Abstract
Background:
microRNAs (miRNAs) are non-coding RNAs with important roles in the progression of human cancers, including gastric cancer. Exosomes are extracellular vesicles, which could transfer numerous noncoding RNAs, such as miRNAs. Here, in our study, we intended to investigate the role of exosomal miR-122-5p in gastric cancer progression.
Methods:
Exosomes were isolated utilizing commercial kit or ultracentrifugation. Biomarkers of exosomes or epithelia-mesenchymal transition (EMT) were monitored by western blot. Expression levels of miR-122-5p and G-protein-coupled receptor kinase interacting protein-1 (GIT1) were evaluated by quantitative reverse transcription-polymerase chain reaction (qRT-PCR) or western blot. Cell proliferation and apoptosis were assessed by colony formation assay, methyl thiazolyl tetrazolium assay and flow cytometry. Cell metastasis was evaluated via Transwell assay. The interaction between miR-122-5p and GIT1 was validated by dual-luciferase reporter assay. Furthermore, tumor growth in vivo was detected by xenograft assay.
Results:
Exosomes were successfully isolated. MiR-122-5p was downregulated in exosomes derived from the serum of gastric cancer patients. Exosomal miR-122-5p could hinder gastric cancer cell proliferation and metastasis in vitro and tumor growth in vivo. Knockdown of GIT1 also inhibited gastric cancer cell proliferation and metastasis. Exosomal miR-122-5p targeted GIT1 to alter cellular behaviors of gastric cancer cells.
Conclusion:
Exosomal miR-122-5p suppressed gastric cancer progression by targeting GIT1.
Introduction
Gastric cancer (GC) is a common, deadly, solid tumor, severely threating human health. 1 Every year, more than 1 million newly diagnosed cases and deaths occur across the world. 2 Though early diagnosis, surgery, radiotherapy, and chemotherapy have worked a great deal, most GC cases diagnosed at advanced stages still have a poor prognosis. 3 Therefore, it is extremely urgent to deeply explore the molecular mechanism of GC progression, to search novel avenues for GC therapy.
Exosomes are little extracellular vesicles (diameter: 40–150 nm), which could be released by most cell types, including cancer cells, and could impact tumor invasion and aggravation. 4 GC-derived exosomes were reported to function in remodeling the pre-metastatic microenvironment and supply novel mechanistic pathways for peritoneal metastasis of GC. 5 A growing body of evidence has proved the important roles of exosomal microRNAs (miRNAs) in tumor progression. 6 For example, exosomal miR-423-5p contributed to the growth and metastasis of GC via targeting the SUFU gene. 7 Exosomal miR-21-5p derived from GC was demonstrated to be an oncogenic miRNA, promoting mesothelial-to-mesenchymal transition of peritoneal mesothelial cells and tumor peritoneal dissemination by targeting SMAD7. 8 In addition, miR-122-5p has been uncovered to play an important role in GC, 9 while the biological functions of exosomal miR-122-5p in GC have not been fully corroborated.
G-protein-coupled receptor kinase interacting protein-1 (GIT1) is associated with focal adhesion and cell metastasis of diverse human cancers, including GC.10, 11 MiRDB (http://mirdb.org/) forecasts that miR-122-5p endows with binding region with the 3′ untranslated region (3′UTR) of GIT1. However, the effect of GIT1 on exosomal miR-122-5p mediated in GC tumorigenicity is still unclear.
In this project, we investigated the functional effects of exosomal miR-122-5p on GC tumorigenesis in vitro and in vivo, and the underlying mechanistic pathway was also explored.
Materials and methods
Serum sample collection
Prior to this study, permission was obtained from the Ethics Committee of The First People’s Hospital of Lanzhou City. Blood samples were obtained from GC patients (GC) and healthy volunteers (HC) recruited at The First People’s Hospital of Lanzhou City. All participators signed written, informed consent before blood sampling. Serum was isolated from blood samples via centrifugation at 4°C, 400 × g for 15 min, then preserved at −80°C.
Cell culture
The human gastric mucosal cell line GES-1 (BNCC353464) was commercially acquired from BeNa Culture Collection (Beijing, China). Human gastric cancer cell lines AGS (CL-0022) and MKN45 (CL-0292) were purchased from Procell (Wuhan, China). Human gastric cancer cell line HGC27 (TCHu 22) was supplied by Cell Bank of the Chinese Academy of Sciences (Shanghai, China). Human gastric cancer cell line MKN74 (CBP60490) was afforded by COBIOER (Nanjing, China). All cell lines were maintained in Roswell Park Memorial Institute 1640 Medium added with 10% fetal bovine serum, 10 mM HEPES, and 1% penicillin-streptomycin (all provided by Invitrogen, Carlsbad, CA, USA) at 37°C in a humidified atmosphere including 5% CO2.
Exosome isolation and characterization
The isolation of exosomes in serum samples was implemented using ExoQuick exosome precipitation kit (System Biosciences, Palo Alto, CA, USA) following the user’s manual. In brief, ExoQuick reagent was added into the serum sample, the mixture was incubated at 4°C for 30 min and centrifugated at 4°C, 1500 × g for 30 min. After removal of supernatant and centrifugation again for 5 min, exosomes were procured and re-suspended in 50 μL phosphate-buffered saline (PBS), and quantified exploiting a bicinchoninic acid protein assay kit (Tanon, Shanghai, China).
Supernatant of cell culture medium of GC cells was collected, then centrifuged at 2000 × g for 30 min to remove cells. Cell-free culture supernatant was treated with Total Exosome Isolation Reagent (Invitrogen) at 4°C for 12 h. After ultracentrifugation (100,000 × g), exosomes were obtained and re-suspended in PBS.
Transmission electron microscopy was used to monitor the shape of isolated exosomes with JEM-1200 EXII transmission electron microscope (TEM; JEOL, Tokyo, Japan) referring to the recommended instructions.
Western blot analysis
Radio-Immunoprecipitation Assay (RIPA; Vazyme, Nanjing, China) lysis buffer was applied to separate protein samples from exosomes and cultured cells. The protein samples were subjected for electrophoresis via 10% sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) and wet electrophoretic transfer onto polyvinylidene fluoride membranes (EMD Millipore, Billerica, MA, USA). After blockage in 5% fat-free milk for 1 h, all membranes were immersed in diluted primary antibodies, then incubated with horseradish peroxidase (HRP) combined secondary antibody (ab205718, 1:5000; Abcam, Shanghai, China). In addition, protein bolts were visualized using an electrochemiluminescent kit (PerkinElmer Life Science, Waltham, MA, USA) and quantified by Image J software (NIH, Bethesda, MD, USA). All primary antibodies were purchased from Abcam: anti-HSP70 (ab181606, 1: 1000), anti-Tsg101 (ab125011, 1: 1000), anti-CD63 (ab134045, 1: 500), anti-Twist1 (ab49254, 1: 1000), anti-E-cadherin (ab40772, 1: 2000), anti-GIT1 (ab156001, 1: 2000), and anti-glyceraldehyde-phosphate dehydrogenase (GAPDH) (ab181602, 1: 2500). Among these, GAPDH performed as loading control.
Exosome disposition and cell transfection
AGS and HGC27 cells were incubated with 40 μg/mL exosomes isolated from corresponding culture medium or extra exosome inhibitor GW4869 (MedChemExpress, Shanghai, China). Furthermore, cells treated with PBS served as control. Cellular behaviors were then examined.
miR-122-5p mimic (miR-122-5p), small interference RNA (siRNA) against GIT1 (si-GIT1), overexpression vector of GIT1 (GIT1) and their corresponding negative control miR-NC, si-NC, and pcDNA were all supplied by GenePharma Co. Ltd. (Shanghai, China). These oligonucleotides or plasmids were introduced into AGS or HGC27 cells utilizing Lipofectamine 3000 (Invitrogen) for 1 d, 2 d, or 3 d. Exosomes isolated from these transfected cells were added into corresponding cell culture medium. The nucleotide sequence of si-GIT1 was 5′-GUGCCAAUAUGAGCUCAGUTT-3′ (sense) and 5′-AGUGAGCUCAUAUUGGCACTT-3′ (anti-sense), and the nucleotide sequence of si-NC was 5′-UUCUCCGAACGUGUCACGUTT-3′ (sense), and 5′-ACGUGACACGUUCGGAGAATT-3′ (anti-sense).
Quantitative reverse transcription-polymerase chain reaction
Total RNA was extracted from cultured cells and exosomes using TRIzol Reagent (Invitrogen) and Qiazol and RNeasy MinElute Spin Column (Qiagen, Germantown, MD, USA) according to the manufacturer’s guideline, respectively. Reverse transcription was conducted with PrimeScript RT Reagent kit (Takara, Otsu, Japan) to generate complementary DNA (cDNA). Then SYBR Premix ExTaq kit was applied for quantitative reverse transcription-polymerase chain reaction (qRT-PCR) assay to test the relative expression level of GIT1, with GAPDH as internal control. With regard to miRNA, TaqMan MicroRNA Reverse Transcription Kit (Thermo Fisher Scientific, Waltham, MA, USA) and TaqMan MicroRNA Assays (Thermo Fisher Scientific) were used for reverse transcription and qRT-PCR assay, for determination of miR-122-5p level, normalized to U6. The relative expression of GIT1 and miR-122-5p was computed by the 2-ΔΔCt formula. Primers used for qRT-PCR assay were: miR-122-5p forward primer 5′-TATTCGCACTGGATACGACACAAAC-3′ and reverse primer 5′-GCCCGTGGAGTGTGACAATGGT-3′; U6 forward primer 5′-GCTTCGGCAGCACATATACTAAAAT-3′ and reverse primer 5′-CGCTTCACGAATTTGCGTGTCAT-3′; GIT1 forward primer 5′-AGGACGACGCCATCTATTCA-3′ and reverse primer 5′-CCCACTTTGCGTGTTCTCAT-3′; GAPDH forward primer 5′-TGCACCACCAACTGCTTAGC-3′ and reverse primer 5′-GGCATGGACTGTGGTCATGAG-3′.
Colony formation assay
To assess the colony formation capacity of GC cells, colony formation assay was carried out. After digestion with trypsin, treated AGS and HGC27 cells were plated onto 6-well plates (500 cells/well), then routinely cultured for 2 weeks until visible colonies generated. Then, colonies were dyed with Crystal violet (Beyotime, Shanghai, China) for 15 min, then observed and counted under a microscope (Olympus X-71; Olympus, Tokyo, Japan).
Methyl thiazolyl tetrazolium (MTT) assay
The current assay was employed to test cell viability of GC cells. Treated AGS and HGC27 cells seeded onto 96-well plates (1 × 103 cells/well) were cultured for the indicated time (0 d,1 d, 2 d, or 3 d) and then disposed with 20 μL 5 g/L MTT solution (Beyotime) for 4 h. After removing MTT, 150 μL dimethyl sulfoxide (Sigma-Aldrich, St. Louis, MO, USA) was dropped into each well. Five min later, absorbance of each well at 490 nm was recorded using a Microplate Reader (Bio-Rad, Hercules, CA, USA).
Flow cytometry
After different treatment, AGS and HGC27 cells were seeded in 6-well plates for 48 h, the harvested, stained with Annexin V-fluorescein isothiocyanate (FITC) reagent from Annexin V-FITC Apoptosis Detection Kit (Beyotime), then treated with propidium iodide for 20 min in a dark place. Apoptotic cells were monitored exploiting a flow cytometry (BD Biosciences, Franklin Lakes, NJ, USA). The data were processed utilizing CellQuest software (BD Biosciences).
Transwell assay
Treated AGS and HGC27 cells at an exponential growth phase were subjected for Transwell migration and invasion assays to measure cell metastasis utilizing Transwell chamber (BD Biosciences). For invasion assay, GC cells suspended in serum-free Medium (5 × 104 cells per well) were plated onto upper chamber pre-coated with Matrigel (BD Biosciences), while medium with 10% serum was added in the lower chamber. After 24-h routine culture, cells invaded to the bottom of the upper chamber were immobilized with methanol, stained by Crystal violet, then observed using a microscope (100 ×). The cell number was counted and averaged from five randomly selected fields. For the migration assay, the experiment procedures were similar to the invasion assay, except that the upper chamber was Matrigel-free and contained 1 × 104 cells each.
Dual-luciferase reporter assay
The possible target genes of miR-122-5p were forecasted by online tool miRDB (http://mirdb.org/), and the 3′ untranslated region (3′UTR) of GIT1 contained a binding region with miR-122-5p. To confirm the interaction between them, wild type luciferase reporter vector of GIT1 3′UTR was built by inserting its fragment containing binding sites into luciferase reporter vector pMirGLO, namely GIT1-wt. Likewise, mutant type luciferase reporter vector, GIT1-mut, was constructed after binding sites were mutated. Subsequently, AGS and HGC27 cells were co-transfected with GIT1-wt or GIT1-mut and miR-122-5p or miR-NC utilizing Lipofectamine 3000. Forty-eight hours later, luciferase activity was detected using the Dual-Luciferase reporter system (Beyotime) based on the manufacturer’s instructions.
Xenograft model assay
Experiment on animals was ratified by the Ethics Committee of The First People’s Hospital of Lanzhou City. Female nude mice (4–6 weeks old; BALB/c) were bought from Beijing Laboratory Animal Center (Beijing, China). AGS cells (1 × 106) stably transfected with miR-122-5p or miR-NC were subcutaneously inoculated into the right flank of each mouse (n=5). The size of formed tumors was recorded once a week. After 4 weeks, all mice were killed in order to excise the tumors for weigh, qRT-PCR, and western blot assay. Additionally, the volume of formed tumors was computed by the following formula: 0.5 × (short diameter) 2 × (long diameter).
Statistical analysis
All experiments were performed at least three times with three biological repetitions. Experimental data were processed SPSS 22.0 (IBM Corp., Armonk, NY, USA) and shown as mean ± SD. Differences were analyzed by the Student’s t-test (for two groups) or one-way analysis of variance (for three or more groups). When P < 0.05, the difference was regarded as statistically significant.
Results
MiR-122-5p was downregulated in exosomes derived from the serum of GC patients and GC cells
At first, exosomes were successfully isolated from GC patients (EXO-GC) and healthy volunteer controls (EXO-HC) using commercial kit. The images were taken using TEM. As displayed in Figure 1(a), isolated exosomes were round vesicles. Next, western blot analysis manifested that the protein levels of exosome biomarkers (HSP70, Tsg101, and CD63) in EXO-GC were significantly higher than those in EXO-HC (Figure 1(b)). Meanwhile, we also examined miR-122-5p expression in EXO-HC and EXO-GC. Data from qRT-PCR assay showed that miR-122-5p was downregulated in EXO-GC when compared to that in EXO-HC (Figure 1(c)). In addition, qRT-PCR assay also confirmed the downregulation of miR-122-5p in AGS, HGC27, MKN45, and MKN74 cells relative to GES-1 cells (Figure 1 (d)). Taken together, miR-122-5p was downregulated in EXO-GC and GC cells.
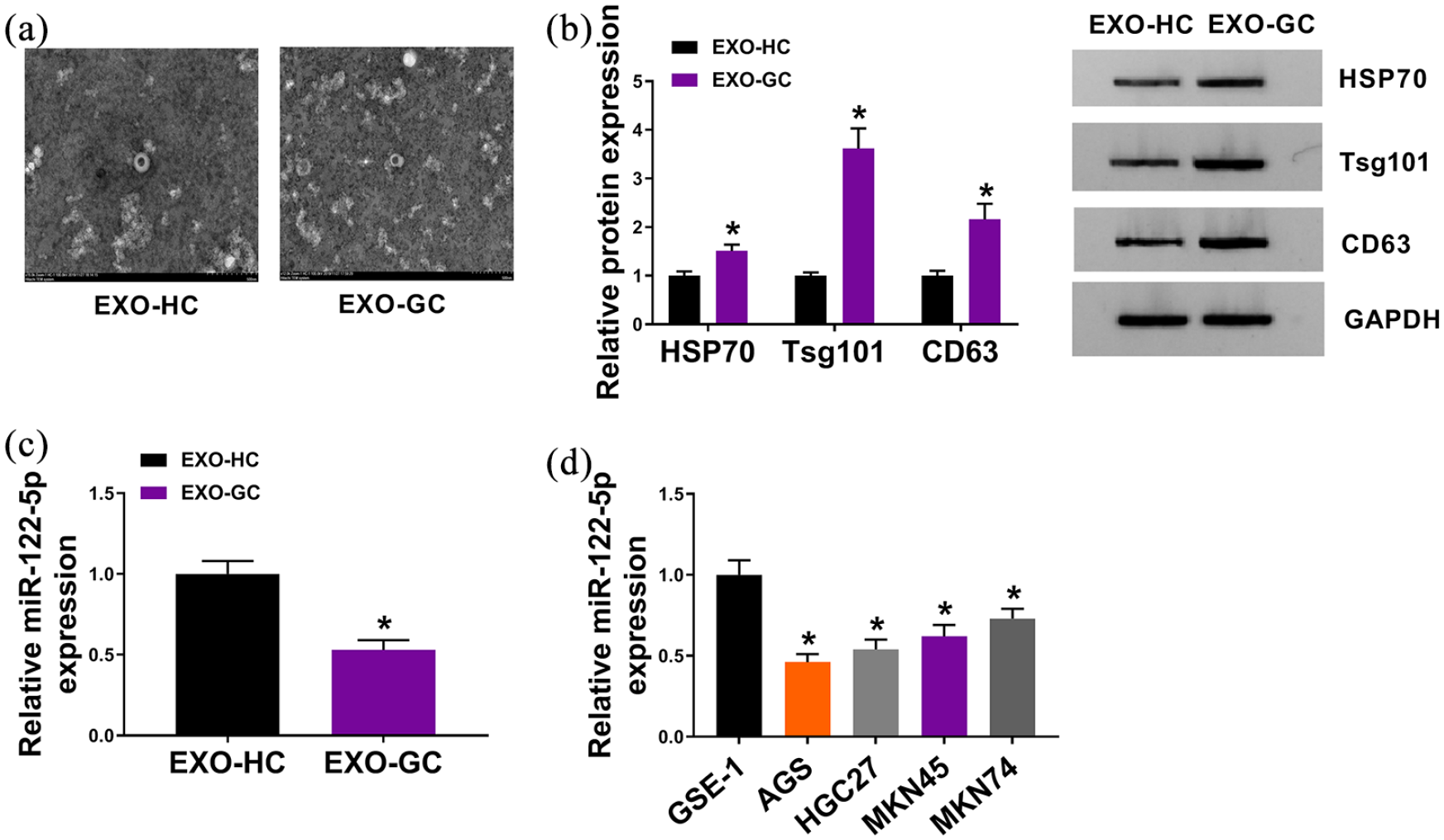
MiR-122-5p was downregulated in exosomes derived from the serum of GC patients and GC cells. (a) Electron micrograph of exosomes isolated from GC patients (EXO-GC) and healthy volunteer controls (EXO-HC). The scale bar represents 100 nm. (b) Western blot analysis for exosome biomarkers (HSP70, Tsg101, and CD63) in EXO-GC and EXO-HC. (c–d) qRT-PCR assay for expression level of miR-122-5p in EXO-GC and EXO-HC, as well as in GES-1, AGS, HGC27, MKN45, and MKN74 cells. *P < 0.05.
Exosomes derived from GC cells promoted cell proliferation, metastasis, and inhibited apoptosis of GC cells
To identify the function of exosomes in GC progression, exosomes (EXO) were also isolated from GC cells. AGS and HGC27 cells were disposed with PBS (control), EXO or EXO + GW4869 (exosome inhibitor) (Figure 2(b)). Results from western blot analysis revealed that GW4869 obviously reduced the expression of HSP70, Tsg101, and CD63 proteins, suggesting its inhibitory roles (Figure 2(a)). As displayed in Figure 2(c), introduction of EXO remarkably elevated the number of colonies, while GW4869 reversed the promoted effects. Furthermore, GW4869 also reversed EXO-mediated elevated cell viability of AGS and HGC27 cells (Figure 2(d) and (e)). Flow cytometry disclosed that EXO greatly repressed apoptosis of the two cell lines, which was facilitated by GW4869 (Figure 2(f)). Transwell assay and western blot analysis suggested that EXO treatment evidently enhanced cell metastasis of AGS and HGC27 cells, which was weakened by GW4869 (Figure 2(g) to (j)). Therefore, exosomes derived from GC cells promoted GC development in vitro.
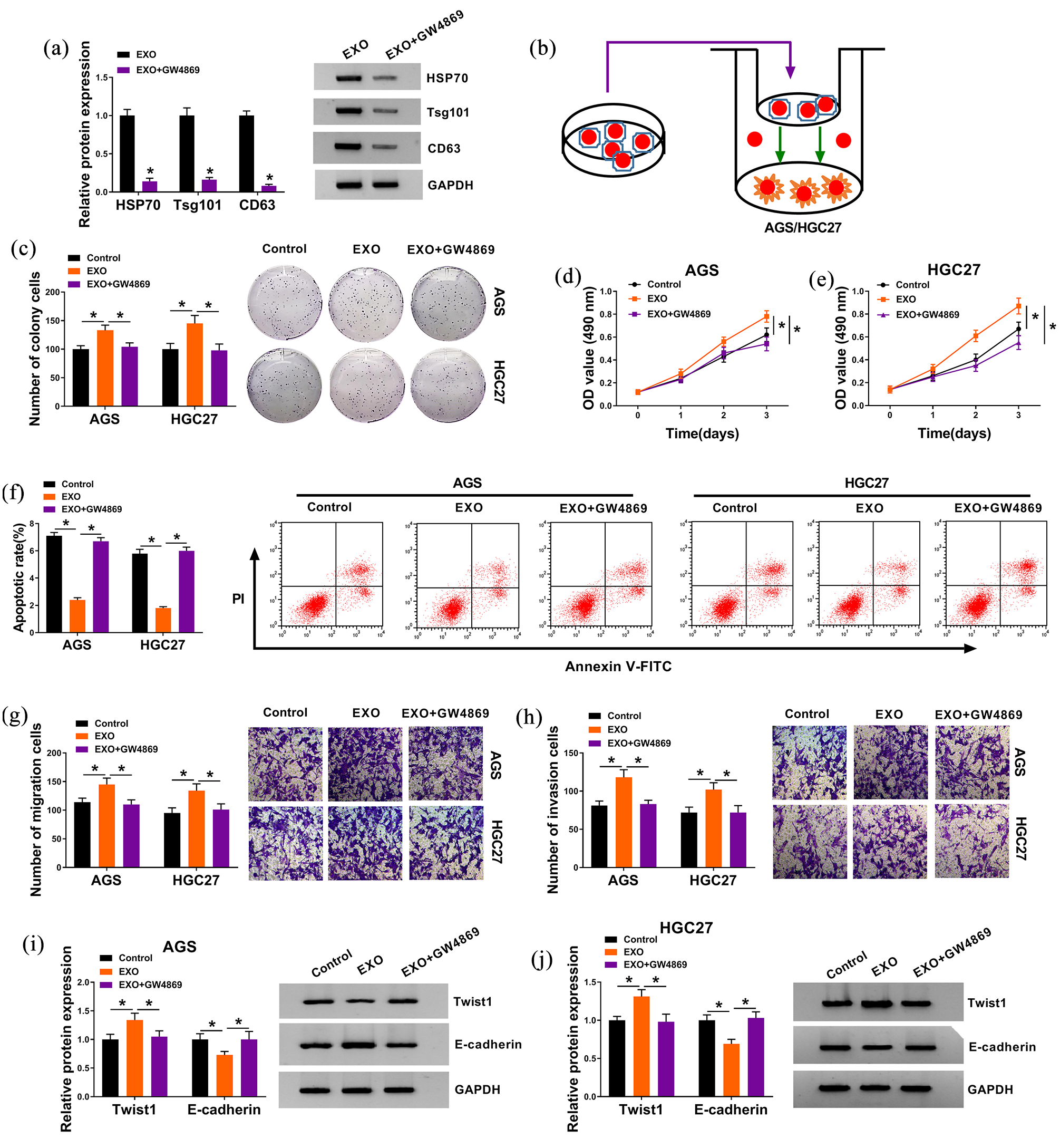
Exosomes derived from GC cells promoted cell proliferation, metastasis and inhibited apoptosis of GC cells. (a) Western blot assay for exosome biomarkers (HSP70, Tsg101, and CD63) in GC cells treated with EXO or EXO + GW4869. (b) AGS and HGC27 cells were disposed with PBS (control), EXO or EXO + GW4869 (exosome inhibitor). (c) Colony formation assay for the colony formation ability of treated cells. (d) and (e) MTT assay for the viability of treated cells. (f) Flow cytometry assay for the apoptosis of treated cells. (g) and (h) Transwell assay for the migration and invasion capacities of treated cells. (i) and (j) Western blot assay for EMT biomarkers (Twist1 and E-cadherin) in treated cells. *P < 0.05.
Exosomal miR-122-5p could block tumorigenicity of GC in vitro
In order to investigate the molecular mechanism of exosomes derived from GC cells, we constructed GC cells with miR-122-5p overexpression, the transfection efficiency is displayed in Figure 3(a). We also discovered that miR-122-5p was upregulated in EXO derived from GC cells treated with miR-122-5p mimic, compared to cells introduced with miR-NC (Figure 3(b)). Prior to functional assays, AGS and HGC27 cells were disposed with EXO derived from GC cells transfected with miR-NC or miR-122-5p (Figure 3(d)). Furthermore, miR-122-5p expression in AGS and HGC27 cells treated with EXO was gradually increased with the accumulation of processing time (Figure 3(c)). Colony formation and MTT assays indicated that gain of miR-122-5p suppressed proliferation of AGS and HGC27 cells (Figure 3(e) to (g)). In addition, overexpression of miR-122-5p apparently promoted cell apoptosis (Figure 3(h)), while hindered metastasis of AGS and HGC27 cells (Figure 3(i) to (l)). Collectively, exosomal miR-122-5p distinctly impeded development of gastric cancer in vitro.
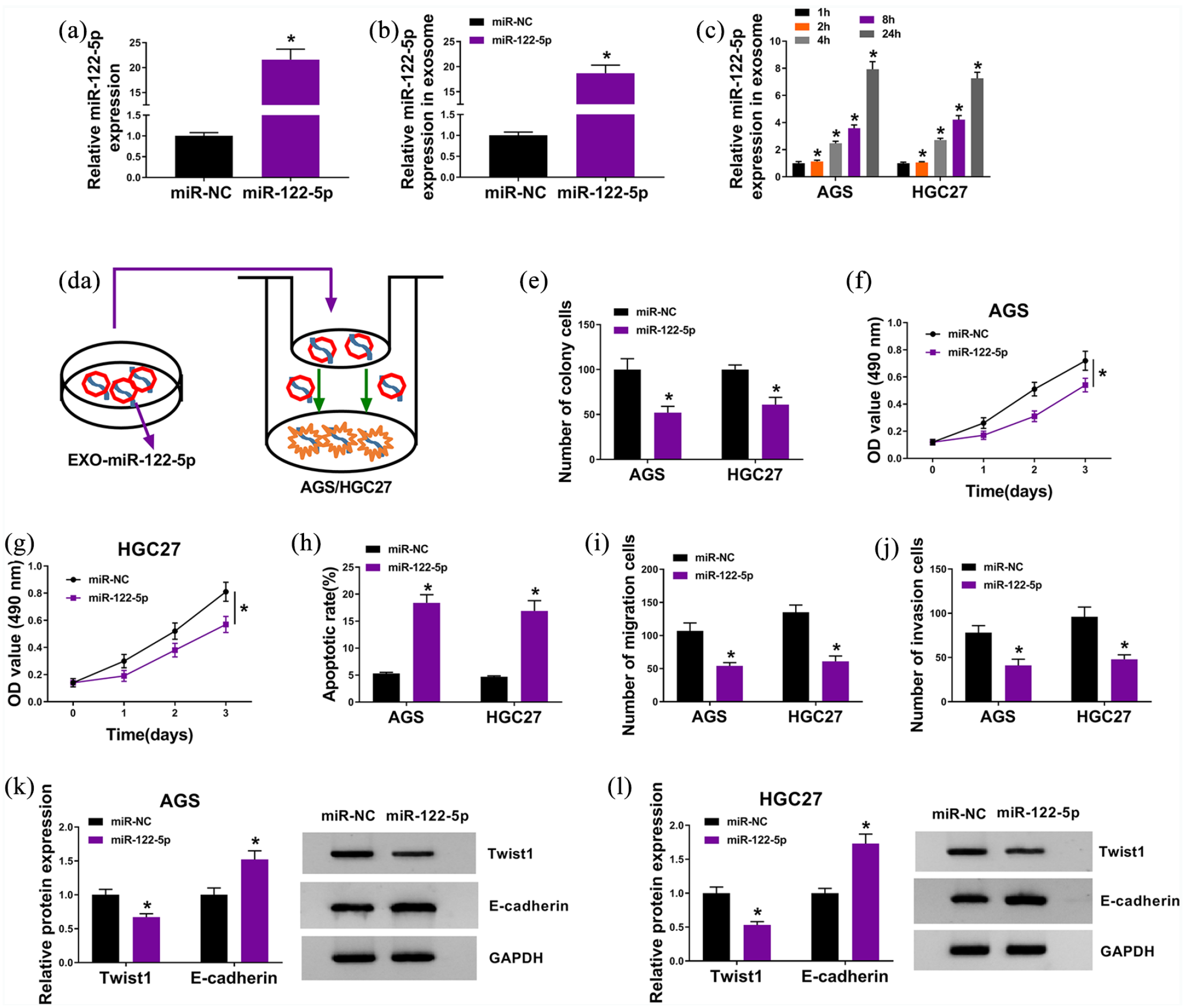
Exosomal miR-122-5p could block tumorigenicity of GC in vitro. (a) qRT-PCR assay for expression level of miR-122-5p in GC cells transfected with miR-122-5p or miR-NC. (b) qRT-PCR assay for expression level of miR-122-5p in EXO derived from GC cells treated with miR-122-5p or miR-NC. (c) qRT-PCR assay for expression level of miR-122-5p in cells treated with EXO for 1 h, 2 h, 4 h, 8 h, or 24 h. (d) AGS and HGC27 cells were disposed with EXO derived from GC cells transfected with miR-NC or miR-122-5p. (e) Colony formation assay for the colony formation ability of treated cells. (f) and (g) MTT assay for the viability of treated cells. (h) Flow cytometry assay for the apoptosis of treated cells. (i) and (j) Transwell assay for the migration and invasion capacities of treated cells. (k) and (l) Western blot assay for EMT biomarkers (Twist1 and E-cadherin) in treated cells. *P < 0.05.
GIT1 was a downstream target of miR-122-5p
To clarify the mechanistic pathway of exosomal miR-122-5p in GC development, the target gene of miR-122-5p was searched using online tool miRDB, and GIT1 3′UTR was found to have binding sites to miR-122-5p (Figure 4(a)). We then performed dual-luciferase reporter assay to validate the interaction between miR-122-5p and GIT1. As shown in Figure 4(b) and (c), the upregulation of miR-122-5p effectively reduced the luciferase activity of GIT1-wt in AGS and HGC27 cells, in contrast to cells co-transfected with miR-NC; the luciferase activity of GIT1-mut was unchanged. Additionally, we found that GIT1 was downregulated in EXO derived from GC cells transfected with miR-122-5p, in reference to cells introduced with miR-NC (Figure 4(d) and (e)). Furthermore, GIT1 was upregulated in EXO-GC compared to that in EXO-HC (Figure 4(f) and (g)), as well as in AGS and HGC27 cells with respect to GES-1 cells (Figure 4(h) and (i)). In conclusion, miR-122-5p could target GIT1.
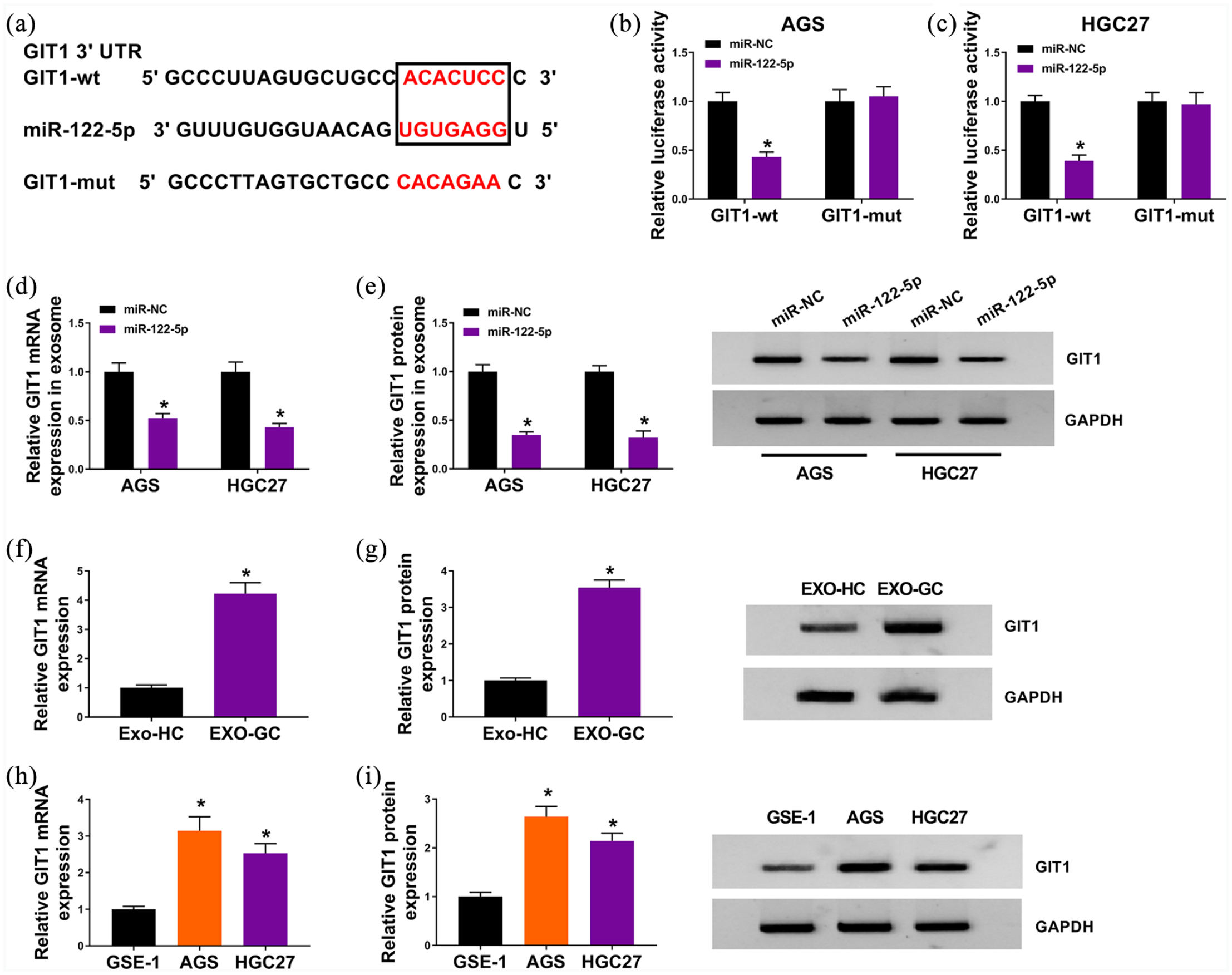
GIT1 was a downstream target of miR-122-5p. (a) The binding sited between miR-122-5p and GIT1 3’UTR, as well as the mutant. (b) and (c) Dual-luciferase reporter assay for the luciferase activity of GIT1-wt and GIT1-mut in AGS and HGC27 cells co-transfected with miR-NC or miR-122-5p. (d) and (e) qRT-PCR and western blot assays for GIT1 expression at mRNA and protein levels in EXO derived from GC cells transfected with miR-NC or miR-122-5p. (f) to (i) qRT-PCR and western blot assays for GIT1 expression at mRNA and protein levels in EXO-GC and EXO-HC, as well as in GES-1, AGS, HGC27, MKN45, and MKN74 cells. *P < 0.05.
Exosomal GIT1 knockdown could block GC development in vitro
For investigating the functional effects of GIT1 on GC development in vitro, GIT1 was specially silenced in GC cells, and knockdown efficiency was determined by qRT-PCR assay and western blot analysis (Figure 5 (a) and (b)). Moreover, the silencing of GIT1 also appeared in EXO derived from GC cells transfected with si-GIT1, in contrast to cells introduced with si-NC (Figure 5 (c) and (d)). Then, AGS and HGC27 cells were treated with EXO derived from GC cells transfected with si-NC or si-GIT1 (Figure 5(e)). Following functional assays uncovered that downregulation of GIT1 inhibited cell proliferation (Figure 5(f) to (h)) and metastasis (Figure 5(j) to (m)), while promoted cell apoptosis (Figure 5 (i)) in AGS and HGC27 cells. Thus, exosomal GIT1 knockdown impeded GC development in vitro.
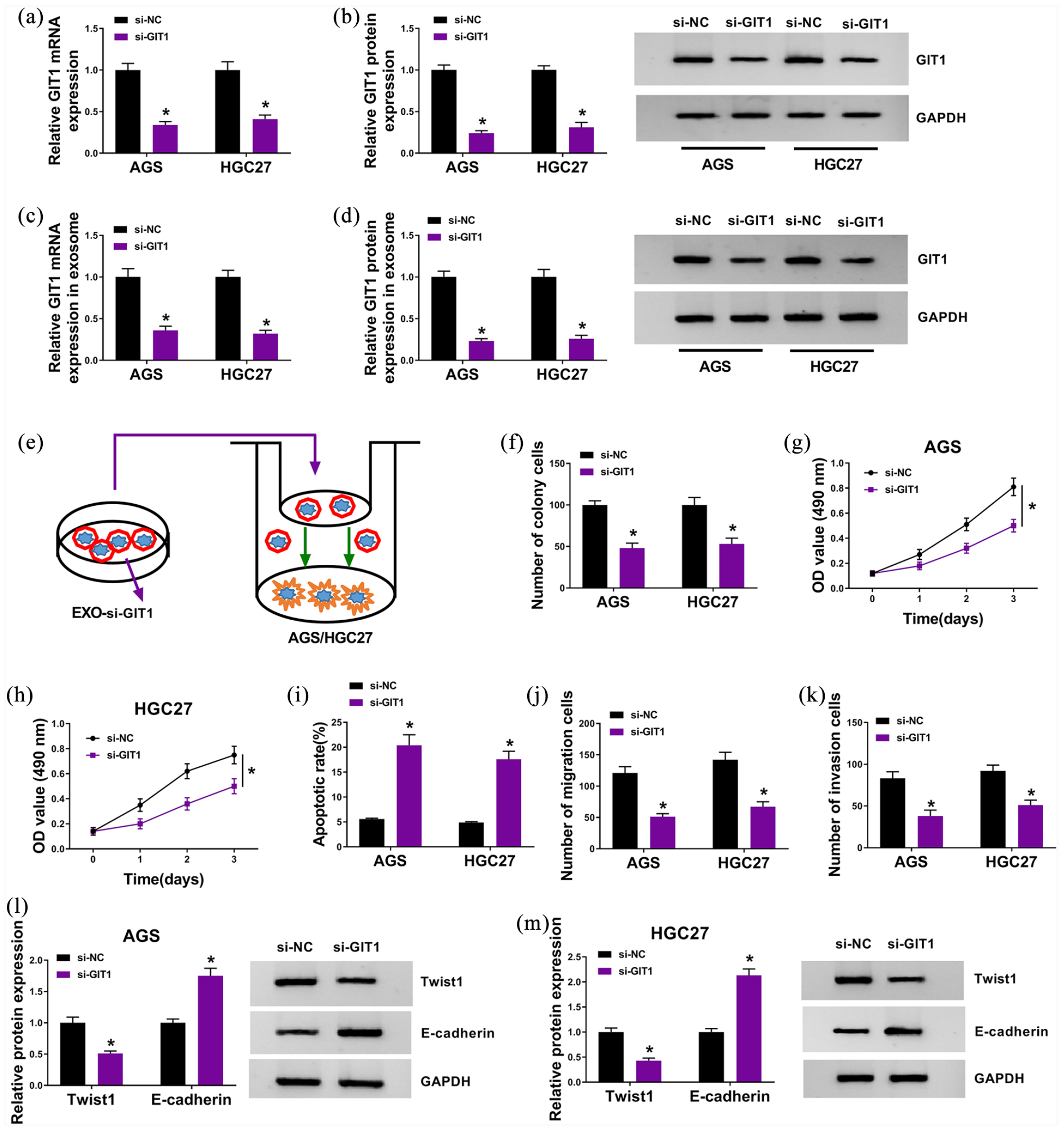
Exosomal GIT1 knockdown could block GC development in vitro. (a) and (b) qRT-PCR and western blot assays for GIT1 expression at mRNA and protein levels in GC cells transfected with si-NC or si-GIT1. (c) and (d) qRT-PCR and western blot assays for GIT1 expression at mRNA and protein levels in EXO derived from GC cells transfected with si-NC or si-GIT1. (e) AGS and HGC27 cells were disposed with EXO derived from GC cells transfected with si-NC or si-GIT1. (f) Colony formation assay for the colony formation ability of treated cells. (g) and (h) MTT assay for the viability of treated cells. (i) Flow cytometry assay for the apoptosis of treated cells. (j) and (k) Transwell assay for the migration and invasion capacities of treated cells. (l) and (m) Western blot assay for EMT biomarkers (Twist1 and E-cadherin) in treated cells. *P < 0.05.
Exosomal miR-122-5p suppressed GC development by downregulating GIT1
Having known that miR-122-5p could target GIT1, we then conducted rescue experiments to explore the role of GIT1 in exosomal miR-122-5p-mediated cellular behaviors of GC cells. As shown in Supplemental Figure 1(a) and (b), gain of miR-122-5p greatly inhibited the mRNA and protein levels of GIT1 in exosomes derived from AGS and HGC27 cells, but addition of GIT1 overturned the inhibitory effects. Furthermore, overexpressed miR-122-5p-mediated the declined clonogenicity (Supplemental Figure 1(c), cell viability (Supplemental Figure 1(d) and (e), metastasis (Supplemental Figure 1(f) and (g) and (i) and (j), as well as the elevated apoptotic rate (Supplemental Figure 1(h)) of AGS and HGC27 cells were all attenuated by additional GIT1. Collectively, exosomal miR-122-5p suppressed GC development by reducing GIT1 expression.
Exosomal miR-122-5p hampered tumor growth of GC in vivo
The function of exosomal miR-122-5p in tumorigenicity of GC in vitro was determined by the mouse xenograft model. AGS cells stably expressing miR-122-5p or miR-NC were subcutaneously injected into BALB/c nude mice (n=5). The volume of generated tumors was measured once a week. As depicted in Supplemental Figure 2(a), the volume of xenograft tumors in miR-122-5p group was smaller than that in miR-NC group. At 4 weeks post injection, generated tumors were resected and weighed. Enforced expression of miR-122-5p also notably reduced the weight of generated tumors (Supplemental Figure 2(b)). Additionally, higher expression of intracellular miR-122-5p (Supplemental Figure 2(c)) and lower expression of GIT1 (Supplemental Figure 2(d) and (e)) were discovered in miR-122-5p group, when compared with the miR-NC group. Herewith, exosomal miR-122-5p could hinder tumor growth of GC in vivo.
Discussion
Former reports have revealed the importance and molecular mechanisms of exosomal miRNAs in diverse human malignancies. 12 In our project, we found that exosomal miR-122-5p was downregulated in the serum of GC patients and GC cells. Furthermore, exosomal miR-122-5p acted as a tumor suppressor in GC by targeting GIT1.
Exosomes are extracellular vesicles, exerting vital roles in transferring intercellular signal networks, and intracellular fragment, such as DNAs, proteins, RNAs (coding and noncoding), etc.13, 14 Of these, miRNAs are a group of well-studied noncoding RNAs, with regulatory functions in many biological processes through repressing the translation and stability of mRNAs. 15 Previous research papers have alleged that exosomal miRNAs are closely related to tumorigenesis, metastasis, and drug resistance of different human cancers, serving as tumor suppressors or promoters.16, 17 Nowadays, accumulating researchers focus on exosomal miRNAs associated with GC. For example, exosomal miR-1290 was upregulated in GC patients and cell lines, and could contribute to proliferation, migration, and invasion of GC cells via downregulating NKD1, a direct target of miR-1290. 18 MiR-27a was obviously upregulated in exosomes derived from GC cells, and was confirmed to expedite proliferation and motility, which could act as a potential therapy target. 19
When it comes to miR-122-5p, it was a down-regulated miRNA in GC, gallbladder carcinoma, and bladder cancer. 9 Moreover, in breast cancer, miR-122-5p exerted suppressive function by targeting ADAM10. 20 Bai et al. 21 pointed out that miR-122-5p was downregulated in primary hepatocellular carcinomas (HCCs) and could repress tumorigenic properties of HCC cells. In our study, downregulation of miR-122-5p in GC cells and EXO-HC was disclosed, which was similar to previous reports.9, 22 A functional assay then revealed that the introduction of miR-122-5p notably suppressed GC progression in vitro and in vivo. Thus, exosomal miR-122-5p served as a GC inhibitor, and has the potential to be a therapeutic target for GC patients.
We then explored the molecular mechanism by which exosomal miR-122-5p exerted its anti-proliferation and anti-metastasis roles. As we all know, miRNAs could trigger the post-transcriptional silencing of target mRNAs to block the functions of targeted genes. 23 Additionally, the miRNA-target mRNA interactions and their regulatory networks widely existed in tumorigenesis and the development of human cancers. 24 In this study, GIT1 was shown to be a target of miR-122-5p. GIT1 is a member of PIX-GIT1-Paxillin protein complex, participating in the regulation of cell migration in diverse cell types, such as neurons, fibroblasts, immune cells, and endothelial cells.25, 26 GIT1 was up-regulated in oral squamous cell carcinoma, with the potential to be a biomarker for tumor prognosis. 27 In addition, the upregulation of GIT1 was closely implicated with the poor prognosis of medullary thyroid carcinoma patients. 28 GIT1 could facilitate the metastasis of non-small cell lung cancer cells via provoking the activity of Rac1/Cdc42. 26 Overexpressed expression of GIT1 was also observed in breast cancer, and miR-149 suppressed breast cancer progression in vitro and in vivo via targeting GIT1. 29 To sum up, GIT1 might exert a tumor-promoting role in human cancers. In concordance with a previous report, 11 we also found the upregulation of GIT1 in GC cells. Moreover, the silencing of GIT1 triggered GC cell proliferation and metastasis inhibition. Also, overexpression of GIT1 could undermine exosomal miR-122-5p-mediated inhibitory impact on GC cell proliferation and metastasis. Taken together, exosomal miR-122-5p functioned as GC suppressor by downregulating GIT1.
Collectively, we found that exosomal miR-122-5p was downregulated in the serum of GC patients and GC cells. Also, exosomal miR-122-5p remarkably inhibited proliferation, migration, and invasion of GC cells in vitro, as well as tumor growth in vivo by targeting GIT1. Our data are the first to prove the tumor-suppressive role of exosomal miR-122-5p in GC development, and developed a potential molecular mechanism, enhancing our comprehension about GC tumorigenesis and progression.
Supplemental Material
sj-pdf-1-jbm-10.1177_1724600821990677 – Supplemental material for Exosomal miR-122-5p inhibits tumorigenicity of gastric cancer by downregulating GIT1
Supplemental material, sj-pdf-1-jbm-10.1177_1724600821990677 for Exosomal miR-122-5p inhibits tumorigenicity of gastric cancer by downregulating GIT1 by Yigang Jiao, Li Zhang, Jun Li, Yuqi He, Xin Zhang and Jingzhe Li in The International Journal of Biological Markers
Footnotes
Acknowledgements
None
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Ethical approval
The research related to human use has been complied with all the relevant national regulations, and has been approved by the Ethics Committee of The First People’s Hospital of Lanzhou City.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
