Abstract
BACKGROUND:
Oral squamous cell carcinoma (OSCC) usually originates from oral potentially malignant disorders (OPMD), such as oral leukoplakia (OLK) and oral lichen planus (OLP). Identifying biomarkers for the early diagnosis and evaluation of malignant transformation in OPMD could improve the survival rate of OSCC patients.
OBJECTIVE:
The present study aimed to screen for potential salivary biomarkers for evaluating the malignant transformation of OPMD.
METHODS:
Salivary proteases from OLK and OSCC patients or healthy donors and proteases in cultural medium from DOK and Cal-27 cells were detected with a human protease array kit. The concentrations of the salivary Kallikrein 5 (KLK5) and urokinase-type plasminogen activator (uPA) proteases were measured by ELISA. Receiver operating characteristics (ROC) to determine the potential value of these proteases in clinical diagnosis were calculated using SPSS software. Immunohistochemistry was used to detect the KLK5 and uPA expression in the oral organizations.
RESULTS:
The salivary protease spectrum was different among patients with OLK and OSCC and healthy donors. KLK5 and uPA levels in saliva tended to increase as the disease progressed (healthy
CONCLUSIONS:
Salivary KLK5 and uPA are potential biomarkers for evaluating OLK and OLP malignant transformation and early diagnosis of OSCC.
Keywords
Introduction
Oral squamous cell carcinoma (OSCC) is the most predominant type of oral cancer [1]. Although comprehensive strategies for its recognition and management have advanced in the past decade, the 5-year survival rate for OSCC has not significantly improved [2, 3]. This lack of improvement might be partly due to the lack of reliable diagnostic biomarkers or early detection methods, which delays the diagnosis of OSCC.
The histopathological development of OSCC occurs via oral potentially malignant disorders (OPMD) that go through stages, including epithelial hyperplasia and dysplasia, followed by OSCC stages consisting of early invasive and lymph node metastatic stages [4]. OPMD, which include oral leukoplakia (OLK), oral erythroplakia (OE), proliferative verrucous leukoplakia (PVL), oral submucous fibrosis (OSMF), oral lichen planus (OLP), and actinic cheilitis (AC) [5, 6], are morphologically altered tissues in which oral cancer is more likely to occur compared to the normal oral mucosa. As the most common OPMD, OLK is a pathological condition with white patches or plaques that could not be attributed to any clinically or histologically definitive lesion, while OLP is defined as a common chronic mucocutaneous disease with unknown etiology [7]. The mean malignant transformation rate of OLK for the total population amounts to 3.5% (405/11423), with a wide range of rates (0.13–34.0%) presented in a recent systematic review study [8]. OLP has a malignant transformation rate of 1.14% (95% CI
Saliva contains a variety of hormones, antibodies, microorganisms, proteins, enzymes, and cytokines. Its components reflect the physiological state of the body [14, 15]. With the development of salivaomics, an increased number of biomarkers related to various systemic and oral diseases, such as gastric cancer [16], type 2 diabetes [17], breast cancer [18], systemic lupus erythematosus [19], oral squamous cell carcinoma [20, 21, 22], periodontal disease [23], and caries [24, 25, 26], have been identified in saliva.
Proteases drive the life cycle of all proteins, ensuring the activation of newly synthesized proteins and degradation of unneeded proteins [27]. The abnormal release and activation of proteases can generate pathological changes in cells, tissues, and organs, especially in the formation and metastasis cancers [28]. Many kinds of proteases can be detected in human saliva, and changes in the salivary protease spectrum can be used as a potential biomarker for oral diseases [21, 29].
In this study, we analyzed the protease spectrum of saliva from healthy donors and patients with OLK and OSCC and in the supernatants from dysplastic oral keratinocytes (DOK, human mucosa precancerous lesion cell line) and Cal-27 (oral squamous cell carcinoma cell line) cells using human protease arrays. Based on the results of the protease array analysis, we measured the concentrations of the kallikrein 5 (KLK5) and urokinase-type plasminogen activator (uPA) proteases in the saliva of healthy donors and patients with OPMD (OLK and OLP) or OSCC by ELISA. The expression levels of these proteases were also evaluated in tissues by immunohistochemistry (IHC). The ELISA data were used to calculate the sensitivity and specificity for diagnosing OLK, OLP, and OSCC using receiver operating characteristic (ROC) curves. Our results demonstrated that salivary KLK5 and uPA are potential biomarkers for evaluating OLK and OLP malignant transformation and the early diagnosis of OSCC.
Materials and methods
Ethics statement
All volunteers were informed that their saliva would be collected to assay the concentration of proteases. All data, information, and samples were used for our research study only. Written informed consent for collecting saliva and tissues and identifying individual medical information was obtained from each subject. The Committee for Ethics Approval of the West China School of Stomatology, Sichuan University and State Key Laboratory of Oral Diseases approved this study (IRB Reference Number: WCHS-IRBD-2015-147).
Saliva collection
Stimulated saliva was collected with salivette tubes (Sarstedt AG & Co., Numbrecht, Germany). Before sample collection, participants were prohibited from eating and drinking for 2 h. They were then asked to gargle three times with 10 mL of clear water. After 30 min, they put the cotton from the salivette tube into their mouths without chewing for 10 min, and then spit the cotton back into the tubes. The tubes were kept on ice following collection. The samples were centrifuged at 10,000
A total of 120 saliva samples were collected from patients with OSCC (
Cell culture and supernatant collection
Dysplastic oral keratinocytes (DOK) and Cal-27 cells were cultured as previously described [30]. To analyze secreted proteases, the cells were seeded in six-well plates at a density of 2
Protease array and image analysis
Twelve saliva samples (
RL: Relative Level of Proteases; IDs: Integrated Density of samples; IDn: Integrated Density of negative control; IDrs: Integrated Density of reference spots.
Quantitative assessment of salivary proteases by ELISA
ELISA analysis was performed on 120 saliva samples from OSCC (
The ELISA data were used to determine the optimal cutoff values, specificity, sensitivity, and area under the curve (AUC) with 95% confidence intervals (CI) with the ROC curve using SPSS 19.0 software.
Immunohistochemical analysis
A panel of 56 formalin-fixed, paraffin-embedded human OLK, OLP, and OSCC specimens were obtained from the West China Stomatological Hospital. For IHC analysis, all slides were deparaffinized twice in xylene (15 min each) and rehydrated through a series of graded alcohol solutions. Antigen retrieval was performed by heating the slides with boiled water for 20 min in 10 mmol/L sodium citrate buffer (pH 6.0). After cooling, endogenous peroxidase activity was blocked with 3% hydrogen peroxide for 15 min and normal serum for 30 min at room temperature. Subsequently, the slides were incubated with rabbit anti-human uPA (cat. no. ET1703-26; 1:100; HUABIO) or KLK5 (cat.no. 10514-2-AP; 1:50; Proteintech) antibody at 4
Image analysis was performed using ImageJ 1.48v software with the IHC Profiler plugin. For the ImageJ settings, the mode was set for cytoplasmic stained images, and the vector was H DAB. The staining intensity was evaluated according to the following ratings: 0, no staining; 1, weak staining; 2, medium staining; 3, strong staining. The H-score was calculated as follows: H-score
Statistical analysis
SPSS 19.0 software (Chicago, IL, USA) was used for data analysis. The statistical power for comparing KLK5 and uPA between groups was calculated using one-way analysis of variance F-tests in the means with G*Power (3.1.9.2) software. The 1-
Coordinates of proteases located on the human protease array membrane
Coordinates of proteases located on the human protease array membrane
RS: reference spot, NC: negative control.
Saliva from OSCC and OLK patients contain more proteases than from healthy donors
We analyzed 12 saliva samples from healthy individuals and patients with OLK and OSCC using human protease array kits that detect 35 human proteases (Fig. 1A and Table 1). The salivary protease spectrum was quite different between the three groups. Saliva from healthy donors contained 17 proteases (Fig. 1B). In contrast, the saliva from OLK (Fig. 1C) and OSCC (Fig. 1D) patients contained 26 and 30 proteases, respectively. The relative levels of each protease calculated from scanned images are presented in Fig. 1G. Most of the salivary proteases (e.g., cathepsin V, KLK5, KLK7, KLK13, MMP-1, MMP-2, MMP-3, MMP-13, proteinase3, and uPA) increased gradually as the disease progressed from OLK to OSCC. In the saliva from OSCC patients, the relative KLK5 and uPA expression levels were 11.57- and 40.07-fold higher than those observed in healthy volunteers, respectively. Moreover, these expression levels were 2.88- and 11.86-fold higher than the levels in OLK patients, respectively. Furthermore, the relative KLK5 and uPA expression levels in the saliva of OLK patients were 4.02- and 3.38-fold higher than found in healthy volunteers. These results indicated that the identities and concentrations of salivary proteases, especially KLK5 and uPA, were associated with OSCC disease progression.
Protease spectrum in saliva and cell culture supernatants. Saliva proteases were detected using human protease array kits. (A) Each dot on the membrane represents a single protease. (B) The salivary protease spectrum of healthy donors. (C) The salivary protease spectrum of OLK patients. (D) The salivary protease spectrum of OSCC patients. (E) The protease spectrum of DOK cell culture supernatant. (F) The protease spectrum of Cal-27 cell culture supernatant. (G) Quantitative analysis of saliva proteases. (H) Semi-quantitative analysis of supernatant proteases. Representative pictures for each type of sample are shown. 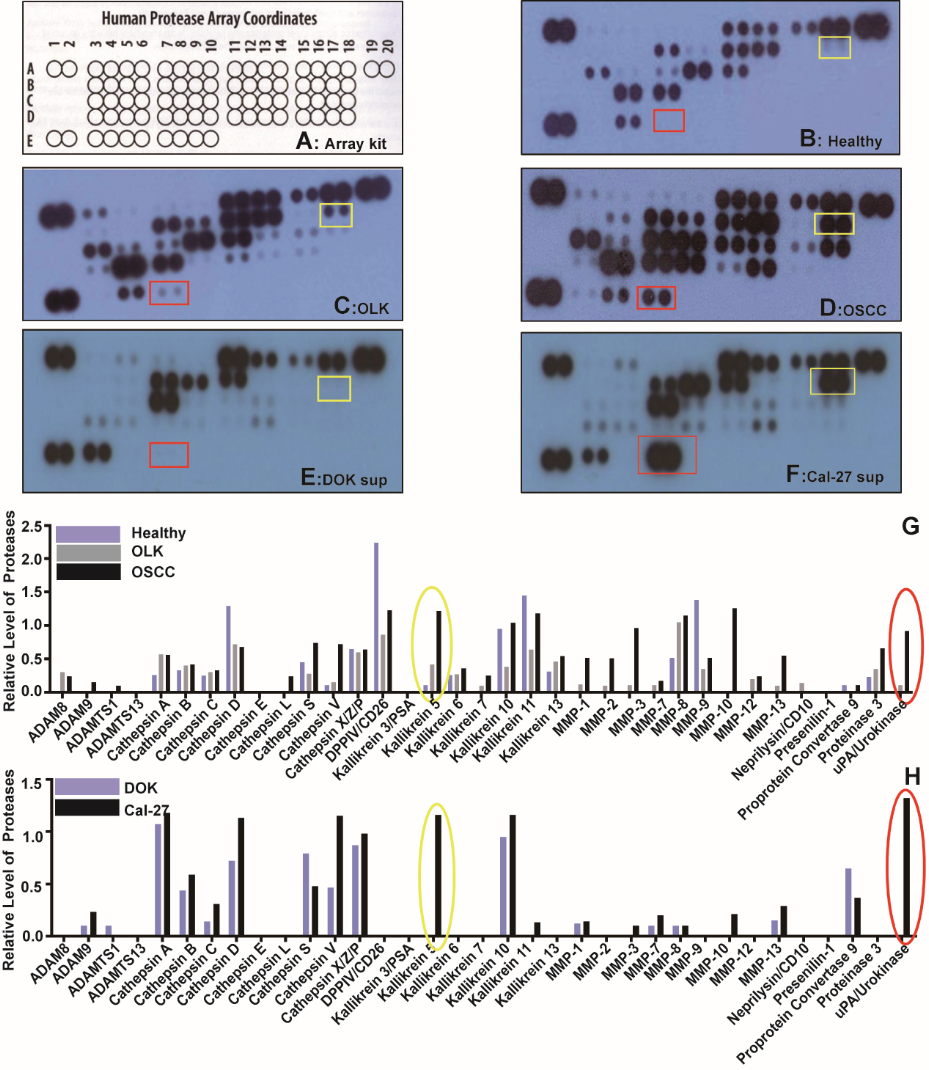
Concentration of salivary KLK5 and uPA detected by ELISA
OLK-T: OSCC which had undergone malignant transformation originating from OLK. OLP-T: OSCC which had undergone malignant transformation originating from OLP.
Cutoff point, AUC, 95% CI, specificity and sensitivity of KLK5 and uPA for diagnosis of diseases
a: Standard error, b: Asymptotic significance.
The culture medium from DOK and Cal-27 cells were also analyzed with the human protease array kit. Cal-27 cells released 19 different proteases into the culture medium compared to 14 by DOK cells (Fig. 1E and F). Consistent with the saliva data, the relative expression levels of most proteases in the supernatant of Cal-27 cells (e.g., ADAM9, Cathepsin B, Cathepsin C, Cathepsin D, Cathepsin V, KLK5, KLK10, KLK11, MMP-3, MMP-7, MMP-10, MMP-13, and uPA) were higher than in the supernatant from DOK cells (Fig. 1H). In particular, the relative KLK5 and uPA expression levels in the Cal-27 medium were 12.86- and 499.41-fold higher than that in the DOK medium, respectively. These data suggested that KLK5 and uPA might be potential biomarkers to evaluate precancerous oral mucosa lesion cancerization.
Quantitative analysis of patient salivary KLK5 and uPA levels. KLK5 and uPA levels were measured in the saliva from healthy donors and OLK, OLK-T, OLP, OLP-T, and OSCC patients by ELISA. Data are presented as the mean 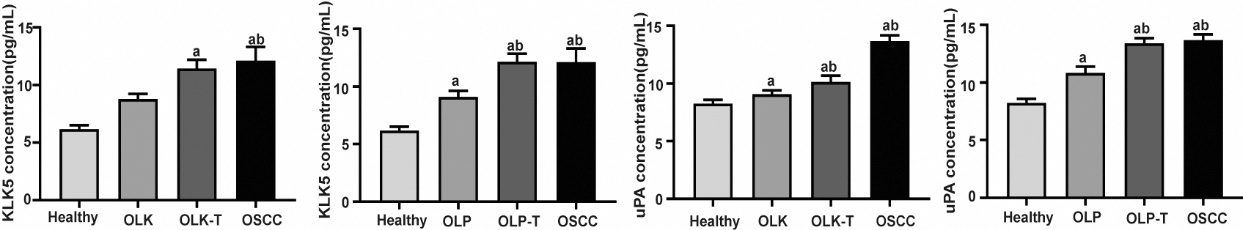
Based on the human protease array results from saliva and cell culture supernatants (Fig. 1), we evaluated the KLK5 and uPA concentrations in saliva samples from healthy individuals and patients with OLK, OLP, OLK-T, OLP-T, or OSCC. The levels of KLK5 and uPA gradually and significantly increased from healthy donors to patients with OLK or OLP to patients with OSCC (OLK-T, OLP-T, and OSCC) (Fig. 2 and Table 2). There were no significant differences between OLK-T, OLP-T and OSCC patients.
ROC curves showed the optimal KLK5 diagnostic cutoffs to be 5.97 pg/mL for OLK, 6.03 pg/mL for OLP, 8.36 pg/mL for OLK-T, 8.80 pg/mL for OLP-T, and 9.45 pg/mL for OSCC (Fig. 3A). The optimal uPA diagnostic cutoffs were 17.19 pg/mL for OLK, 17.26 pg/mL for OLP, 19.45 pg/mL for OLK-T, 22.29 pg/mL for OLP-T, and 20.96 pg/mL for OSCC (Fig. 3B). The corresponding AUC value, 95% CI, sensitivity, and specificity for KLK5 and uPA are presented in Table 3. Both KLK5 and uPA showed high sensitivity as a biomarker for the diagnosis of OLP, OLP-T, OLK, OLK-T, and OSCC, and their combination improved the diagnostic accuracy compared to either test alone, especially the specificity of diagnosis (Fig. 3C–G, Table 4).
ROC curves for salivary KLK5 and uPA alone and in combination for the diagnosis and differential diagnosis of OLK, OLK-T, OLP, OLP-T, and OSCC. (A–B) ROC curves for KLK5 and uPA for patients with OLK, OLK-T, OLP, OLP-T, and OSCC versus healthy donors. (C–G) ROC curves for KLK5 and uPA alone and in combination for the diagnosis of OLK, OLP, OLK-T, OLP-T, and OSCC. (H–K) ROC curves for KLK5 and uPA alone and in combination for patients with OLK or OLP versus OSCC.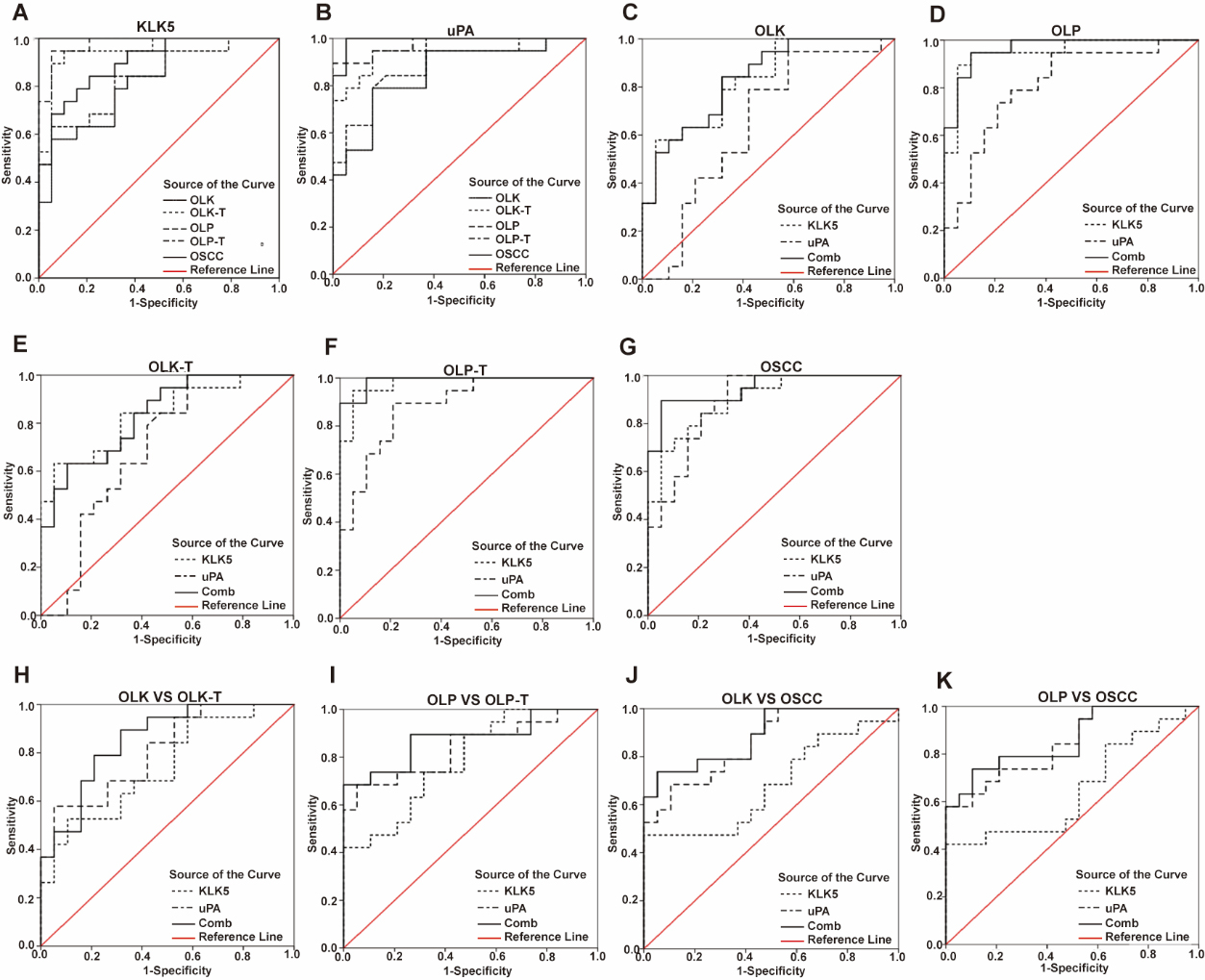
AUC, 95% CI, specificity and sensitivity of KLK5, uPA and their combination for diagnosis of diseases
a: Standard error, b: Asymptotic significance.
We found that raised salivary uPA concentrations could separately distinguish patients with OSCC from those with OLK or OLP, while salivary KLK5 alone could not. The individual tests had low AUC values (0.65–0.78) and low specificity (21.0–52.6%). When KLK5 and uPA were tested together, the differential diagnostic accuracy between OSCC and OLK/OLP improved, with AUC values of 0.85 (95% CI
Cutoff point, AUC, 95% CI, specificity and sensitivity of KLK5, uPA and their combination for discriminating OLK and OLP from OSCC
a: Standard error, b: Asymptotic significance.
Altogether, the ROC analysis illustrated that salivary KLK5 and uPA were potential biomarkers for the diagnosis of OLK, OLP, and OSCC, and their combination was better for distinguishing OSCC from OLK and OLP.
KLK5 and uPA expression levels were analyzed in 16 OLK, 5 OLP, and 35 OSCC specimens. As shown in Fig. 4A, both KLK5 and uPA were expressed at low levels in oral epithelial tissue from OLK and OLP patients. In contrast, OSCC tissues were more intensely stained. Semi-quantitative analysis showed that the KLK5 and uPA levels were significantly increased in OSCC compared to OLK and OLP (Fig. 4B–C;
KLK5 and uPA levels in OLK, OLP, and OSCC specimens. (A) KLK5 and uPA expression levels were analyzed by immunohistochemistry in OLK, OLP, and OSCC specimens; (B) H-score analysis of KLK5 in different oral tissues. (C) H-score analysis of uPA in different oral tissues. *** 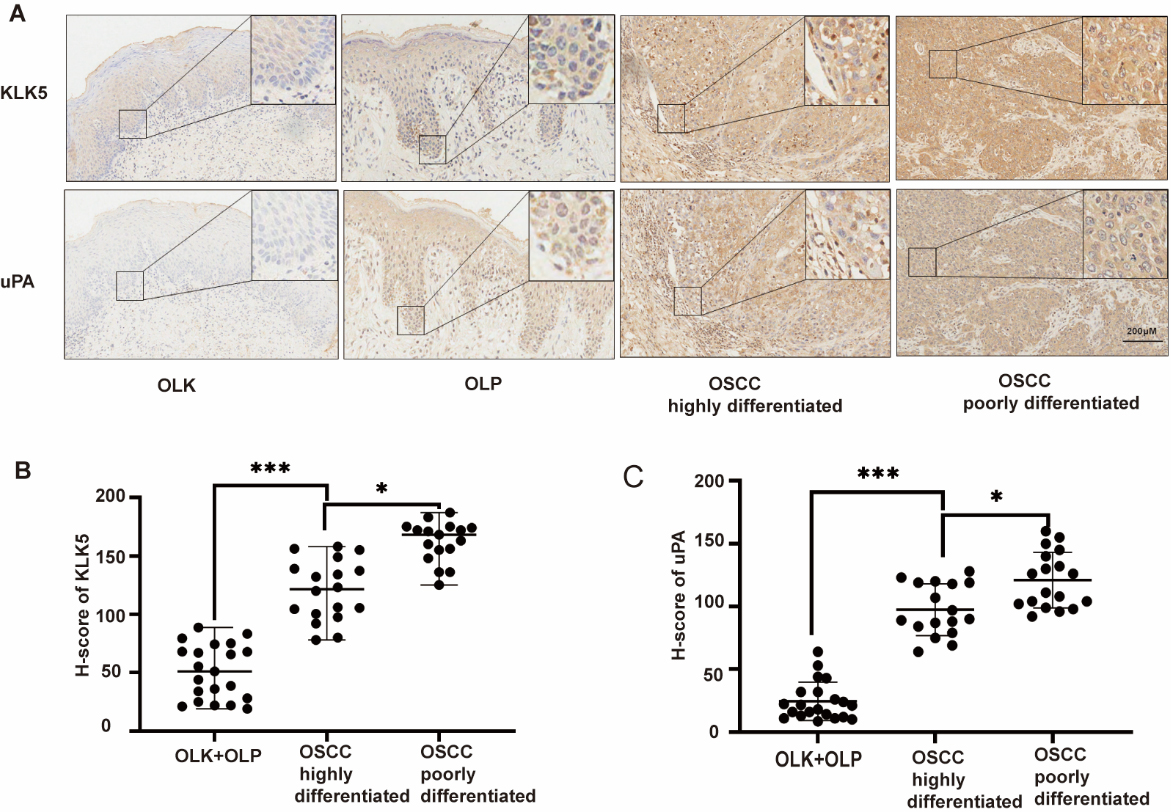
The identification of biomarkers for diverse tumor types in the blood, urine, cerebrospinal fluid, or other body tissues has attracted significant attention in the last decade. In theory, tumor biomarkers can be used to screen, diagnose, stage, and monitor cancers [31]. Several proteases have been extensively studied as potential biomarkers, including uPA/uPAR and matrix metalloproteinases (MMPs) [32]. The abnormal activation or secretion of proteases can give rise to pathological processes in cells, tissues, and organs. Many proteases play important roles in the invasion and metastasis of cancers because of their ability to degrade the extracellular matrix (ECM) barrier surrounding a tumor [33]. Saliva is an important body fluid that can reflect the body’s physiological state. An increasing number of disease-related salivary biomarkers have been discovered, and the analysis of the salivary protease spectrum has been reported as a perfect way to evaluate oral health status [34, 35].
OLK and OLP are the most common OPMD. Many methods exist to prevent and treat OLK, such as drugs, conventional scalpel excision, photodynamic therapy, cryosurgery, and laser ablation, while the main method for treating OLP is medication [4]. Despite the continuous advancements in treatment technology, OLK and OLP still have a high malignant transformation rate [8, 9]. Recurrence and malignant transformation of OLK and OLP are major clinical obstacles. Identification of biomarkers with high specificity and sensitivity for identifying malignant transformation of OLK and OLP is a major focus in the field. Such markers could allow for surgical intervention at an early stage.
OLK, OLP, and OSCC are usually located on the surface of oral mucous and contact saliva directly. As a result, the saliva of those patients will contain all kinds of molecules secreted and released by tumor cells or cells of precancerous lesions, suggesting that it would be easier to find a specific and sensitive biomarker for oral diseases in saliva than in serum. Our results demonstrated that the salivary protease spectrum was quite different between healthy donors and OLK or OSCC patients. Indeed, both the number of proteases and their levels significantly increased with disease progression (i.e., health donors
Furthermore, we selected two important proteases from the protease spectrum, KLK5 and uPA for further research as they were both clearly increased in the saliva samples and supernatants. Human kallikreins (KLKs) constitute a subgroup of 15 secreted trypsin or chymotrypsin-like serine proteases [36]. KLK5 is abundantly expressed in human OSCC and correlates with the formation of more aggressive OSCC tumors in a murine orthotopic OSCC model [37]. It may promote the metastatic dissemination of OSCC by promoting the loss of junctional integrity through the cleavage of desmoglein1 [38]. KLK5 can mediate PAR-2 activation, which regulates the expression of inflammation-associated mRNAs and microRNAs, thereby modulating the progression of oral tumors [39]. uPA belongs to the plasminogen activator system, which includes the plasminogen activators (uPA, tPA), the cell membrane receptor for uPA (uPAR), the plasminogen activator inhibitors (PAI-1, PAI-2), and plasmin [40, 41]. The uPA/uPAR complex plays a significant role in the invasive and metastatic potential of various tumors [42]. Active uPA increases the production of plasmin from plasminogen, leading to ECM degradation. This degradation facilitates cancer cell invasion into the surrounding tissue. Most head and neck cancer studies have shown that uPA/uPAR overexpression enhances tumor cell proliferation, migration, and invasion [43, 44]. The upregulation of uPA contributes to the promotion of oral tumor invasion by extracellular matrix metalloproteinase inducer (EMMPRIN) [44]. It also promotes ESCC cell proliferation, migration, and invasion via the PI3K/AKT and ERK signaling pathways [45]. Those studies indicated that KLK5 and uPA are important proteins in the development and metastasis of OSCC, but rare study of the two in precancerous lesions are performed.
In current study, we detected the contents of saliva KLK5 and uPA in healthy peoples, patients with OLK, OLP, OLK-T, OLP-T, and OSCC. Simultaneously, we tested the expression of them in OLK, OLP and OSCC histologic sections. The results confirmed that their expression levels progressively increased with the development of the OSCC. These data strongly demonstrated that KLK5 and uPA play an important role in the process of OLK and OLP cancerization. Furthermore, ROC curves analysis suggested that both KLK5 and uPA have high AUCs in the diagnosis of OLK and OLP, as well as OLK-T, OLP-T and OSCC. Based on the generated cutoff values for diagnosing OLK, OLP, OLK-T, OLP-T and OSCC, they all have high sensitivity but low specificity. However, the combined analysis of KLK5 and uPA not only significantly increased the specificity for the diagnosis of all five diseases but also allowed the differential diagnosis of OSCC from OLK and OLP. These results suggested that the combination of KLK5 and uPA could represent a potential biomarker for determining the malignant transformation of OLK or OLP into OSCC.
Conclusions
Our current study demonstrated that salivary KLK5 and uPA could be potential biomarkers for evaluating the malignant transformation of OLK and OLP and early diagnosis of OSCC. Importantly, their combination could aid in the differential diagnosis of OSCC from OLK and OLP. Unfortunately, our sample size was limited. The potential value of KLK5 and uPA as biomarkers requires a multicenter study involving a larger number of patients to verify their predictive efficacy.
Footnotes
Acknowledgments
This research was funded by the National Natural Science Foundation of China (No. 81372892) and the Interdisciplinary integration Foundation (2020KXK0405) from the State Key Laboratory of Oral Disease.
Conflict of interest
The authors declare that they have no conflicts of interest to disclose.
Author contributions
Conception: Yun Feng and Ping Zhang
Interpretation or analysis of data: Ping Zhang, Yingzhu Kang, Jiao Chen, Xiaoying Li, Min Luo, Hongli Chen, Bomiao Cui, Liwei Wang, Jingnan Wang, and Die Lv
Preparation of the manuscript: Ping Zhang and Yingzhu Kang
Revision for important intellectual content: Ping Zhang, Jiao Chen, and Xiaoying Li
