Abstract
BACKGROUND:
Circular RNA (circRNA) is a class of non-coding RNA that is vital for regulating gene expression and biological functions. Mounting studies demonstrate that circRNA is crucial for human cancer development. However, the role of circ_0000039 in gastric cancer (GC) remains uncertain.
METHODS:
Normal human gastric tissues and GC tissue samples were collected, and quantitative real-time polymerase chain reaction (qRT-PCR) was employed to detect the expression levels of circ_0000039, miR-1292-5p, and DEK. GC cell lines with overexpression and low expression of circ_0000039 were constructed. Cell counting kit-8 (CCK-8), scratch healing and Transwell experiments were used to assess the function of circ_0000039 on the proliferation, migration and invasion of GC cells. Bioinformatics analysis and dual-luciferase reporter assays were employed to detect the targeting relationship between circ_0000039 and miR-1292-5p.
RESULTS:
Circ_0000039 expression was up-regulated in GC tissues and cell lines, and it was significantly related with poor differentiation of tumor tissues. In addition, circ_0000039 overexpression enhanced the proliferation, migration and invasion of GC cells, while circ_0000039 depletion inhibited these malignant biological behaviors. In terms of mechanism, it was found that circ_0000039 promoted the proliferation and progression of GC cells by adsorbing miR-1292-5p and up-regulating the expression of DEK.
CONCLUSION:
Circ_0000039 is a new oncogenic circRNA in GC, which regulates the miR-1292-5p/DEK axis to modulate the malignant biological behaviors of GC.
Introduction
Gastric cancer (GC) is a common malignancy of the human digestive system, which is the fourth most common malignancies, as well as the third leading cause of cancer-related death in the world [1, 2, 3, 4]. Although radical gastrectomy and chemotherapy are widely used, the prognosis of the patients in advanced stage is still disappointing [5, 6]. Therefore, it is particularly important to understand the molecular mechanism of GC progression.
Circular RNA (circRNA) is a non-coding RNA commonly found in the cytoplasm of various eukaryotic cells [7]. It is formed by back-splicing from immature RNA, lacking 5’ caps and 3’ poly (A) tail [8]. Moreover, it possesses higher stability and evolutionary conservation than linear RNA [9]. CircRNAs have been recognized as promising biomarkers for the diagnosis of diverse diseases in recent years, and they participate in regulating the biological behaviors of cancer cells [10, 11]. For instance, knockout of the circCCDC66 inhibits tumor growth and invasion in colonic cancer [12]. Circ_0000190 expression is down-regulated in GC tissues, and dysregulation of circ_0000190 is remarkably linked to tumor diameter, lymph node metastasis, and TNM staging [13]. Circ_0000745 expression in GC tissue is closely related to the degree of tumor tissue differentiation [14]. However, the role of circ_0000039 in GC has yet to be fully elucidated.
Accumulating studies show that circRNA functions as a miRNA sponge to modulate the expression of genes, and this mechanism is associated with tumorigenesis and cancer progression [15]. For instance, circ_101280 promotes the carcinogenesis of hepatocellular carcinoma via adsorbing miR-375 and up-regulating JAK2 expression [16]. Circ-BPTF facilitates bladder cancer progression through the miR-31-5p/RAB27A axis [17]. In addition, recent research indicates that abnormal regulation of the circRNA-miRNA interaction is implicated in the occurrence and progression of GC [18]. For example, the expression of circRNA_100269 is down-regulated in GC and circRNA_100269 inhibits tumor cell growth via targeting miR-630 [11].
DEK is a nuclear protein with a molecular weight of 42 KDa, and involved in many biological processes, including heterochromatin integrity, transcriptional regulation, and DNA damage repair and so on [19, 20, 21, 22]. Moreover, DEK is significantly associated with the tumorigenesis and metastasis of diverse tumors [23, 24]. In GC, it is reported that miR-1292-5p targets DEK to suppress cancer cell proliferation, migration and invasion [25].
In this study, we found that circ_0000039 and DEK expressions were up-regulated in GC tissues, and showed a positive correlation. Interestingly, bioinformatics database Circinteractome (
Materials and methods
Clinical samples
In this study, 27 GC tissues and matched adjacent non-tumor tissues were obtained from the Caoxian People’s Hospital, and none of the patients enrolled in this study received preoperative radiotherapy or chemotherapy. The samples were obtained during surgery and then stored in liquid nitrogen for further analysis. The adjacent tissue, 5 cm from the margin of the tumor, was evaluated by two pathologists and non-tumor tissue was observed. Written informed consent was obtained from all patients. The experiments were conducted with the approval and guidance of the Ethics Committee of Caoxian People’s Hospital (Approval Number: CXRMYY-2017075).
Cell lines and culture
Four human GC cell lines (MGC-803, SGC-7901, BGC-823, and AGS) and human normal gastric mucosal epithelial cell lines (GES-1) were available from the Chinese Academy of Sciences Biochemistry and Cell Biology Research (Shanghai, China). All cells were cultured in Dulbecco Modified Eagles Medium (DMEM) (Gibco, Carlsbad, CA, USA) containing 10% fetal bovine serum (FBS, HyClone, Logan, UT, USA), 100 U/mL penicillin and 100
RNA extraction and quantitative real-time polymerase chain reaction (qRT-PCR)
In accordance with the instructions, total RNA in tissues and cells was extracted using TRIzol reagent (Invitrogen, Shanghai, China) and miRNA was extracted employing mirPremier
Cell transfection
Circ_0000039 overexpressing plasmid pcDNA3.1-circ_0000039 (pcDNA3.1-circ), DEK overexpressing plasmid pcDNA3.1-DEK (DEK) and control plasmid pcDNA3.1, siRNA against circ_0000039 (si-circ) and control siRNA (si-NC), miR-1292-5p mimics and inhibitors, and controls (NC mimics or NC inhibitors) were available from GenePharma (Shanghai, China). AGS and SGC-7901 cells were inoculated in 24-well plates at 3
Cell proliferation detection
Cell proliferation was assessed using the Cell Counting Kit 8 (CCK-8) experiment. Transfected cells were inoculated in 96-well plates at 1
Scratch healing experiment
Scratch healing experiment was employed to detect the motility of cancer cells. In short, after the cells cultured in a 6-well plate covered the bottom of the wells, a sterilized pipette was used to make a wound on the bottom of the wells. After that, the cells were gently washed with PBS twice, and serum-free DMEM was added. Then the scratch was observed and photographed under a microscope. After cell culture was continued for 24 h, the healing of scratch was observed and photographed, and the average scratch healing rate was calculated. Scratch healing rate (%)
Cell migration and invasion experiment
Circ_0000039 and DEK expression were up-regulated in the GC tissues A-B. qRT-PCR was used to detect the expression levels of circ_0000039 and DEK in GC tissues and matched adjacent non-tumor tissues of GC patients, the results of which showed that both circ_0000039 and DEK were up-regulated in GC samples. C. Expression correlation between circ_0000039 and DEK in GC samples was analyzed by Pearson’s correlation analysis, which indicated that circ_0000039 and DEK were positively correlated in GC samples. D. qRT-PCR was used to detect the expression level of circ_0000039 in normal gastric mucosal epithelial cell lines (GES-1) and GC cell lines (MGC-803, SGC-7901, BGC-823, and AGS), which implied that the expression of circ_0000039 was up-regulated in GC cell lines. *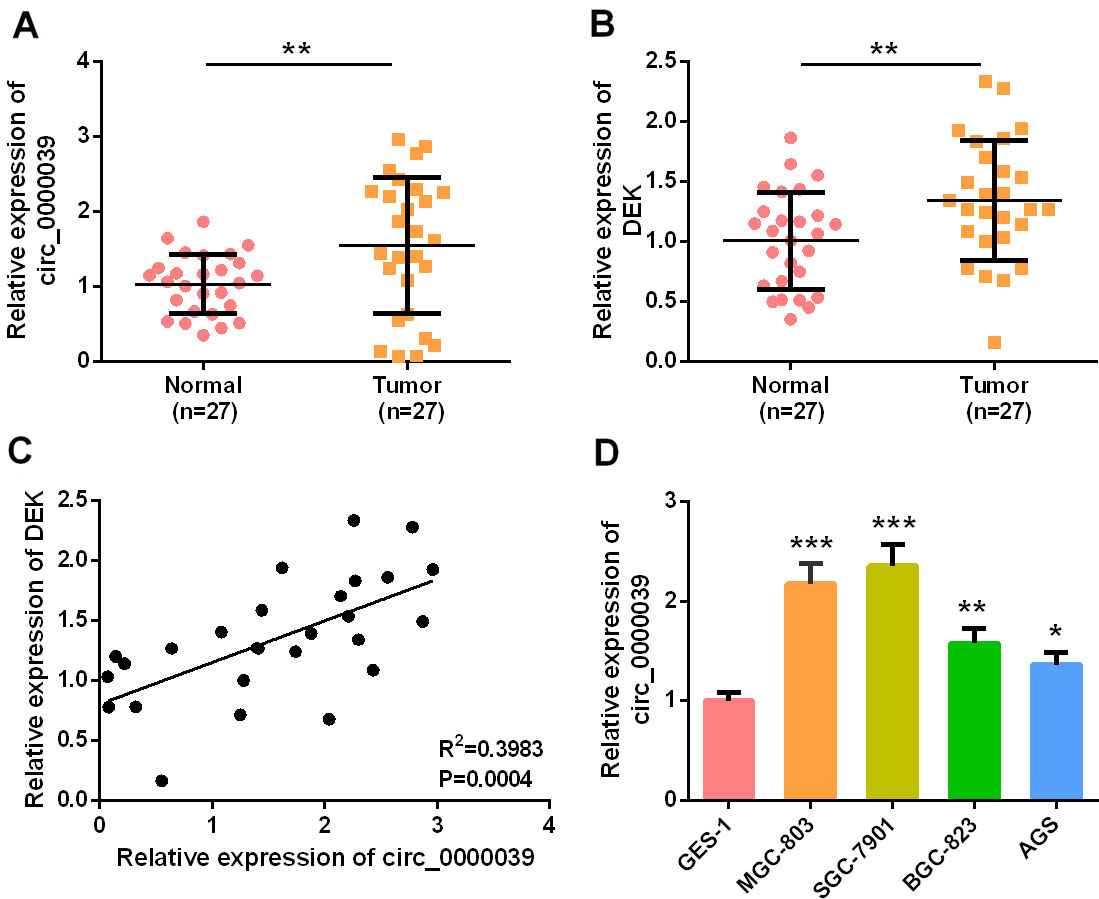
In migration experiment, after AGS and SGC-7901 cells were dispersed with 0.25% trypsin, the cells were centrifuged, resuspended, and dispersed with serum-free medium. 8
Luciferase reporter gene assay was performed using the dual-luciferase reporter assay system (Promega, Madison, WI, USA). The target fragments of wild type circ_0000039 and mutant circ_0000039 were constructed and integrated into pGL3 vector (Promega, Madison, WI, USA) to construct pGL3-circ_0000039-wild type (circ_0000039-WT) and pGL3-circ_0000039-mutant (circ_0000039-MUT) reporter vectors. Circ_ 0000039-WT or circ_0000039-MUT, and miR-1292-5p mimics or NC mimics were co-transfected into AGS and SGC-7901 cells. After 48 h of transfection, the luciferase activity of the cells in each group was determined according to manufacturer’s instruction.
Western blot
AGS cells transfected with pcDNA-3.1-circ and miR-1292-5p mimics, and SGC-7901 cells transfected with si-circ and miR-1292-5p inhibitors were collected. The medium was discarded and the cells were rinsed once with PBS. The total protein was extracted using RIPA lysis buffer (Beyotime Biotechnology, Shanghai, China). After the protein samples were denatured, SDS-PAGE was performed, and the protein was transferred to a PVDF membrane (Millipore, Bedford, MA, USA) by electrophoresis. After blocking the PVDF membrane with 5% skim milk solution for 2 h, primary antibodies DEK antibody (ab166624, 1: 1000, Abcam, Shanghai, China) and
Effect of circ_0000039 overexpression on the growth, migration and invasion of AGS and SGC-7901 cells. A. Circ_0000039 overexpression and knockdown cell line were constructed, and the expression level of circ_0000039 in cells was detected by qRT-PCR. B. CCK-8 experiments were used to analyze the effect of circ_0000039 on the proliferation of AGS and SGC-7901 cells, which indicated that circ_0000039 overexpression promoted the proliferation of AGS cells, and circ_0000039 knockdown repressed the proliferation of SGC-7901 cells. C. The effect of circ_0000039 expression on AGS and SGC-7901 cell movement was analyzed by scratch healing experiment, which indicated that circ_0000039 overexpression promoted the motility of AGS cells, and circ_0000039 knockdown inhibited the motility of SGC-7901 cells. D. Transwell assay was used to analyze the effect of circ_0000039 on the migration and invasion of AGS and SGC-7901 cells, and the results suggested that circ_0000039 overexpression promoted the migration and invasion of AGS cells, and circ_0000039 knockdown inhibited the migration and invasion of SGC-7901 cells. *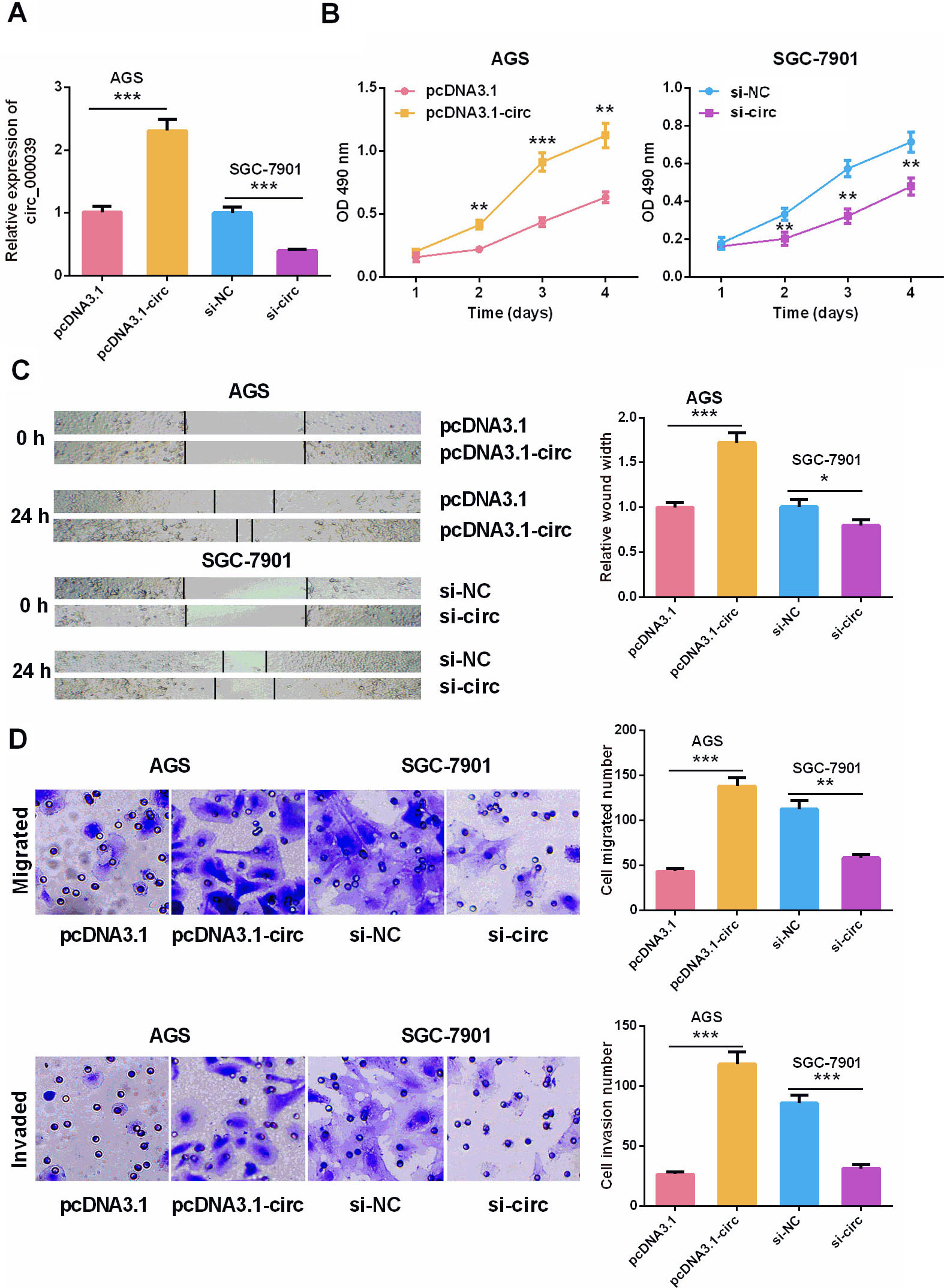
All experiments were conducted in triplicate and repeated three times. All statistical analyses were performed using SPSS 20.0 software (IBM, SPSS, Chicago, IL, USA). Data were expressed as mean
Association between circ_0000039 expressions and clinicopathologic features
Association between circ_0000039 expressions and clinicopathologic features
Circ_0000039 and DEK expressions were up-regulated in the GC tissues
First of all, the expression level of circ_0000039 in GC samples and adjacent non-tumor tissue samples was detected by qRT-PCR. It was revealed that circ_0000039 was highly expressed in GC tissues (Fig. 1A). Meanwhile, the expression of DEK was also analyzed by qRT-PCR, and the data demonstrated that DEK expression in GC tissue was also remarkably higher than that of non-tumor tissues (Fig. 1B). Importantly, correlation analysis implied that circ_0000039 expression was positively correlated with DEK expression in GC tissues (Fig. 1C). In addition, the expression of circ_0000039 in GC cell lines was significantly increased compared to in human normal gastric mucosal epithelial cell line GES-1, further confirming that circ_0000039 expression was up-regulated in GC (Fig. 1D). Subsequently, the relationship between circ_0000039 expression and different clinicopathological characteristics was analyzed by Chi-square test, and the results showed that the expression of circ_0000039 was closely related to poor differentiation of tumor tissues (Table 1). Among the GC cell lines, the expression of circ_0000039 is relative higher in SGC-7901 cell line, and relatively lower in AGS cell line. Thus, AGS and SGC-7901 cells were chosen to be applied to the following gain-of-function and loss-of-function experiments respectively.
Circ_0000039 targeted miR-1292-5p. A. Sequence of targeted binding sites of circ_0000039 and miR-1292-5p. B. qRT-PCR was used to detect the expression level of miR-1292-5p in GC tissues and matched adjacent non-tumor tissues of GC patients, and the results suggested that miR-1292-5p was significantly reduced in GC tissues. C. The correlation between circ_0000039 and miR-1292-5p expressions was analyzed, and the results showed that circ_0000039 expression and miR-1292 expression were negatively correlated. D. qRT-PCR was used to detect the expression level of miR-1292-5p in AGS and SGC-7901 cell lines after circ_0000039 was overexpressed or knocked down, and as shown, circ_0000039 overexpression repressed the expression of miR-1292-5p and circ_0000039 promoted the expression of miR-1292-5p. E. Dual-luciferase reporter assay was used to verify the relationship between circ_0000039 and miR-1292-5p in AGS and SGC-7901 cells. *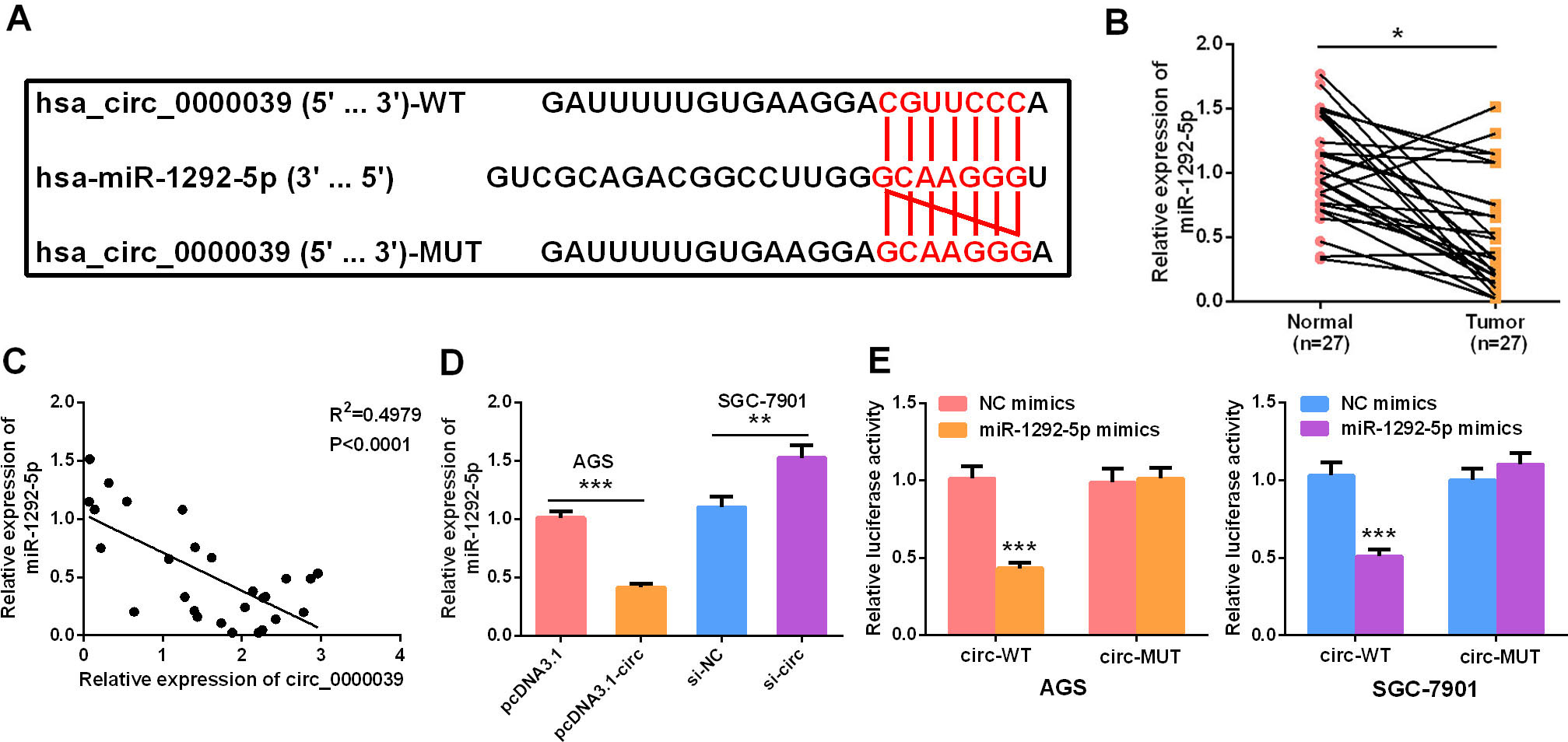
Next, pcDNA3.1-circ_0000039 overexpressing vector was transfected into AGS cells and circ_0000039 overexpression model was constructed. Circ_0000039 siRNA was transfected into GC cell line SGC-7901, and the low expression model of circ_0000039 was constructed. The transfection efficiency was determined 24 h after the transfection (Fig. 2A). CCK-8 experiments implied that overexpression of circ_0000039 facilitated the growth of AGS cells, and knocking down circ_0000039 inhibited the proliferation of SGC-7901 cells (Fig. 2B). The results of scratch healing and Transwell experiments revealed that circ_0000039 overexpression enhanced the motility, migration and invasion of AGS cells, while circ_0000039 knockdown inhibited these malignant biological behaviors of SGC-7901 cells (Fig. 2C–D). These results suggested that circ_0000039 was a cancer-promoting factor in GC.
Circ_0000039 acted as a miRNA sponge for miR-1292-5p in AGS and SGC-7901 cells
The potential targets of circ_0000039 were searched in the Circinteractome. Circ_0000039 was predicted to have a binding site for miR-1292-5p (Fig. 3A). The expression of miR-1292-5p in GC tissues and adjacent non-tumor tissues was detected by qRT-PCR. Compared with in adjacent non-tumor tissues, miR-1292-5p expression in GC tissues was significantly reduced (Fig. 3B), and the expression of circ_0000039 in GC tissues was negatively correlated with the expression of miR-1292-5p (Fig 3C). Moreover, circ_0000039 overexpression in AGS cells markedly inhibited miR-1292-5p expression, and inhibition of circ_0000039 significantly increased the expression of miR-1292-5p in SGC-7901 cells (Fig 3D). Furthermore, dual-luciferase reporter gene experiments confirmed that miR-1292-5p mimics significantly inhibited the luciferase activity of wild-type circ_0000039 reporter in AGS and SGC-7901 cell lines, but could not significantly reduce the luciferase activity of the mutant circ_0000039 reporter, validating the binding relationship between circ_0000039 and miR-1292-5p (Fig. 3E).
MiR-1292-5p overexpression partly reversed the oncogenic effect of circ_0000039
To pinpoint the role of circ_0000039/miR-1292-5p axis in GC cells, miR-1292-5p mimics were transfected into AGS cells overexpressing circ_0000039, and SGC-7901 with circ_0000039 knocked down cells was transfected with miR-1292-5p inhibitors (Fig. 4A). After that, the biological behaviors of the AGS and SGC-7901 cells were detected by CCK-8 experiment, scratch healing experiment, and Transwell experiment, respectively. The results implied that overexpression of miR-1292-5p significantly inhibited the promotion of AGS cell proliferation, movement, migration and invasion induced by circ_0000039 overexpression (Fig. 4B–D). Conversely, the knockdown of circ_0000039 caused the inhibition of SGC-7901 cell proliferation, movement, migration, and invasion, which was partially attenuated by miR-1292-5p inhibitors (Fig. 4B–D). These results further indicated that circ_0000039 participated in regulating GC cell proliferation and metastasis via adsorbing miR-1292-5p.
MiR-1292-5p overexpression inhibited the oncogenic effect of circ_0000039. A. circ_0000039 overexpressed AGS cells were transfected with miR-1292-5p mimics; SGC-7901 cells with circ_0000039 knockdown were transfected with miR-1292-5p inhibitors. qRT-PCR was used to detect the expression level of miR-1292-5p. B. CCK-8 experiments were used to analyze the proliferation of AGS and SGC-7901 cells, and the results showed that miR-1292-5p mimics partly abolished the function of circ_0000039 overexpression, and miR-1292-5p inhibitors partly reversed the effects of circ_0000039 knockdown. C. Scratch healing experiments were used to detect the movement of AGS and SGC-7901 cells, and the results showed that miR-1292-5p mimics partly abolished the function of circ_0000039 overexpression, and miR-1292-5p inhibitors partly reversed the effects of circ_0000039 knockdown. D. Transwell assay was used to analyze the migration and invasion of AGS and SGC-7901 cell lines, and the results showed that miR-1292-5p mimics partly abolished the function of circ_0000039 overexpression, and miR-1292-5p inhibitors partly reversed the effects of circ_0000039 knockdown. *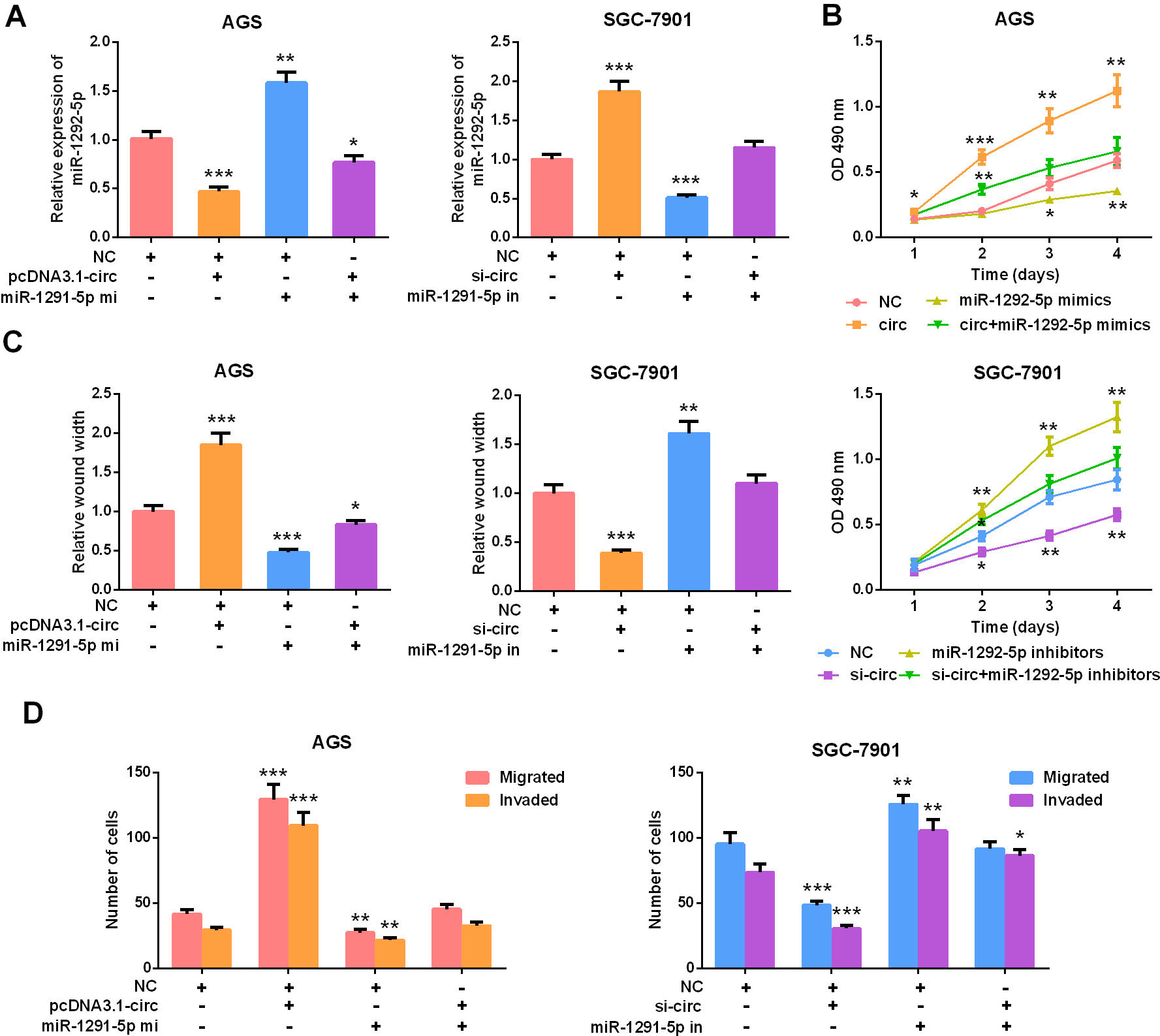
Targeted binding between the 3’UTR of DEK and miR-1292-5p is validated by a previous study [25] (Fig. 5A). In this work, qRT-PCR and Western blot were employed to detect DEK expression in AGS and SGC-7901 cell lines after circ_0000039 and miR-1292-5p expressions were artificially regulated. The results showed that overexpression of circ_0000039 enhanced the expressions of DEK mRNA and protein, while the transfection of miR-1292-5p mimics significantly inhibited the expression of DEK (Fig. 5B–C). Subsequently, pcDNA3.1-DEK overexpressing plasmid was transfected into the SGC-7901 cells with circ_0000039 knockdown (Fig. 5D). The results of the CCK-8 experiment, the scratch healing experiment, and the Transwell experiment showed that the knockdown of circ_0000039 inhibited the proliferation, motility, migration, and invasion of SGC-7901 cells (vs. NC SGC-7901 cell line), while the overexpression of DEK partially restored these biological processes of SGC-7901 cells (vs. NC
DEK overexpression restored the oncogenic effect of circ_0000039. A. The sequence of targeted binding sites of DEK and miR-1292-5p was predicted by bioinformatics analysis. B-C. qRT-PCR and Western blot were used to analyze the expressions of DEK mRNA and protein in AGS and SGC-7901 cells after the transfection, and the results showed that miR-1292-5p mimics and inhibitors repressed and promoted the expression of DEK respectively, and circ_0000039 overexpression and knockdown respectively counteracted the function of miR-1292-5p mimics and inhibitors. D. pcDNA3.1-DEK was transfected into SGC-7901 cells with circ_0000039 knocked down, and the expression of DEK was detected by qRT-PCR, and the results showed that circ_0000039 knockdown repressed the expression of DEK, and after pcDNA3.1-DEK was transfected, the expression of DEK was restored. E. CCK-8 was used to detect the proliferation of SGC-7901 cells, and the results suggested circ_0000039 knockdown repressed the proliferation, while DEK overexpression reversed it. F. Scratch healing experiment was used to analyze the movement of SGC-7901 cells, and the results suggested circ_0000039 knockdown repressed the motility, while DEK overexpression reversed it. G. Transwell assay was used to analyze the migration and invasion of SGC-7901 cells, and the results suggested circ_0000039 knockdown repressed the migration and invasion, while DEK overexpression reversed it. *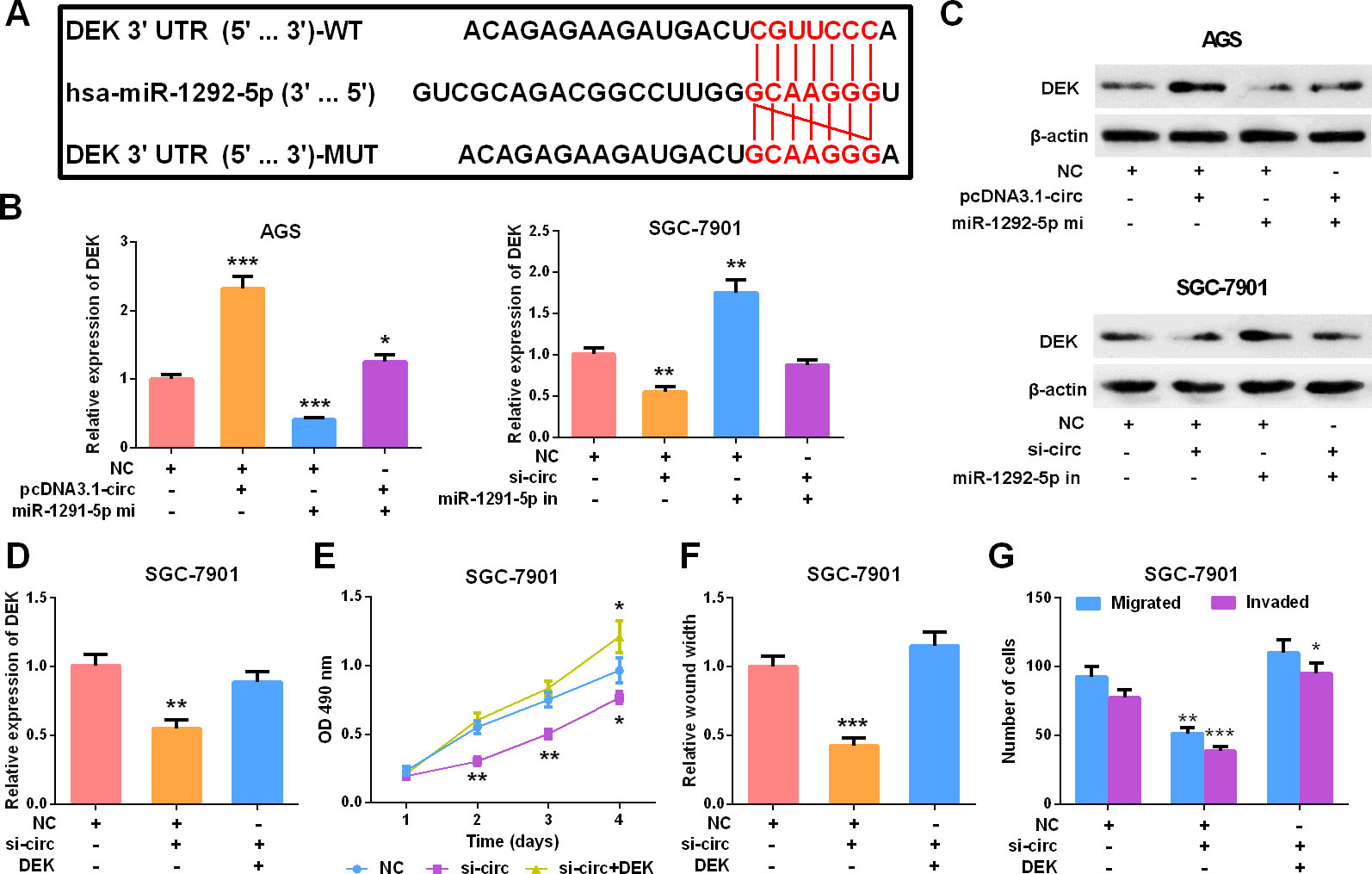
CircRNAs were originally thought to be byproducts of RNA transcription and splicing and have low expression abundance. However, recent studies authenticate that circRNA possesses a highly stable and specific circular structure and are promising biomarker for the diagnosis and prognosis evaluation of cancers [26, 27]. Functionally, circRNAs can regulate key biological behaviors in human malignancies, such as cell proliferation, migration, and invasion [28, 29, 30, 31]. For instance, it is revealed that circ_0067934 is up-regulated in esophageal squamous cell carcinoma and is involved in promoting cancer cell proliferation and migration [32]. The inhibition of circPVRL3 enhances the proliferation and migration of GC cells [33]. Nevertheless, the role of circ_0000039 in GC has not been studied previously. In this study, for the first time, it was found that circ_0000039 expression in human GC tissues was remarkably higher than that in non-tumor tissues; up-regulation of circ_0000039 expression was significantly related to poor differentiation of tumor tissues. Further experiments showed that over-expression of circ_0000039 enhanced the proliferation, motility, migration and invasion of GC cells. These demonstrations implied that circ_0000039 played a role as a novel oncogenic circRNA in promoting the development of GC.
Competitive endogenous RNAs (ceRNAs) include mRNA, pseudogenic RNAs, long noncoding RNAs (lncRNAs) and circRNAs, which are transcripts that competitively sponge miRNAs to reduce their expression and function, and regulate the expression of downstream genes. Recent studies reveal that circRNAs figure prominently in regulating gene expression by targeting specific miRNAs [34]. Accumulating data demonstrate that ceRNA mechanism in which circRNAs participate is involved in promoting or inhibiting the tumorigenesis and development of malignancies [35]. For instance, circ_103809 can sponge miR-620 and negatively regulate its expression, thereby further inhibiting the proliferation and invasion of liver cancer cells [36]. In addition, circPSMC3 is identified as a molecular sponge for miR-296-5p to regulate the expression of PTEN and further inhibit the tumorigenesis of GC [37]. In this study, bioinformatics analysis revealed that circ_0000039 could probably target miR-1292-5p, and dual-luciferase and in vitro experiments validated this prediction. It was also demonstrated that miR-1292-5p overexpression repressed the oncogenic effect of circ_0000039. This phenomenon indicated that miR-1292-5p was a crucial downstream miRNA by which circ_0000039 exerted its function in GC.
Proto-oncogene DEK is a highly conserved chromatin-bound protein, and its high expression helps keep genome stability and facilitates the DNA double strand breaks repair of cells [22, 38]. It is reported that DEK is involved in the tumorigenesis and development of diverse tumors. DEK is verified to facilitate the invasion of breast cancer cells via modulating
In conclusion, this study finds that circ_0000039 expression is significantly elevated in GC tissues and closely associated with the differentiation status of tumor tissues. Our data also reveal that circ_0000039 is an oncogenic factor in GC, which can modulate miR-1292-5/DEK axis to affect the malignant biological behaviors of GC cells, providing a better understanding of the mechanism of GC carcinogenesis and development.
Footnotes
Funding
This study is supported by the Research Funding for Clinical Researchers of Caoxian People’s Hospital (Approval No. CXRMYY-17075).
