Abstract
By using a meta-analytical approach, this study aimed to analyse the diagnostic capacity of protein-based biomarkers in saliva for the differential diagnosis of oral potentially malignant disorders (OPMDs) and oral squamous cell carcinoma (OSCC) from healthy individuals as control group (HCG).
Articles on protein-based biomarkers in saliva, which provided quantitative expression in individuals with clinical and histopathological diagnosis of OPMD or oral leukoplakia (OL) were considered eligible. Searches were conducted in eight electronic databases. The methodological quality was assessed using the Quality Assessment of Diagnostic Studies tool (QUADAS-2). Functional analysis was also performed. Meta-analyses were performed using the OpenMeta tool (Analyst).
Meta-analysis was possible for 4 of the 11 biomarkers studied. Only the carcinoembryonic antigen (CEA) and the soluble fragment of cytokeratin 19 (CYFRA21) were significant for the OSCC/OPMD subgroup, both with a very low heterogeneity. CEA had an OE
The CEA and CYFRA21 markers proved very useful when differentiating OSCC from OPMD. The CYFRA21 was the only protein that was capable of distinguishing between OPMD and healthy controls.
Keywords
Introduction
The oral squamous cell carcinoma (OSCC) represents more than 95% of all malignant neoplasms of the mouth [1, 2] and it has a high prevalence and morbidity, with 354,900 new cases and 177,400 deaths registered per year worldwide [3]. Despite advances in the understanding and treatment protocols for different malignant tumours, the five-year survival rate for patients with OSCC does not exceed 50–60% overall [1, 4]. As a result, prevention and early diagnosis are considered to be highly necessary in order to improve the prognosis and survival rates.
The oral potentially malignant disorders (OPMD) are clinical conditions with a variable malignant transformation capacity. This group of lesions is comprised of leukoplakia, erythroplakia, erythroleukoplakia, lichen planus, nicotinic stomatitis, actinic cheilitis, amongst others [2, 5, 6]. The OL is the main lesion within the OPMD group, with an annual global average of malignant transformation rate of between 1 and 2% [2, 5, 7], nonetheless, these estimations may be higher, depending on the geographical region observed and the level of dysplasia, therefore meaning that it is possible to achieve rates that are equal to or greater than 39% [2]. The OL is clinically defined as a “white plaque with a questionable risk of malignancy once all other similar clinical lesions that carry no risk have been excluded” [2, 8]. This lesion may present microscopic aspects, which can vary from hyperkeratosis, with or without various levels of dysplasia (low or high), to carcinoma in situ [2, 4, 8]. Therefore, a biopsy is considered to be strictly necessary in order to establish a definitive diagnosis.
Currently, the diagnoses based on these clinical and histopathological criteria represent a problem given that they are poorly reproducible and have high inter-observer variability, therefore impeding a real standardisation in the early detection and prevention of the development of OSCC [2, 9, 10, 11]. As a result, one of the most important challenges in the diagnosis of OL is to predict which of the lesions will progress to OSCC.
Saliva is a complex biological matrix, the composition of which can vary depending on multiple factors [12, 13, 14]. The idea of using saliva as a method for the diagnosis and control of different diseases (periodontitis, cancer, recurrent aphthous stomatitis, OPMD, etc.) [13, 14] has proved to be of great interest, not only due to the varying molecular content, the low invasiveness of the procedure, and the fact that it is easy to collect, but also given the low cost that is involved [13]. The advances that have been made into the study of saliva have meant that it has been possible to establish its usefulness as a source of biomarkers that is comparable to blood, cerebrospinal fluid, pleural fluid and urine [13, 14, 15, 16], therefore allowing for the early diagnosis and screening of different diseases through “liquid biopsies” [14, 15, 17].
Despite innovations in the field of protein identification that involve several highly sensitive procedures (such as mass spectrometry techniques) [15, 16, 18, 19, 20], the ELISA method is still one of the most widely used, given its wide dissemination, reproducibility and low cost in comparison with the new omics techniques. [21, 22, 23, 24]. As a consequence, the search for protein markers in saliva offers further possibilities for understanding the structural interrelationships of OPMD and how these might determine progression to oral carcinoma.
The main goal was to perform a systematic review and meta-analysis with the markers that met the established requirements. The secondary goals were 1) To obtain information on the protein-based salivary markers described in the literature for OPMD, 2) To study the expression differences compared to healthy controls and malignant OSCC lesions.
Methods
A systematic review of the literature was conducted by a multidisciplinary team between the 1
PICO question
The study was organised based on the following PICO question: Patient, index test, comparison, outcome. P: patients diagnosed with an OPMD or OL. I: protein biomarker in saliva. C: patients with OSCC and the control group of healthy individuals (HCG). O: expression differences between OPMD and OSCC-HCG. Based on this the following scientific question was developed: Are there any protein-based markers in saliva that make it possible to quantitatively obtain diagnostic differences between healthy patients, OPMD and OSCC?
Criteria for considering studies for this review
Types of accuracy study
Studies of protein-based molecular markers (index tests) in saliva that revealed quantitative expression results related to clinically and histopathologically diagnosed diseases (reference standard). Therefore, the following were included 1) Human studies and controls, in the English language, which considered the OL or the OPMD and that included cases of OL as the reference standard; 2) Studies that included protein and saliva-based markers exclusively; 3) Studies that used ELISA and/or equivalent methods of analysis with quantitative expression determination (Bromocresol green method, Immunoturbidometry and LUMINEX). The exclusion criteria included: 1) Patients with explicit diagnosis, systemic disease, or a disorder that interfered with the conditions of interest; 2) Studies that only evaluated blood or tissue markers; 3) Studies in which the biomarker was not validated and was only discovered in label-free analysis; 4) Studies that only tried to validate a technique for diagnosis; 5) Betel quid chewing population as the only risk factor; 6) Studies that did not define the biomarker; 7) Studies that investigated changes in cytokines in saliva pre/post-treatment and that did not distinguish a potential biomarker; 8) Studies that did not define or include groups with the conditions of interest; 9) Studies that only included the OSCC or OSCC/HCG group. 10) Studies that quantified protein expression in units of measurement other than weight/capacity (i.e. mg/dl, ng/dl or pg./dl), such as Sequence coverage, mass/load ratio (m/z), Ion score, Fold-change o Mascot score. 11) Studies in which the marker was expressed under a minimum breakpoint.
Participants
The participants included in the study were those with clear and validated diagnoses, both clinically and histopathologically. Studies on experimental animals or in vitro models were excluded.
Control and target conditions
The patients included had been diagnosed according to the WHO Classification for Head and Neck Tumours. OPMD and OL were considered as the target condition and OSCC and HCG were considered as the healthy control group, both positive and negative according to the compared subgroups. The HCG group included patients without systemic diseases and who were not treated with immunosuppressive medication.
Reference standard
The reference standard for the diagnosis of OPMD, OL and OSCC was defined based on clinical and histopathological parameters (dysplasia and malignancy). Studies that did not detail any reference standard for the diagnosis of the described pathologies were not considered eligible for inclusion in this review. Given the great diversity of studies, clear inclusive diagnostic criteria were established. For the OSCC group: clinically diagnosed patients who had been verified through a histopathological study, regardless of the degree of differentiation, and excluding cases of warty carcinomas were included. For the OPMD group: cases of oral lichen planus, erythroplakia, oral submucous fibrosis and OL were included. We considered as OPMD any studies that did not mention a clear clinical diagnosis, but that did mention a non-malignant lesion with some degree of dysplasia [28, 29]. When only OL was found within the OPMD group, this had to be: white, non-detachable, homogeneous and non-homogeneous (mottled or warty) lesions with hyperkeratosis, acanthosis and/or histological squamous hyperplasia, with any degree of dysplasia (except carcinoma in situ, which was excluded).
Index test(s)
Any single protein-based biomarker detected in saliva, which was analysed from a quantitative point of view in order to differentiate between controls and targets. Studies analysing multiple biomarker panels were excluded because of the impossibility of analysis.
Search methods for the identification and selection of studies
Information sources and search strategy
Electronic searches in MEDLINE were performed using PubMed, EMBASE, OVID, Web of Science, Scopus, Cochrane Library, ClinicalTrials and the WHO regional bibliographic databases (AIM, IMEMR, IMSEAR, WPRIM) and the Conference Citation Index databases. Following the recommendations established by the Cochrane Systematic Reviews of Diagnostic Test Accuracy Group [30], the search strategy used to identify precision studies included three sets of terms related to the target condition (OPMD or OL), the index tests (protein-based biomarkers), and the type of oral sample (saliva) (Supplementary 2).
Data collection and analysis
The search was conducted by two observers (MPS and EA). Any ongoing discrepancies were resolved through discussion with two different reviewers (XMM and CMCP). The reasons for excluding studies were detailed. During the first phase, the title and abstract of the retrieved articles were read, and any studies that complied with the inclusion criteria, as well as those that did not provide sufficient data for a clear decision to be made were judged in full text. Subsequently, the entire content of all of the studies that were considered eligible was reviewed in their entirety in a second-round, and a final decision was made as to whether they were to be included or not.
Assessment of methodological quality
Two reviewers (MPS and EA) assessed the quality of the included studies separately using the revised Quality Assessment of Diagnostic Studies tool (QUADAS-2) [31]. This tool consists of four domains: 1) patient selection, 2) index test, 3) reference test, 4) flow and time. In addition, each domain was evaluated in terms of its risk of bias, and the first three domains were also evaluated in terms of their applicability.
Qualitative analysis
The unit of analysis was the expression of each protein biomarker in saliva, supplemented by the author and the year of publication. In most of the studies, the quantitative expression or level of a quantifiable protein was reported. Accordingly, these results were interpreted according to the presence or absence of disease (OSCC, OPMD or OL) based on a numerical measurement that was classified using a specific threshold (predetermined or not). The expression estimates were shown as integer values and their standard deviation for each biomarker and subgroup. The weighted mean difference (WM) between the case group and the control group was obtained for each model, giving the overall expression (OE) and the 95% confidence interval (CI95%). We distributed the meta-analysis in two subgroups, the first OSCC (standard reference)/ OPMD and the second OPMD (standard reference)/HCG, including the isolated OL reports within OPMD group.
Quantitative analysis
A meta-analysis was performed when the number of data for a biomarker was at least two papers and when the units of measurement of the marker were the same. The analysis strategy was to include all of the expression studies, regardless of their threshold value. The statistical analysis of the results was performed using the OpenMeta tool (Analyst).
Functional networks, and pathway mapping
Analyses of biological pathways within the differential protein markers were performed using the PANTHER enrichment analysis tool [32], only for interactions with high confidence (0.7). The Reactome Biological Pathway enrichment analysis of proteins of interest was performed using the STRING tool [33]. In both cases, we used proteins that presented statistically significant differences in our study, and in some cases, we performed node enrichment to be able to obtain more protein interactions. Finally, we used a systematic bioinformatics analysis including Database for Annotation, Visualisation and Integrated Discovery (DAVID) and gene ontology enrichment [34].
Assessment of heterogeneity
Firstly, the threshold effect of expression was evaluated graphically by looking at the forest plot and secondly, the threshold effect was evaluated statistically using the
Results
Study selection
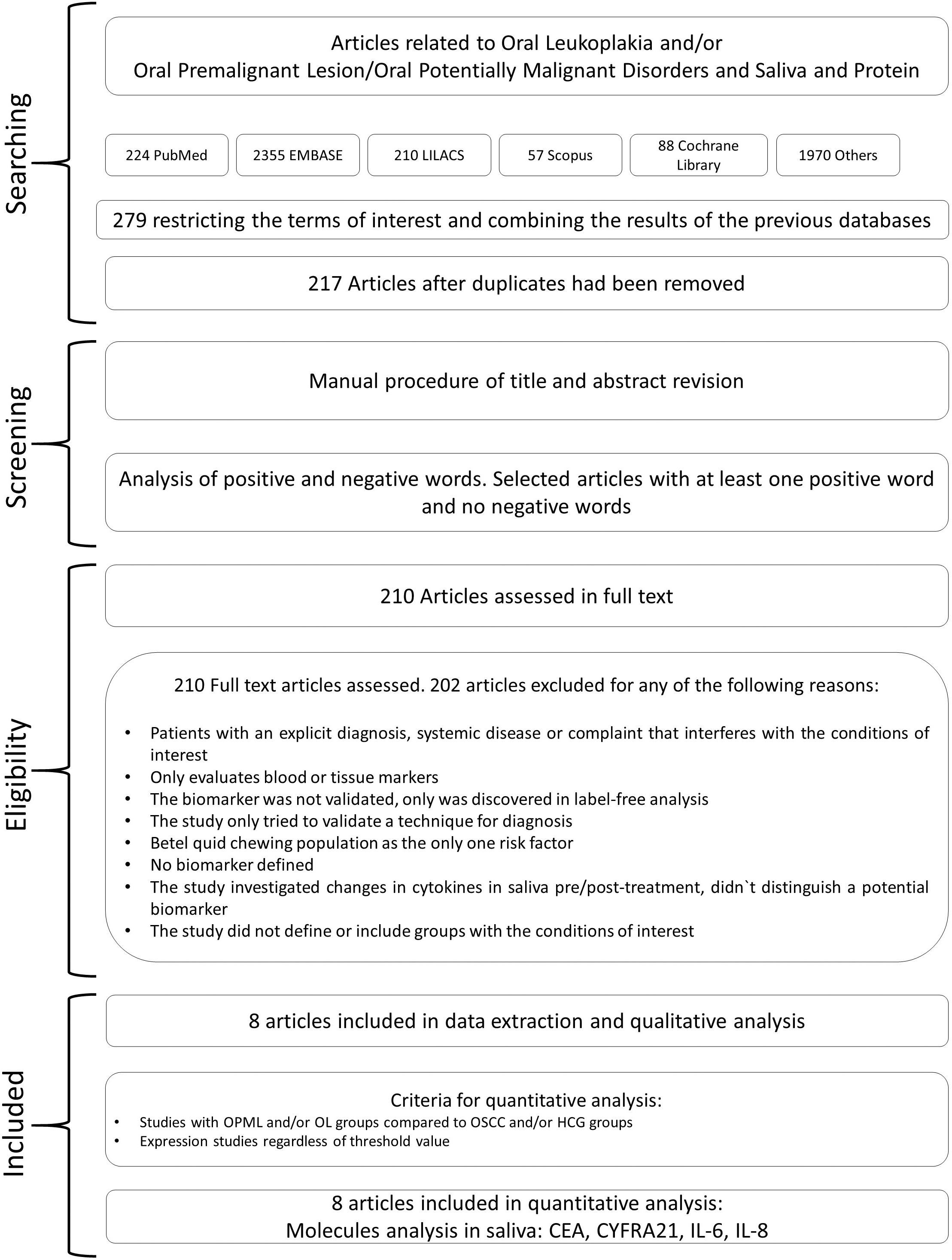
The overall combined result of the references obtained from the aforementioned databases was 279 citations. After removing the duplicates, we obtained 217 unique citations. The biomarkers described in the literature for OPMD included: carcinoembryonic antigen (CEA), C-reactive protein (CRP), soluble fragment of cytokeratin 19 (CYFRA21), human epidermal growth factor receptor 2 (Her-2/neu; erbB-2), interleukin-1
Characteristics of protein-based studies in saliva
Table 1 summarises the characteristics of the eight selected studies, disaggregated by biomarkers. The
Descriptive data from the studies included in the meta-analysis
Descriptive data from the studies included in the meta-analysis
Note: CEA: carcinoembryonic antigen, CYFRA 21: soluble fragment of cytokeratin 19, IL-6: interleukin-6, IL-8: interleukin-8.
Flow chart describing the management of data from the search, data processing and selection of articles for meta-analysis.

Overall expression values
Note: Overall expression values (OE) with a 95% confidence interval (CI95%) and the heterogeneity (
Forest plot with the weighted mean differences, overall expression (OE), 95%.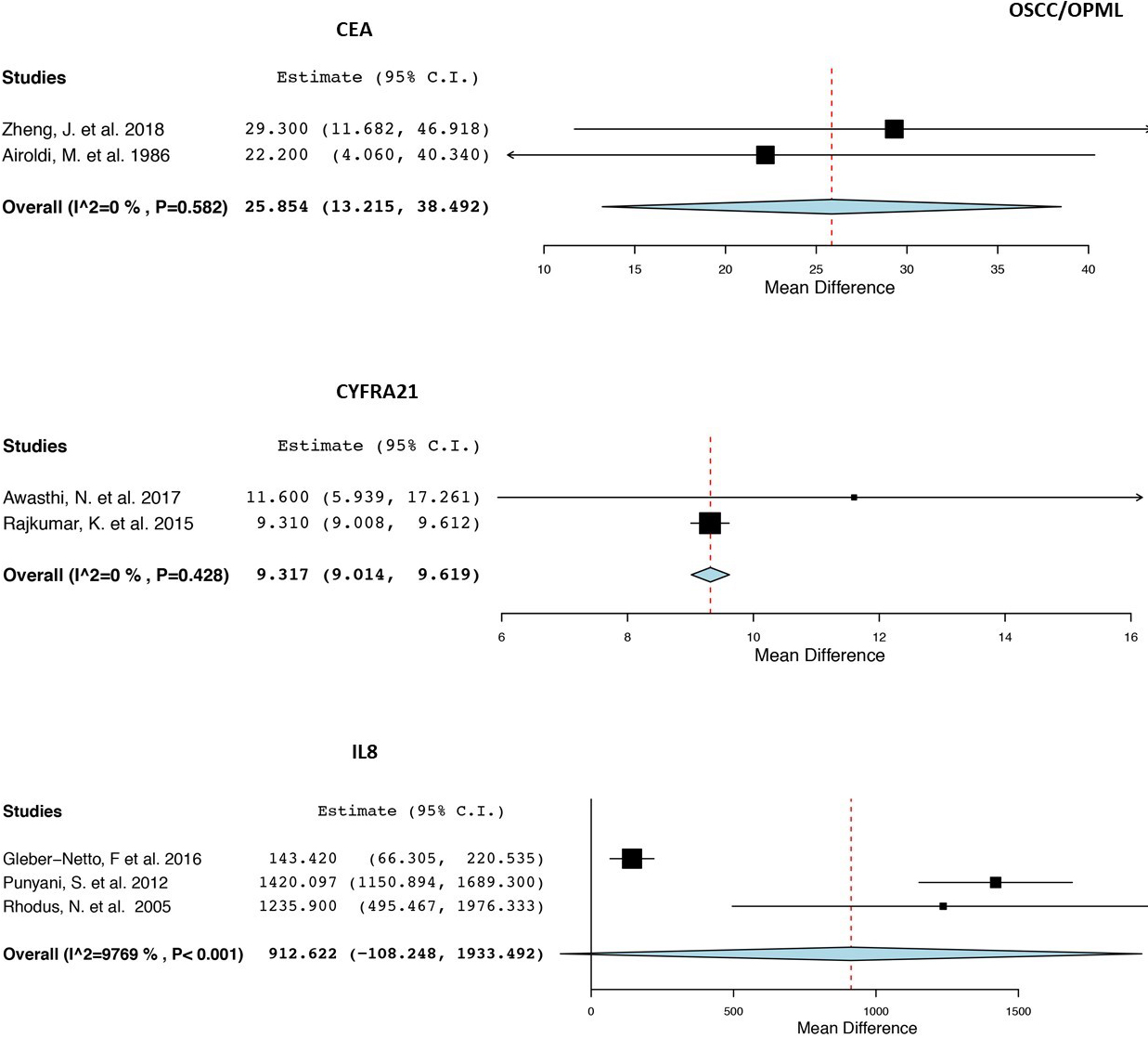
Forest plot with the weighted mean differences, overall expression (OE), 95% confidence interval and heterogeneity for each protein biomarker in saliva for the OPMD/HCG subgroup analysis A) CEA. B) CYFRA21. C) IL8. D) IL6.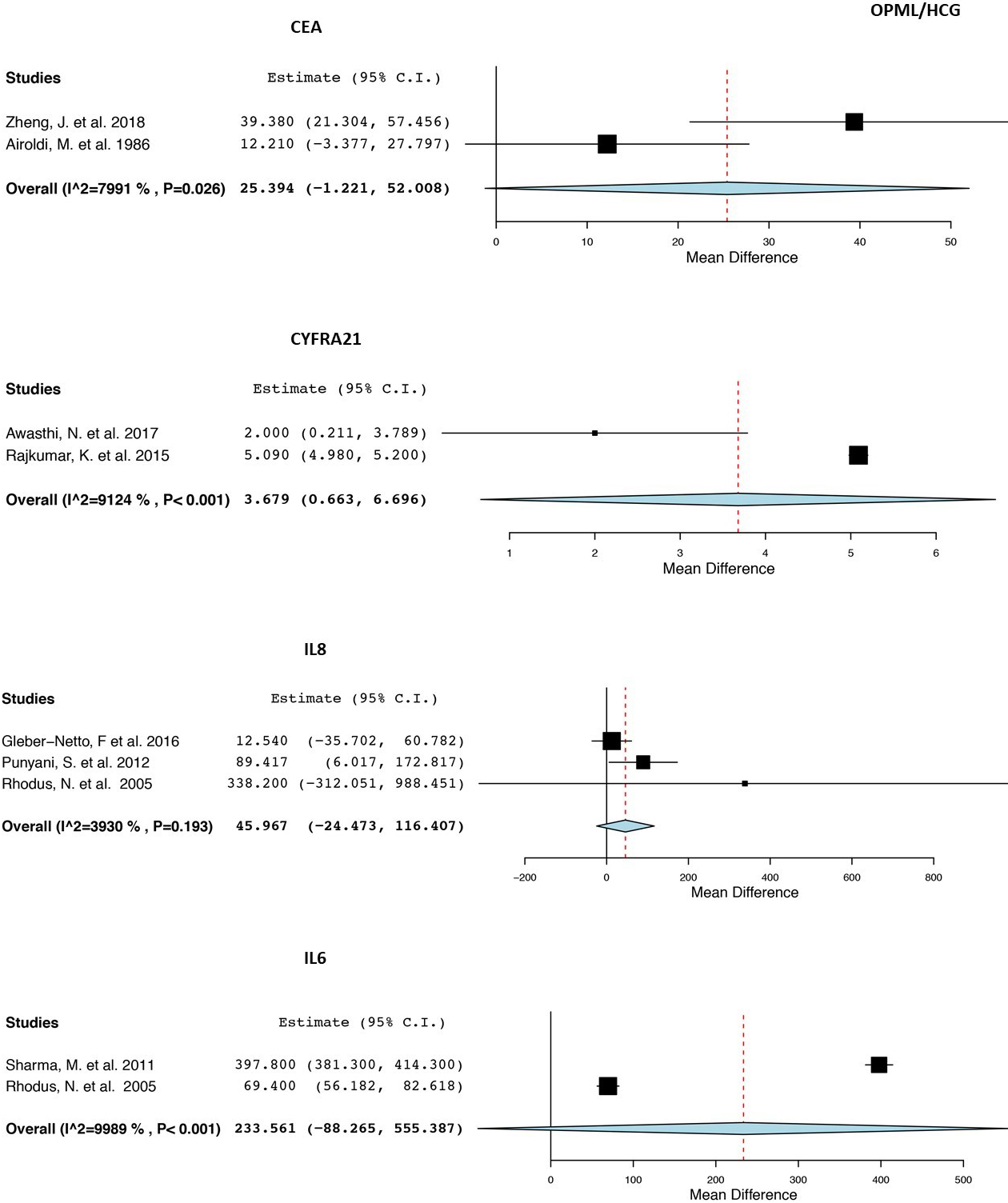
eight selected studies were conducted in five different countries: China [36], Italy [43], India [37, 39, 40, 41], USA [42] and Taiwan [38]. A total of 986 patients were included who were distributed in the following groups: HCG (
With regards to the methodological quality of the studies (Supplementary 3), after applying the QUADAS-2 scale regarding the risk of bias, it was determined that this was very high for patient selection (100%) and very low (0%) for Flow and Timing and Reference Standard. With respect to concerns regarding the applicability, the methodological risk was generally very low, especially for the patient selection and index test fields, although the study by Zheng et al. (1 study of 8, 12.5%) had an uncertain evaluation of the reference standard, given that it did not define the number of cases for each type of diagnosis included in the OPMD group.
Synthesis of the qualitative and quantitative analyses of four salivary biomarkers
In terms of the different markers and study subgroups, Table 3 shows all the values of OE and
Once these proteins had been obtained from the eight selected references, we performed interaction studies to determine the possible relationship between them. To do this the STRING program was used, and as can be seen in Fig. 3, there is a direct relationship between IL-6 and IL-8 (CXCL-8), forming a small cluster of pro-inflammatory cytokines with two proteins, the CXC chemokine receptor type 2 (CXCR2) and the IL-6 receptor (IL6R), which are potent chemotherapeutics targets and neutrophil activators (Supplementary 4).
Given the relationship between the cytokines, which showed a well-defined cluster, several studies were performed to determine the position of these proteins in the immune system’s response to the inflammatory process. Using the DAVID program, we were able to observe how these proteins are found in the TNF signalling pathway activated by macrophages (Supplementary 5). The study, which used databases such as PHANTER provided us with a global view of the biological processes in which the selected proteins are involved. According to Supplementary 6, proteins are involved in signalling processes, the immune system, regulation, localisation, locomotion, etc. A new protein-protein interaction analysis was performed, in which it was observed that the selected biomarkers presented a theorical direct molecular relationships between IL-8 (or CXCL-8), IL-6 and CYFRA21 (KRT19). The data gathered from STRING indicated that KRT19 is involved in the organisation of myofibrils alongside KRT8, while CEA plays a very important role in cell adhesion and intracellular signalling processes (Fig. 3, Supplementarys 7 and 8). Only CYFRA21 can be considered as a good marker for the OPMD/HCG subgroup. By asking the program to indicate more possible nodes/interactions related to this protein, we determined a strong cluster composed of cytoskeleton proteins and Mesothelin, a membrane-binding protein that plays a very important role in the cell adhesion process (Supplementary 7 and Supplementary 9).
Discussion
The hypothesis of this study proposed the idea that there are protein-based biomarkers in saliva whose quantitative expression in OPMD is different to that of healthy control groups and malignant lesions, therefore making it possible for reference points of expression to be established. Although we found studies in which 11 different biomarkers had been analysed, it was only possible to perform a meta-analysis on eight studies with four biomarkers.
Protein interaction network of the four protein markers selected using the STRING databases.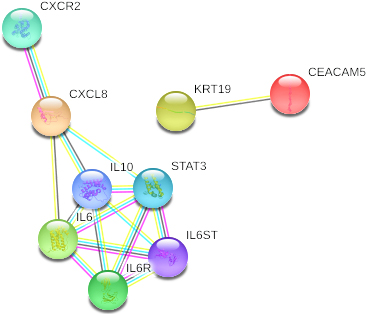
CEA is a transmembrane glycoprotein that has multiple functions. In OSCC of the tongue, its overexpression correlates with neutrophil infiltration and both are associated with poor prognosis [44]. In our meta-analysis, CEA presented differences in expression between OSCC and OPMD (
Historically, CYFRA21 has been researched as a biomarker for non-small cell lung carcinoma and it was subsequently evaluated in other malignant [45] and potentially malignant disorders [46]. It may also be useful as a biomarker in the head and neck region [47]. The OE for CYFRA21 was highly significant in OSCC compared to OPMD (
In studies that were performed to determine the interaction between CEA and CYFRA21, the results demonstrated that both present a strong association (Fig. 3) and that they are directly related to a whole cluster of keratins that are involved in the cellular cytoskeleton, one of the key factors in oral carcinogenesis [48]. All this was in line with studies in which it has been determined that both proteins can be used as diagnostic markers in other epithelial tumours such as lung cancer [49] and head and neck cancer [50], therefore concurring that these two proteins are differentially expressed in healthy and potentially malignant tissues.
The IL-6 salivary mRNA was previously identified as a promising biomarker for OSCC [51], however, studies into the role of protein expression in saliva or OPMD are limited. There was no significant difference in the IL-6 cytokine salivary OE between the OPMD group and HCG. The anticipated means of expression for the studies included in the meta-analysis were considerably different, the highest being 397.8 (CI95%: 381.3–414.3) in the study by Sharma et al. [41], compared to 69.4 (CI95%: 56.1–82.6) in the study by Rhodus et al. [42], which resulted in a value of extreme heterogeneity. In the study by Rhodus et al, no clinical diagnoses were indicated in the OPMD group, only different histological degrees of dysplasia [42]. The concentration of IL-6 was significantly higher in OSCC and OPMD when compared to HCG (
Changes in the expression of pro-inflammatory cytokines may play a role in the malignant transformation in OSCC and are likely to be used as diagnostic markers in the serum of patients, due to their excessive production by tumour cells [14], and, likewise, they could also act as possible pharmacological targets. IL-8 plays an important role in cancer invasion, angiogenesis and metastasis [52]. In this meta-analysis, we found no significant difference for IL-8 between OSCC and OPMD, nor between OPMD and HCG. Likewise, we did not find any relationship between these pro-inflammatory cytokines and the previously described markers (CEA and CYFRA21), however, as can be seen in Fig. 3, the pro-inflammatory cytokines would form a strong signalling cluster by themselves, through the functional analysis that has been performed with the DAVID program, corresponding to the immune response and inflammation signalling pathway (Supplementary 4). The studies included in the meta-analysis showed a very high value of heterogeneity in all of the subgroups (Figs 1C and 3C). The study by Gleber-Netto et al. [38] covered an almost exclusively male population in Taiwan, with a percentage of smokers that exceeded 80% in all groups. The corresponding clinical diagnoses were not reported in the OPMD group, only those lesions with different degrees of dysplasia were included. On the other hand, the study conducted by Punyani et al. [40] in India reported that the OPMD group was comprised of 13 oral submucous fibrosis and 12 OL, and that 5 of these had mild dysplasia. The identification of IL-8 was significantly higher in OSCC compared to OPMD (
The main constraints of this study included the lack of clinical-pathological information on the lesions that are mentioned as dysplastic in the OPMD group, and likewise, the degree of dysplasia has not been characterised. Additionally, there were considerable discrepancies between the number of analysed cases of OL and OPMD, compared with the cases of OSCC. Very few studies have analysed pure OL groups, independent of other OPMD, which therefore makes it difficult for specific conclusions to be drawn. Another aspect that must be taken into consideration is that the gender and age of the individuals in the meta-analysis was quite heterogeneous, without there being a strictly matched selection in terms of gender or similar age intervals between all of their study groups. Moreover, not all of the studies reported the presence of risk factors such as smoking (a key factor in oral carcinogenesis) and others included habits that are typical of the geographical region under study, such as chewing betel nut, therefore meaning that extrapolation to a global level was not possible. In terms of the strengths of this study, it is important to highlight the clear and strict inclusion criteria that made it possible to limit analysis biases. Furthermore, even though there were different techniques such as proteomic techniques that allowed for massive quantitative analyses of the literature, this systematic review only included studies in which the quantitative values were representative, reproducible and comparable.
This systematic review and meta-analysis provides a summary of the quantitative expression of biomarkers based on salivary proteins. Based on the results, we can conclude that it is quantitatively possible to discriminate the OPMD compared to malignant lesions and healthy controls. The CEA and CYFRA21 markers proved to be very useful when differentiating OSCC from OPMD, however, the usefulness of exclusively differentiating the group of leukoplakia from OSCC does not seem totally clear. Besides, the CYFRA21 marker proved to be the only protein that was capable of distinguishing between OPMD and healthy controls. The definition of these protein-based salivary markers as a molecular signature, which identifies potentially malignant lesions, would allow for new, less invasive protocols to be established for early diagnosis and screening, as well as making it possible to determine therapeutic targets that would improve the prognosis of patients with potentially malignant lesions and those at risk.
Author contributions
Esteban Arroyo: Conception, Interpretation and analysis of data, Preparation of the manuscript, Revision for important intellectual content.
Sergio Piñeiro Donís: Conception, Interpretation and analysis of data, Preparation of the manuscript, Revision for important intellectual content.
Cintia Chamorro: Interpretation and analysis of data, Preparation of the manuscript.
Monica Oliveira Alves: Interpretation and analysis of data, Preparation of the manuscript.
Xabier Marichalar Endia: Interpretation and analysis of data, Preparation of the manuscript.
Darcy Fernandes: Interpretation and analysis of data.
Alejandro Lorenzo Pouso: Interpretation and analysis of data, Preparation of the manuscript, Supervision.
Andreia Bufalino: Conception, Supervision.
Susana Bravo López: Conception, Preparation of the manuscript, Supervision.
Pilar Gandara Vila: Conception, Supervision.
Mario Pérez Sayáns: Conception, Interpretation and analysis of data, Preparation of the manuscript, Revision for important intellectual content, Supervision.
Sources of support in the form of grants
Esteban Arroyo received funding for this research from Programa Estudantes-Convênio de Pós-Graduação – PEC-PG, CAPES/CNPq, Brazil (88881.154490/2017-01); Andreia Bufalino received funding for this research from São Paulo Research Foundation (FAPESP) (grant #2017/01438-0) and the National Council for Scientific and Technological Development CNPq (423945/2016-5); Darcy Fernandes received funding for this research from São Paulo Research Foundation (FAPESP) (grant #2017/01798-7). This study was financed in part by the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior – Brasil (CAPES).
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-203043.
sj-pdf-1-cbm-10.3233_CBM-203043.pdf - Supplemental material
Supplemental material, sj-pdf-1-cbm-10.3233_CBM-203043.pdf
