Abstract
Background:
EGFR mutation is a common oncogene driver in East Asians with lung adenocarcinoma (LUAD), conferring a favorable prognosis with effective targeted therapy. However, the EGFR mutation is a weak predictor of long-term survival. Therefore, a powerful predictive tool is urgently needed to estimate disease prognosis and patient survival for East-Asian patients with LUAD.
Methods:
In this first systematic analysis of the relationships among EGFR mutation, immunophenotype, and prognosis in LUAD samples from East-Asian patients, we constructed a prognostic signature consisting of EGFR-associated immune-related gene pairs (EIGPs). The predictive performance for overall survival (OS) and the clinical significance of this signature were then comprehensively investigated.
Results:
Based on transcriptome data analysis of a training set, we proposed the EIGP index (EIGPI), represented by five EIGPs, which was significantly associated with the OS of East-Asian patients with LUAD. It was also well validated in a test set. Furthermore, the prognostic performance of the EIGPI was further verified using protein levels in an additional independent set. Stratification analysis and multivariate Cox regression analysis revealed that the EIGPI was an independent prognostic factor. When combined with stage, the composite immune-clinical prognostic model index (ICPMI) showed improved prognostic accuracy in all datasets.
Conclusion:
This study was the first to systematically investigate the relationships among EGFR mutation, immunophenotype, and prognosis in East Asians with LUAD and develop a composite clinical and immune model associated with EGFR mutation. This model may be a reliable and promising prognostic tool and help further personalize patient management.
Introduction
As the most predominant type of lung malignancy, lung adenocarcinoma (LUAD) seriously endangers human health and life, with more than 1 million annual deaths worldwide.1,2 Although many established treatments, such as cytotoxic chemotherapy, molecular targeted therapy, and immunotherapy, have been applied in LUAD, patients’ long-term survival remains unsatisfied, with an average 5-year survival rate of 16%.3,4 Therefore, we need reliable biomarkers for estimating disease prognosis and patient survival to guide the curative management of LUAD.
Several studies have reported gene expression-based signatures for prognostic evaluation in patients with LUAD5–9 with the development of multi-omics profiling. Nevertheless, the prognostic signatures specifically for East-Asian populations with LUAD rarely are reported. LUAD differs between populations sampled from East-Asian and Western countries.10,11 For instance, LUAD tends to occur in male smokers in Western countries, while female never-smokers dominate East-Asian patients with LUAD. 12 Many large-scale genomic studies have found differences in the frequency of common somatic mutations.10,13–15 In Western countries, at least 35% of patients with LUAD have Kirsten rat sarcoma viral oncogene homolog (KRAS) mutations compared with 5–10% in East-Asian countries.16–18 Moreover, mutations in the epidermal growth factor receptor (EGFR) are found in 40–60% of East-Asian patients with LUAD, but only 7–10% of Caucasian patients.19–23 Recent advancements in targeted drugs against EGFR confer patients with LUAD and the EGFR mutation a favorable prognosis. Until now, KRAS-mutant patients with LUAD have a poor prognosis and are without effective targeted therapies.24–26 Thus, we need to develop a novel powerful discrimination tool specifically for East-Asian populations with LUAD to perform risk stratification.
Recently, there has been an increasing notion that the immune system plays an important part in cancer development.27,28 Cancers are often able to evade immune destruction, and diverse components of the immune system are key factors related to carcinogenesis and cancer progression.29–32 Also, many immune-associated parameters are able to predict prognosis in patients with LUAD.6,33–35 However, few studies have comprehensively investigated the immune-related characteristics regarding their prognostic potential in LUAD based on East-Asian patients’ data. Moreover, considering the high frequency of this mutation, the influences of EGFR mutation on the immune phenotype also require exploration.
We analyzed the immune phenotype related to EGFR mutation in LUAD samples from patients of East-Asian descent. Based on immune-related genes that were differentially expressed between EGFR wild-type and EGFR-mutant tumors, we then applied two gene expression datasets to build and validate a prognostic signature composed of EGFR-associated immune-related gene pairs (EIGPs). The signature’s prognostic power was also further validated using protein values in a cohort recruited from the Cancer Hospital/Institute, Chinese Academy of Medical Sciences (CICAMS). Also, to take full advantage of the complementary effect of clinical and molecular characteristics, we combined the immune signature and clinical parameters to construct a composite immune-clinical prognostic model index (ICPMI) according to the results of multivariate prognosis analyses. We also tested the predictive accuracy of ICPMI for overall survival (OS) in the validation dataset. Finally, we compared this composite signature with other existing prognostic predictors to verify this signature’s predictive robustness and reliability.
Materials and methods
Study design and sample collection
As shown in the analysis pipeline (Supplemental Figure S1), 574 LUAD patients from East-Asian countries (169 patients from the GIS2019 cohort for signature training, 226 patients from the GSE31210 cohort for signature testing, and 179 patients from the CICAMS cohort for signature validation) were included in development and validation of the prognostic model.
We collected the raw RNA-seq reads from 172 tumor tissue and 88 normal tissues, the normalized RSEM-estimated count data, the tumor mutation burden (TMB) data, and the genome instability index (GII) data from 169 tumor tissues of the GIS2019 cohort (https://src.gisapps.org/OncoSG_public/study/summary?id=GIS031). 10 To determine EGFR-associated immune-related genes (EIGs), we first identified 6223 genes that were differentially expressed between tumors and normal tissues at the thresholds of adjusting p-value < 0.05 and log2(fold change) >1 using the DESeq2 R package. 36 We then identified 649 differential genes between EGFR mutant and wild-type samples using the same method among these genes. Finally, 85 EIGs out of 2240 immune-related genes extracted from the Immport database (https://immport.niaid.nih.gov) were identified in the GIS2019 cohort. 37 Microarray data from the GSE31210 cohort were downloaded from the Gene Expression Omnibus (GEO, http://www.ncbi.nlm.nih.gov/geo) under accession number GSE31210. 38 The CICAMS cohort enrolled 179 LUAD patients who underwent radical operations in the Cancer Hospital, Chinese Academy of Medical Sciences (Beijing, China) from January 2012 to December 2013. None of the patients underwent preoperative treatments (e.g. chemotherapy, radiotherapy, and immunotherapy). Moreover, the paraffin-embedded samples and EGFR mutation status of all enrolled patients were available. The Ethics Committee of CICAMS approved this study (approval number CH-L-043). All enrolled patients signed the written informed consent form before the study, and the study had local ethics committee oversight. All corresponding characteristics of included patients and clinical outcomes in each cohort are shown in Supplemental Table S1.
Development and validation of a prognostic signature based on EIGPs
Among the 85 EIGs obtained from the GIS2019 cohort, 58 genes were measured across all datasets. We then use the 58 shared EIGs to construct 699 EIGPs by pairwise comparison. For each sample, the gene expression value underwent pairwise comparison to obtain a score (0 or 1) for each EIGP. For example, if the first gene’s expression value was higher than the second gene in an EIGP, the EIGP score was 1; otherwise, the EIGP score was 0. 6 This method depended entirely on an individualized tumor sample’s gene expression profile without the need for normalization. We then assessed the prognostic value of 699 EIGPs using univariate Cox proportional hazards regression modeling based on normalized RSEM-estimated count data in the GIS2019 dataset. Next, to minimize the risk of overfitting, we applied the least absolute shrinkage and selection operator (LASSO) Cox proportional hazards regression model (glmnet R software), and the minimum criteria were chosen. 39 To render prognostic signature more optimized and practical, we used a multivariate Cox proportional hazards regression model to select the EIGPs that formed the EIGP index (EIGPI) for prediction. EIGPI was calculated through the formula, which included weighting the score of the selected EIGPs by their respective coefficients. We then applied X-tile 3.6.1 software (Yale University, USA) to determine the optimal cutoff value to classify patients into EIGPI-high or EIGPI-low groups. The predictive ability of the novel EIGPI for OS was assessed in three independent cohorts using receiver operating characteristic curve (ROC) and Kaplan–Meier survival analyses. Notably, we generated the protein expression values of the selected EIGs using the immunohistochemistry (IHC) method. We also conducted univariate and multivariate Cox regression analyses to investigate whether EIGPI was an independent prognostic risk factor.
IHC analysis
We collected the paraffin-embedded samples of 179 LUAD patients to examine the protein levels of the selected 10 EIGs in the CICAMS cohort. Expression of 10 genes were assessed with IHC using an angiopoietin 4 (ANGPT4) assay (anti-human ANGPT4 rabbit recombinant polyclonal antibody, TA350852; OriGene, USA), a brain-derived neurotrophic factor (BDNF) assay (anti-human BDNF rabbit recombinant monoclonal antibody, ab108319; Abcam, USA), a fatty acid binding protein 7 (FABP7) assay (anti-human FABP7 rabbit polyclonal recombinant antibody, 14836-1-AP; Proteintech, USA), an inhibin, beta E (INHBE) assay (anti-human INHBE rabbit recombinant polyclonal antibody, ab254687; Abcam, USA), an oxytocin (OXT) assay (anti-human oxytocin rabbit recombinant monoclonal antibody, MAB5296; Millipore, USA), a Peptidase inhibitor 3 (PI3; SKALP) assay (anti-human SKALP rabbit polyclonal recombinant antibody, 15963-1-AP; Proteintech, USA), a S100 calcium binding protein A2 (S100A2) assay (anti-human S100A2 rabbit recombinant monoclonal antibody, ab109494; Abcam, USA), a TEK receptor tyrosine kinase (TEK; TIE2) assay (anti-human TIE2 rabbit polyclonal recombinant antibody, 19157-1-AP; Proteintech, USA), a semaphorin 3G (SEMA3G) assay (anti-human SEMA3G rabbit recombinant polyclonal antibody, TA322270; OriGene, USA), and a serpin family D member 1 (SERPIND1) assay (anti-human SERPIND1 rabbit recombinant polyclonal antibody, TA313999; OriGene, USA). All IHC slides were evaluated by two experienced pathologists blinded to clinical characteristics according to the evaluation criteria of the prior method.40–42 The staining score of every sample was calculated using the following formula: staining score = staining intensity × percentage of positive tumor cells × 100. Scoring based on the staining intensity: no color development was rated as 0 (negative), and color development intensity was pale yellow as 1 (weak), yellow as 2 (moderate), and brown-yellow as 3 (strong). Ten fields were randomly chosen under a high-power microscope (×400). The average value was taken to calculate the percentage of tumor cells that stain positively compared with all tumor cells in view. The results for representative staining images of the 10 genes at different levels are presented in Supplemental Figure S2.
Development and validation of ICPMI
According to the results of multivariate Cox regression analyses in the three cohorts, we integrated the EIGPI and stage to the ICPMI by using LASSO Cox proportional hazards regression model in the GIS2019 dataset. ICPMI was calculated by weighting the score of the selected parameters by their respective coefficients. Using the approach above for determining the cutoff of EIGPI, the optimal cutoff value for ICPMI was defined by X-tile software. The prognostic performance of the ICPMI was evaluated in three cohorts through ROC and Kaplan–Meier survival analyses. Univariate and multivariate Cox regression analyses were also performed to investigate whether ICPMI was an independent prognostic risk factor. Moreover, we compared the prognostic performance of ICPMI with EIGPI or other clinical factors in terms of the area under the curve (AUC) values of the ROC and the concordance index (C-index) to confirm the reliability and practical application of ICPMI.
Statistical analysis
We used R software (version 3.6.0) and GraphPad Prism software (version 5.0) to conduct the statistical analyses. Log-rank tests were used for between-group comparisons of survival curves from the Kaplan–Meier survival analyses. Chi-square and Mann–Whitney U tests were applied to statistical analyses between two groups. All reported p-values were double-tailed—notably, a p-value < 0.05 indicated a statistically significant result for all analyses, unless otherwise specified.
Results
Relationships of immune phenotype with EGFR mutation in LUAD samples from East Asians
Given that EGFR mutation is the most common oncogene driver in East Asians with LUAD, we investigated the association between EGFR status and the prognosis of patients in three independent cohorts. Here, we further verified that EGFR-mutant LUAD patients conferred a favorable prognosis (GIS2019: p = 0.0322; GSE31210: p = 0.0259; CICAMS: p < 0.001; Figure 1a–c). We then performed gene set enrichment analysis (GSEA) using gene expression data of LUAD samples from GIS2019 and GSE31210 to explore the distinct features of biological processes related to EGFR status. The GSEA results of GIS2019 revealed that, compared with the EGFR-mutant cases, EGFR wild-type tumors were significantly enriched in diverse immune pathways, including the complement pathway (NES = −1.54, p = 0.0180), the inflammatory response pathway (NES = −1.50, p = 0.0131), and the interferon-gamma response pathway (NES = −1.10, p = 0.0394) (Figure 1d). As expected, the GSEA results of GSE31210 also showed that EGFR wild-type LUADs were strongly related to positive regulation of immune-associated pathways compared with the mutant cases (Figure 1e).

Relationships of immune phenotype with EGFR mutation in LUAD samples from East Asians. (a–c) Kaplan–Meier survival curves of overall survival based on EGFR mutation status for LUAD patients from the GIS2019 cohort (a), the GSE31210 cohort (b), and the CICAMS cohort (c). (d and e) Compared with EGFR mutant LUADs, significant enrichment of immune-related pathways in LUADs without EGFR mutation in the GIS2019 dataset (d) and the GSE31210 dataset (e). NES: normalized enrichment score. (f) Volcano plot of 85 immune-related genes differentially expressed in LUAD samples with and without EGFR mutation. (g and h) GO analysis (g) and KEGG analysis (h) of the biological processes and pathways enriched for 85 immune-related genes.
To further identify the relationships between EGFR status and immune phenotype in East Asians with LUADs, we conducted an analysis of the expression profiles of 2240 immune-related genes extracted from the Immport database using LUAD cases from the GIS2019 cohort. Among these genes, 85 EIGs were differentially expressed in LUAD samples with and without EGFR-mutant tumors (Figure 1f). Next, we performed gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analyses to identify the biological processes and pathways associated with these 85 significant genes. The results of the GO analysis showed that the genes were mostly related to the humoral immune response process and the leukocyte migration process (Figure 1g). The KEGG analysis results revealed that the genes were mainly involved in the natural killer cell-mediated cytotoxicity pathway and the cytokine–cytokine receptor interaction pathway (Figure 1h).
Construction and definition of the EIGPI for East Asians with LUAD
Considering the significant differences in the immune phenotype between East-Asian patients with LUAD, with and without the EGFR mutation, we attempted to construct a prognostic signature based on the EIGs. Among 85 EIGs obtained from the GIS2019 cohort, 58 EIGs were shared by all datasets, and 699 EIGPs were constructed by pairwise comparison. Using univariate Cox proportional hazards regression modeling, we selected 69 prognostic EIGPs that were significantly associated with OS (p < 0.05). We then applied LASSO Cox proportional hazards regression modeling to identify gene pairs with the greatest predictive power. According to the minimum criteria, 17 EIGPs were selected (Figure 2a and b). Taking the prognostic signature’s optimized and practical value into consideration, multivariate Cox regression analysis was then conducted to further generate the EIGPI for prediction, and a novel prognostic signature consisting of only five gene pairs was built (Figure 2c). The five selected EIGPs and their coefficients are presented in Supplemental Table S2. Subsequently, the EIGPI score for each patient was calculated using the formula: EIGPI score = −1.140 × value of ANGPT4|BDNF + 1.125 × value of FABP7|INHBE −1.040 × value of OXT|PI3 +1.427 × value of S100A2|TEK + 1.102 × value of SEMA3G | SERPIND1. Based on the best cutoff value of 2.0, we then stratified patients into the EIGPI-high group (n = 23) and EIGPI-low group (n = 146).
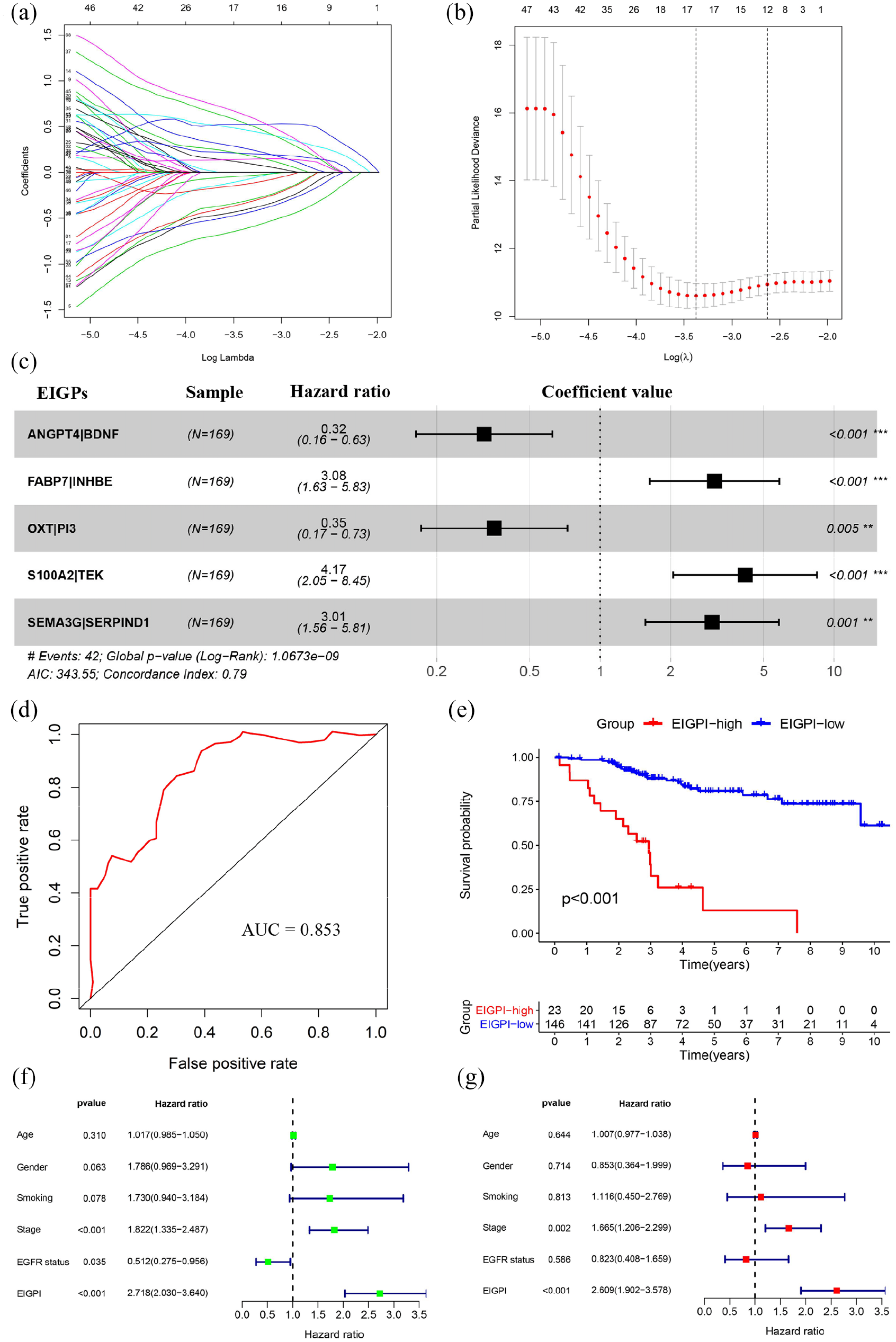
Construction and definition of the EIGPI for East Asians with LUAD. (a and b) LASSO Cox proportional hazards regression analysis identified 17 EIGPs most associated with overall survival. (c) Prognostic values of five selected EIGPs with multivariate Cox proportional hazards regression analysis. (d) ROC analysis of the EIGPI for overall survival. (e) Kaplan–Meier survival curve of overall survival for East Asians with LUAD based on the EIGPI. (f and g) Univariate (f) and multivariate (g) regression analyses of the relationships between EIGPI and clinical factors for the predictive value of overall survival.
To assess the predictive performance of the novel EIGPI, the AUC value of the ROC was calculated, and Kaplan–Meier survival analysis was performed. The results demonstrated that the AUC value at five-year OS was 0.853 (Figure 2d). Patients in the EIGPI-low group had significantly better long-term survival than those in the EIGPI-high group (p < 0.001; Figure 2e). Next, we performed univariate Cox regression analyses of the training dataset and found that EIGPI, stage, and EGFR status were predictor factors (EIGPI: p < 0.001; stage: p < 0.001; EGFR status: p = 0.035; Figure 2f). The results after adjusting for clinicopathological factors indicated that both EIGPI and stage were independent prognostic factors (EIGPI: p < 0.001; stage: p = 0.002; Figure 2g). However, the AUC value of EIGPI was greater than the stage (Supplemental Figure S3F).
Validation of the EIGPI for East Asians with LUAD
To confirm the discriminatory power of EIGPI for East Asians with LUAD, we applied the same formula to the test dataset from the GSE31210 cohort consisting of 226 LUADs. The 226 patients were then stratified into the EIGPI-high group (n = 80) and EIGPI-low group (n = 146) based on the training dataset’s cutoff value. As expected, the fact that its AUC value at a five-year OS was 0.719 revealed that the EIGPI for East Asians with LUAD in the test dataset was a reliable predictive signature (Figure 3a). Kaplan–Meier survival analysis suggested that patients with the EIGPI-high score had significantly worse OS than those with the EIGPI-low score (p < 0.001; Figure 3b). In addition, the results of univariate and multivariate Cox regression analyses showed that both EIGPI and stage were not only predictor factors (EIGPI: p < 0.001; stage: p < 0.001; Figure 3c) but also were the significant independent predictor factors of OS (EIGPI: p = 0.007; stage: p = 0.004; Figure 3d). Consistent with the prior results, the AUC value of EIGPI was greater than the stage (Supplemental Figure S4E).

Validation of the EIGPI for East Asians with LUAD. (a) ROC analysis of the EIGPI for overall survival in the GSE31210 set. (b) Kaplan–Meier survival curve of overall survival for East Asians with LUAD based on the EIGPI in the GSE31210 set. (c and d) Univariate (c) and multivariate (d) Cox regression analyses of relationships between EIGPI and clinical factors for the predictive value of overall survival in the GSE31210 set. (e) ROC analysis of the EIGPI for overall survival in the CICAMS set. (f) Kaplan–Meier survival curve of overall survival for East Asians with LUAD based on the EIGPI in the CICAMS set. (g and h) Univariate (g) and multivariate (h) Cox regression analyses of relationships between EIGPI and clinical factors for the predictive value of overall survival in the CICAMS set.
To further verify the reliability and practical values of EIGPI, we instigated its prognostic performance using protein expression values in an independent cohort (CICAMS) of 179 LUAD patients. In terms of each sample, the protein expression values of 10 EIGs underwent pairwise comparison to obtain a score (0 or 1) for each EIGP. The EIGPI score of each patient was then calculated using the formula above. Notably, the EIGPI for East Asians with LUAD was a robust prognostic signature at the protein level; the AUC value for 5 years of OS was 0.836 (Figure 3e). We then assigned the 179 patients into an EIGPI-high-group (n = 29) and an EIGPI-low group (n = 150) according to the same cutoff value. Kaplan–Meier survival analysis showed a remarkable difference in long-term survival between the two groups (p < 0.001; Figure 3f). We also conducted univariate and multivariate Cox regression analyses and found that EIGPI, stage, and EGFR status were not only predictor factors (EIGPI: p < 0.001; stage: p < 0.001; EGFR status: p < 0.001; Figure 3g) but also were the significant independent predictor factors of OS (EIGPI: p < 0.001; stage: p < 0.001; EGFR status: p = 0.026; Figure 3h). However, the AUC value of EIGPI was greater than the stage and EGFR status (Supplemental Figure S5G).
Stratification analyses of OS for the EIGPI according to clinical factors
Considering that the EGFR mutation was commonly mutated and a weak predictor of long-term survival probability in East Asians with LUAD, we analyzed the prognostic performance of EIGPI in patients with different EGFR mutation status. The results of Kaplan–Meier survival analysis in the GIS2019 cohort showed that among different mutation status, long-term survival times in EIGPI-high groups were remarkably shorter than those of EIGPI-low groups (p < 0.001; Supplemental Figure S3A and S3B). When it comes to the GSE31210 and CICAMS cohorts’ eternal validation sets, all these findings were also well confirmed (p < 0.001; Supplemental Figure S4A, S4B, S5A, and S5B).
Given that clinical staging system was the independent prognostic factors of OS in the above findings, we stratified LUAD patients by stage in the three independent cohorts. Results revealed that in all subgroups, including stage I, stage II, stage III, and stage IV subgroups, patients in the EIGPI-high groups had poorer OS times than those in the EIGPI-low groups (p < 0.05; Supplemental Figure S3C–E, S4C–D, and S5C–F). Notably, we also found that the p-values of stages-specific Kaplan–Meier survival curves in each staging subgroup were not always the same. The statistical significance of stages-specific Kaplan–Meier survival curves in advanced stages was always more reliable.
Composite prognostic model by combining the EIGPI with stage
In multivariate analyses of three independent sets, both EIGPI and stage were independent prognostic risk factors, suggesting a complementary value. Therefore, to further improve the accuracy of our signature, we combined the EIGPI score and stage to build the ICPMI by applying LASSO Cox proportional hazards regression model in the training set (Figure 4a and b). Subsequently, ICPMI was derived as (0.926 × EIGPI score) + (0.449 × stage). An optimal cutoff value of 2.637 for assigning patients was determined based on X-tile software analysis in the training set. According to the above formula, after stratifying patients in three cohorts, the prognostic performance of the ICPMI was evaluated through ROC and Kaplan–Meier survival analyses. The ICPMI for East Asians with LUAD was identified as a reliable prognostic model; their AUC values at a five-year OS were 0.858 for the GIS2019 cohort, 0.740 for the GSE31210 cohort, and 0.850 for the CICAMS cohort (Figure 4c, 5a, and 5e). Kaplan–Meier survival analyses demonstrated that patients with the ICPMI-high score had significantly worse OS times than those with the ICPMI-low score (p < 0.001; Figure 4d, 5b, and 5f). Moreover, the results of multivariate Cox regression analyses suggested that the prognostic ability of ICPMI is an independent predictor of OS when adjusted for age, sex, smoking, and EGFR status (p < 0.001; Figure 4f, 5d, and 5h). Stratified analyses of all subgroups, including EGFR mutant and wild-type subgroup—as well as stage I, stage II, stage III, and stage IV subgroups—revealed that the ICPMI-high score identified high-risk patients with shorter long-term survival times (p < 0.05; Supplemental Figure S6, S7A–S7D, and Figure S8).
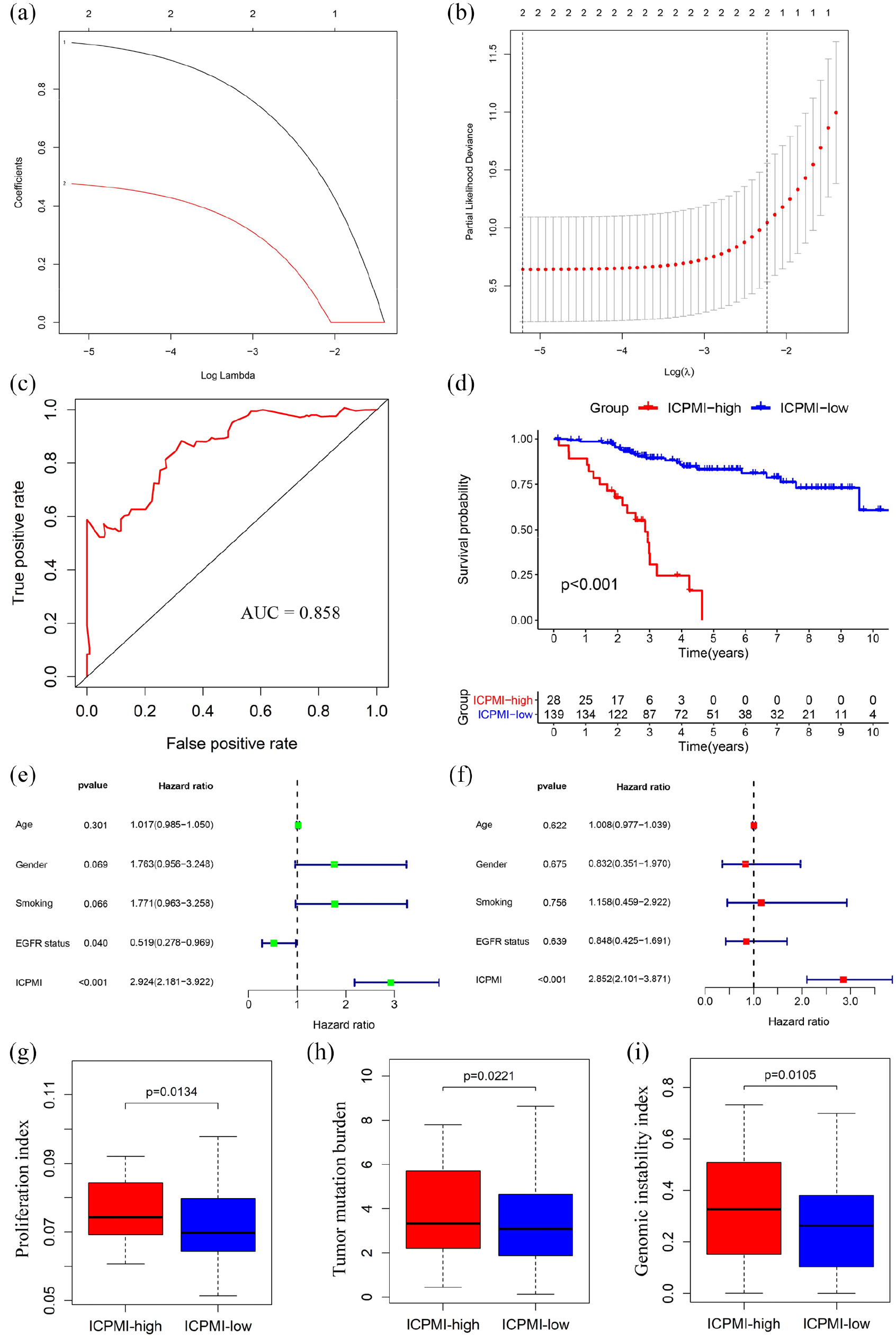
ICPMI by combining the EIGPI with a stage in the training set. (a and b) LASSO Cox proportional hazards regression analysis determined the EIGPI with stage most related to overall survival. (c) ROC analysis of the ICPMI for overall survival. (D) Kaplan–Meier survival curve of overall survival for East Asians with LUAD based on the ICPMI. (e and f) Univariate (e) and multivariate (f) regression analyses of the associations between ICPMI and clinical factors for the predictive value of overall survival. (g–i) Phenotypic differences between the two groups (ICPMI-high and ICPMI-low) focusing on proliferation-related features (g), tumor mutation burden (h), and genome instability index (i).

Validation of the ICPMI for East Asians with LUAD. (a) ROC analysis of the ICPMI for overall survival in the GSE31210 set. (b) Kaplan–Meier survival curve of overall survival for East Asians with LUAD based on the ICPMI in the GSE31210 set. (c and d) Univariate (c) and multivariate (d) Cox regression analyses of associations between ICPMI and clinical factors for the predictive value of overall survival in the GSE31210 set. (e) ROC analysis of the ICPMI for overall survival in the CICAMS set. (f) Kaplan–Meier survival curve of overall survival for East Asians with LUAD based on the ICPMI in the CICAMS set. (g and h) Univariate (g) and multivariate (h) Cox regression analyses of associations between ICPMI and clinical factors for the predictive value of overall survival in the CICAMS set.
LUAD is characterized by features closer to normal tissue, including downregulation of proliferation-associated pathways, lower TMB, and lower GII. These patients had better long-term survival. 10 Therefore, we sought to gain new insights into the associations of ICPMI with the features using the corresponding data from the GIS2019 dataset. The proliferation index was calculated with imsig R package (version.1.0.0), which used a set of gene signatures generated by a network-based deconvolution approach. 43 The TMB and GII data were downloaded from the GIS2019 database. The results indicated that higher levels of proliferation index, TMB, and GII were expressed in the ICPMI-high group in the GIS2019. This explained the rationality of our signature to some extent (proliferation index: p = 0.0134; TMB: p = 0.0221; GII: p = 0.0105; Figure 4g–i). Furthermore, owing to no genomic data in the GSE31210, we only analyzed the correlation between ICPMI and proliferation-associated features. As expected, the ICPMI-high group had a higher level of proliferation index in the GSE31210 (Supplemental Figure S7E).
In addition, we performed the subgroup analysis of the patient characteristics according to the risk stratification of EIGPI and ICPMI. As expected, the significant differences in OS based on the risk stratification of EIGPI and ICPMI were shown in all the three cohorts (Supplemental Table S3). In addition, EGFR status also showed significant differences based on the risk stratification of EIGPI and ICPMI in the three cohorts. Notably, two groups divided by EIGPI had no significant differences in stage, but there were significant differences in stage according to the risk stratification of ICPMI.
Comparison of the ICPMI and other existing prognostic factors
Since the prognostic significance of EGFR status, stage, EIGPI, and ICPMI were well confirmed in the three independent cohorts, we sought to explore whether ICPMI is the strongest predictor of OS of East Asians with LUAD. Therefore, to compare the prognostic performance of ICPMI with other predictors, we first calculated the AUC values of the ROC from four parameters in three datasets. The AUC value of ICPMI was greater than other prognostic factors (Figure 6a–c). For the C-index, ICPMI more accurately predicted long-term survival in all datasets (Figure 6d–f).

Comparison of the ICPMI and other existing prognostic factors. (a–c) The prognostic performance was compared between ICPMI and other existing prognostic factors according to ROC analyses in the GIS2019 cohort (a), the GSE31210 cohort (b), and the CICAMS cohort (c). (e–g) The C-index was applied to evaluate the predictive performance of ICPMI with other existing prognostic factors for survival prediction in the GIS2019 cohort (e), the GSE31210 cohort (f), and the CICAMS cohort (g).
Discussion
Various prognostic predictors for LUAD have been continually proposed as the prevalence of high-throughput sequencing technology for cancer research has increased.5–9 However, the prognostic signatures specifically for East-Asian populations with LUAD rarely are reported. As we all know, racial disparities have long been recognized in LUAD, and are explained by intrinsic genetic predisposition and other environmental factors.44–48 Therefore, a reliable predictive tool is urgently needed to estimate disease prognosis and patient survival in East Asians with LUAD to help personalize patient management.
The EGFR mutation, the most common oncogene driver in East Asians with LUAD, has been recently researched as a prognostic biomarker.49–51 We performed a comprehensive investigation of EGFR mutation in regulating immune phenotype in LUAD samples from East-Asian patients. We found that EGFR wild-type LUADs exhibited better regulation of immune-associated pathways than EGFR-mutant tumors in the current study. However, a recent study reported that patients with LUAD and no EGFR mutation showed a higher proportion of CD8+ T cells and higher levels of PD-L1 compared with the EGFR-mutant patients. 52 Notably, patients with LUAD who presented with de novo resistance to EGFR–tyrosine kinase inhibitor (TKI) therapy exhibited strong PD-L1 expression.53,54 Therefore, patients with LUAD and no EGFR mutation confer an unfavorable prognosis due to the lack of effective targeted therapies. Among patients with EGFR-mutant LUAD, better prognosis also depended on the higher frequency of low-grade tumors, such as adenocarcinoma in situ and minimally invasive adenocarcinoma, which rarely recurred after curative resection. 55
Given that the fact that patients with LUAD and the EGFR mutation had a better prognosis than those without the EGFR mutation, the EGFR mutation appears to be a reliable prognosis biomarker. Unfortunately, we found that the EGFR mutation only weakly predicted long-term survival in three independent cohorts. The contributions of the immune system to carcinogenesis and cancer progression have been increasingly acknowledged in recent years.29–32 Many immune-associated factors also exhibit a tremendous prognostic estimation value in patients with LUAD.6,33–35 Therefore, to leverage the complementary value of EGFR mutation and immune-associated genes, we constructed a novel immune signature (i.e. EIGPI) significantly associated with the OS of East Asians with LUAD. This signature was based on immune-related genes that were differentially expressed between EGFR wild-type and EGFR-mutant LUADs in the training set. This signature was well validated in a test set. The discriminatory power of the EIGPI was further validated using protein values, which were obtained with the IHC method, in an additional independent cohort. Notably, owing to its simple technology and low cost, the IHC technique might be more suitable for clinical application. Next, based on the multivariate analysis results, we further leveraged the complementary value of EIGPI and stage. We found that integrating the two could provide higher accuracy of long-term survival estimation in East Asians with LUAD.
Considering the technical biases inherent across different platforms with RNA-seq, microarray, or IHC data, our prognostic signature (i.e. EIGPI) was derived from the relative ranking of gene or protein expression values of a sample. This eliminated the need for the scaling and normalization of data. Furthermore, our signature provided a specific formula for calculating the EIGPI score and the cutoff value for risk stratification. This could be applied across multiple datasets from different platforms. Based on these advantages, and given the same formula and cutoff value in the test set, our prognostic signature also reached a consistent result. We therefore believe our signature is poised for translation into clinical practice as a promising predictor of survival in patients with LUAD.
Although multivariate Cox analysis found that EIGPI was an independent predictive factor for OS in East Asians with LUAD, our prognostic model (i.e. ICPMI), which integrated the EIGPI and stage, possessed a higher predictive efficacy and accuracy for long-term survival. As expected, ICPMI was also an independent prognostic factor based on multivariate Cox analysis. Notably, strong prognostic performance in important clinical subgroups of LUAD patients could make the signature more suitable for clinical use. Therefore, considering that both EGFR mutation and stage were predictors of long-term survival probability in East Asians with LUAD, we tested the prognostic performance of ICPMI in all subgroups, including EGFR mutant and wild-type subgroup—as well as stage I, stage II, stage III, and stage IV subgroups—and found that the signature was well validated. Interestingly, the prognostic performance of ICPMI was more powerful in advanced stage, which might be because advanced EGFR mutant LUAD patients tended to receive EGFR-TKIs treatments. We also compared the robustness of ICPMI with other existing prognostic factors, including EGFR status, stage, and EIGPI. We found that ICPMI achieved a more accurate prediction of long-term survival in all datasets. These results increase our confidence that ICPMI will be an effective future prognostic tool.
Our study’s limitations, including its retrospective nature, should be acknowledged, although we tried to include as many datasets from different and unusual platforms as possible for stricter validation of our signature.
To gain new insights into the potential underlying mechanism of ICPMI that stratified high-risk and low-risk patients, we investigated the relationships of ICPMI with the features closer to normal tissue, including proliferation index, TMB, and GII. We found that high-risk patients with ICPMI-high scores showed higher proliferation index levels, TMB, and GII in the GIS2019. This finding justifies use of our signature for prognosis estimation.
In conclusion, this study was the first to systematically analyze the relationships among EGFR mutation, immune phenotype, and prognosis in East Asians with LUAD. We also proposed a composite clinical and immune model associated with EGFR mutation, which may be a powerful prognostic tool and help further optimize the personalized cancer therapy paradigm.
Supplemental Material
sj-docx-10-tam-10.1177_17588359211006949 – Supplemental material for Development and external validation of a composite immune-clinical prognostic model associated with EGFR mutation in East-Asian patients with lung adenocarcinoma
Supplemental material, sj-docx-10-tam-10.1177_17588359211006949 for Development and external validation of a composite immune-clinical prognostic model associated with EGFR mutation in East-Asian patients with lung adenocarcinoma by Chengming Liu, Sufei Zheng, Sihui Wang, Xinfeng Wang, Xiaoli Feng, Nan Sun and Jie He in Therapeutic Advances in Medical Oncology
Supplemental Material
sj-docx-11-tam-10.1177_17588359211006949 – Supplemental material for Development and external validation of a composite immune-clinical prognostic model associated with EGFR mutation in East-Asian patients with lung adenocarcinoma
Supplemental material, sj-docx-11-tam-10.1177_17588359211006949 for Development and external validation of a composite immune-clinical prognostic model associated with EGFR mutation in East-Asian patients with lung adenocarcinoma by Chengming Liu, Sufei Zheng, Sihui Wang, Xinfeng Wang, Xiaoli Feng, Nan Sun and Jie He in Therapeutic Advances in Medical Oncology
Supplemental Material
sj-docx-12-tam-10.1177_17588359211006949 – Supplemental material for Development and external validation of a composite immune-clinical prognostic model associated with EGFR mutation in East-Asian patients with lung adenocarcinoma
Supplemental material, sj-docx-12-tam-10.1177_17588359211006949 for Development and external validation of a composite immune-clinical prognostic model associated with EGFR mutation in East-Asian patients with lung adenocarcinoma by Chengming Liu, Sufei Zheng, Sihui Wang, Xinfeng Wang, Xiaoli Feng, Nan Sun and Jie He in Therapeutic Advances in Medical Oncology
Supplemental Material
sj-docx-9-tam-10.1177_17588359211006949 – Supplemental material for Development and external validation of a composite immune-clinical prognostic model associated with EGFR mutation in East-Asian patients with lung adenocarcinoma
Supplemental material, sj-docx-9-tam-10.1177_17588359211006949 for Development and external validation of a composite immune-clinical prognostic model associated with EGFR mutation in East-Asian patients with lung adenocarcinoma by Chengming Liu, Sufei Zheng, Sihui Wang, Xinfeng Wang, Xiaoli Feng, Nan Sun and Jie He in Therapeutic Advances in Medical Oncology
Supplemental Material
sj-tif-1-tam-10.1177_17588359211006949 – Supplemental material for Development and external validation of a composite immune-clinical prognostic model associated with EGFR mutation in East-Asian patients with lung adenocarcinoma
Supplemental material, sj-tif-1-tam-10.1177_17588359211006949 for Development and external validation of a composite immune-clinical prognostic model associated with EGFR mutation in East-Asian patients with lung adenocarcinoma by Chengming Liu, Sufei Zheng, Sihui Wang, Xinfeng Wang, Xiaoli Feng, Nan Sun and Jie He in Therapeutic Advances in Medical Oncology
Supplemental Material
sj-tif-2-tam-10.1177_17588359211006949 – Supplemental material for Development and external validation of a composite immune-clinical prognostic model associated with EGFR mutation in East-Asian patients with lung adenocarcinoma
Supplemental material, sj-tif-2-tam-10.1177_17588359211006949 for Development and external validation of a composite immune-clinical prognostic model associated with EGFR mutation in East-Asian patients with lung adenocarcinoma by Chengming Liu, Sufei Zheng, Sihui Wang, Xinfeng Wang, Xiaoli Feng, Nan Sun and Jie He in Therapeutic Advances in Medical Oncology
Supplemental Material
sj-tif-3-tam-10.1177_17588359211006949 – Supplemental material for Development and external validation of a composite immune-clinical prognostic model associated with EGFR mutation in East-Asian patients with lung adenocarcinoma
Supplemental material, sj-tif-3-tam-10.1177_17588359211006949 for Development and external validation of a composite immune-clinical prognostic model associated with EGFR mutation in East-Asian patients with lung adenocarcinoma by Chengming Liu, Sufei Zheng, Sihui Wang, Xinfeng Wang, Xiaoli Feng, Nan Sun and Jie He in Therapeutic Advances in Medical Oncology
Supplemental Material
sj-tif-4-tam-10.1177_17588359211006949 – Supplemental material for Development and external validation of a composite immune-clinical prognostic model associated with EGFR mutation in East-Asian patients with lung adenocarcinoma
Supplemental material, sj-tif-4-tam-10.1177_17588359211006949 for Development and external validation of a composite immune-clinical prognostic model associated with EGFR mutation in East-Asian patients with lung adenocarcinoma by Chengming Liu, Sufei Zheng, Sihui Wang, Xinfeng Wang, Xiaoli Feng, Nan Sun and Jie He in Therapeutic Advances in Medical Oncology
Supplemental Material
sj-tif-5-tam-10.1177_17588359211006949 – Supplemental material for Development and external validation of a composite immune-clinical prognostic model associated with EGFR mutation in East-Asian patients with lung adenocarcinoma
Supplemental material, sj-tif-5-tam-10.1177_17588359211006949 for Development and external validation of a composite immune-clinical prognostic model associated with EGFR mutation in East-Asian patients with lung adenocarcinoma by Chengming Liu, Sufei Zheng, Sihui Wang, Xinfeng Wang, Xiaoli Feng, Nan Sun and Jie He in Therapeutic Advances in Medical Oncology
Supplemental Material
sj-tif-6-tam-10.1177_17588359211006949 – Supplemental material for Development and external validation of a composite immune-clinical prognostic model associated with EGFR mutation in East-Asian patients with lung adenocarcinoma
Supplemental material, sj-tif-6-tam-10.1177_17588359211006949 for Development and external validation of a composite immune-clinical prognostic model associated with EGFR mutation in East-Asian patients with lung adenocarcinoma by Chengming Liu, Sufei Zheng, Sihui Wang, Xinfeng Wang, Xiaoli Feng, Nan Sun and Jie He in Therapeutic Advances in Medical Oncology
Supplemental Material
sj-tif-7-tam-10.1177_17588359211006949 – Supplemental material for Development and external validation of a composite immune-clinical prognostic model associated with EGFR mutation in East-Asian patients with lung adenocarcinoma
Supplemental material, sj-tif-7-tam-10.1177_17588359211006949 for Development and external validation of a composite immune-clinical prognostic model associated with EGFR mutation in East-Asian patients with lung adenocarcinoma by Chengming Liu, Sufei Zheng, Sihui Wang, Xinfeng Wang, Xiaoli Feng, Nan Sun and Jie He in Therapeutic Advances in Medical Oncology
Supplemental Material
sj-tif-8-tam-10.1177_17588359211006949 – Supplemental material for Development and external validation of a composite immune-clinical prognostic model associated with EGFR mutation in East-Asian patients with lung adenocarcinoma
Supplemental material, sj-tif-8-tam-10.1177_17588359211006949 for Development and external validation of a composite immune-clinical prognostic model associated with EGFR mutation in East-Asian patients with lung adenocarcinoma by Chengming Liu, Sufei Zheng, Sihui Wang, Xinfeng Wang, Xiaoli Feng, Nan Sun and Jie He in Therapeutic Advances in Medical Oncology
Footnotes
Acknowledgements
We are grateful to all the participants who have made this research possible.
Authors’ contributions
L.C.M designed the study, performed experiments, analyzed data, and wrote the manuscript. Z.S.F, W.S.H, W.X.H and F.X.L performed experiments and analyzed data. H.J and S.N conceived and designed the study and wrote the manuscript.
Availability of data and materials
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Conflict of interest statement
The authors declare that there is no conflict of interest.
Consent for publication
The manuscript is approved by all authors for publication.
Ethics approval and consent to participate
The human tissue study protocol was approved by the Ethics Committee of Cancer Hospital, Chinese Academy of Medical Sciences, and Peking Union Medical College (Beijing, China). The approval number was CH-L-043. All formalin-fixed and paraffin-embedded (FFPE) NSCLC tissue samples were obtained using informed consent and agreement.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The National Natural Science Foundation of China (grant numbers 81802299, 81502514), the Fundamental Research Funds for the Central Universities (grant number 3332018070), the CAMS Innovation Fund for Medical Sciences (CIFMS) (grant numbers 2016-I2M-1-001, 2017-I2M-1-005), the National Key R&D Program of China (grant numbers 2018YFC1312100, 2018YFC1312102), the National Key Basic Research Development Plan (grant number 2018YFC1312105), and the Graduate Innovation Funds of Peking Union Medical College (grant number 2019-1002-06).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
