvironment [12, 28]. Correlation analysis has shown that intratumoral heterogeneity and copy number aberrations are significantly associated with the LF. Patients with high copy number aberration are less infiltrated by leukocytes. Overall mutations did not show any correlation with the LF in contrast. However, several individual mutations were associated with the LF. Importantly, clinically relevant mutations such as KRAS and KEAP1 have been more frequently mutated in patients with lower LFs. KRAS mutation has been associated with lower T cell infiltration [29]. These results suggest that individual mutation may play a role in defining the immune microenvironment. The LF was also significantly associated with overall changes in the copy number variation. Many focal CNV were also associated with LF.
In the recent past, DNA methylation has emerged as a major mechanism for regulating genes [30]. We hypothesized that changes in DNA methylation must also regulate the immune microenvironment. We identified 787 CpGs with a strong correlation with the LF. Majority of these CpGs were also differentially methylated in LUAD compared to normal. Such observations suggest that tumor DNA methylation shifts during carcinogenesis, and that the immune microenvironment is regulated according to the specific changes. It is noteworthy, in comparison to genes with a negative correlation, which regulated developmental pathways, that genes that correlated positively with leukocyte infiltration were enriched with transcription-related metabolism. We also observed vast differences in gene expression in patients with high and low LFs. Most of these genes have been differentially regulated in tumors in comparison with normal samples and regulated immune pathways, as have methylation changes. It is possible that changes in gene expression during tumor development regulate the infiltration of immune cells. Next, we used the LF related genes to develop and validate an independent and strong immune risk score for the prognostication of LUAD patients. The signature member genes showed an association with immune pathways. ARNTL2 is a core gene associated with circadian clock [31]. ARNTL2 has been associated with LUAD patients outcome and shown to regulate the pro-metastatic secretome [32]. ITRIP is a transmembrane protein which may be involved in immune modulation [33]. Similarly, TSPAN32 is a T cell associated gene which may function as tumor suppressor [34]. ARHGEF6 is a guanine nucleotide exchange factor for Rho protein which is required for medulloblastoma genesis [35]. In this analysis we also identified two novel prognostic genes RP11-7560H9.5 and C11orf21. The function of these genes is still unknown.
Conclusion
Taken together, we have identified genetic, epigenetic aberrations associated with immune cell infiltration in LUAD tumors. We showed that these changes in LUAD determine the genetic and epigenetic makeup during carcinogenesis and the immune microenvironment. Our data suggest that the genetic and epigenetic nature of the tumors is the significant determinant of immune surveillance in LUAD. We also developed and validated a strong and robust prognostic signature for survival prediction of LUAD patients.
Authors’ contribution
Conception: Sudhanshu Shukla
Interpretation or analysis of data: Sudhanshu Shukla and Seema Khadirnaikar
Preparation of the manuscript: Sudhanshu Shukla, Seema Khadirnaikar, Annesha Chatterjee
Revision for important intellectual content: Sudhanshu Shukla, Seema Khadirnaikar and Annesha Chatterjee
Supervision: Sudhanshu Shukla
All the authors have read and approved the manuscript.
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-203071.
sj-pdf-1-cbm-10.3233_CBM-203071.pdf - Supplemental material
Supplemental material, sj-pdf-1-cbm-10.3233_CBM-203071.pdf
Footnotes
Acknowledgments
The Cancer genome atlas (TCGA) is acknowledged for the clinical and genomic datasets. Sudhanshu Shukla is supported by the grants from DBT, Govt of India (BT/PR27478/MED/30/1954/2018) and SERB, DST, Govt. Of India (ECR/2018/000528) and CSIR, Govt of India (27(0366)/20/EMR-II.
