Abstract
BACKGROUND:
Long non-coding RNAs (lncRNAs) play important roles in cancer development, yet their roles in renal carcinoma remain unclear.
OBJECTIVE:
We performed this study in order to investigate the expression and roles of lncRNAs in renal cell carcinoma.
METHODS:
In this study, we investigated the expression of lncRNAs in renal cell carcinoma through microarray analysis. Quantitative real-time PCR was performed to measure the expression of lncRNAs. Gain- or loss-of-function experiments were performed to investigate the roles of lncRNAs in cell proliferation and apoptosis. RNA pull-down and western blotting were performed to explore the underlying mechanism.
RESULTS:
The microarray analysis identified an upregulated lncRNA MIR4435-1HG in renal carcinoma. The expression level of MIR4435-1HG was correlated with TNM stage, tumor size, and Fuhrman grade. High expression of MIR4435-1HG indicated poor prognosis. MIR4435-1HG knockdown inhibited cell proliferation, and suppressed the migrating and invasive capacity of renal carcinoma cells. RNA pull-down followed by mass spectrometry revealed an interaction between MIR4435-1HG and pyruvate carboxylase, which was later corroborated by western blotting.
CONCLUSIONS:
MIR4435-1HG plays a critical role in the oncogenesis of renal cell carcinoma and may serve as a potential biomarker for renal cell carcinoma.
Introduction
Renal cell carcinoma (RCC) is one of the most deadliest cancers in the world, accounting for 3% of all cancer [1]. Clear cell renal cell carcinoma (ccRCC) accounts for approximately 70% of all RCCs [2]. Some patients already have metastasis at the time of diagnosis. Other patients are eligible for surgery, but recurrence and metastasis of the disease often occur [3]. However, renal cancer is characterized by a poor response to conventional chemotherapy and radiation therapy [4]. Although targeted therapies based on tyrosine kinases and the mammalian target of rapamycin (mTOR) have achieved certain effects, there are still problems with their limited survival benefit in patients and their obvious side effects [5].
Although greater than 75% of the human genome is selectively transcribed, only a small portion of transcripts are eventually translated into proteins. The transcripts that lack protein coding capacity are called non-coding RNAs, and they are further classified based on their length [6]. Long non-coding RNA (lncRNA) is defined as RNA of greater than 200 nucleotides in length, but without protein encoding ability [7]. Numerous studies have now demonstrated the involvement of lncRNAs in key life processes. LncRNAs have been shown to be involved in a variety of tumor pathologies [8]. LncRNAs may become new biomarkers for cancers, which may be helpful for diagnosis, prognosis prediction and even selecting treatment strategies for cancer patients [9]. In addition, for existing targeted therapy or immunotherapy, targeting lncRNAs may increase drug sensitivity or reverse drug resistance [10]. Since the underlying molecular mechanisms regulating RCC progression have not been completely elucidated, exploring the role of lncRNAs in RCC may help explain the mechanism of RCC and even provide alternative therapeutic targets.
In this study, we identified lncRNA MIR4435-1HG which has not been previously recognized in renal carcinoma before. Afterwards, the role of MIR4435-1HG was explored in cell lines and animal studies. This study aimed to determine the effects of MIR4435-1HG and its molecular mechanism, and provide a theoretical basis for MIR4435-1HG as a new biomarker for renal cancer.
Materials and methods
Patients’ samples and microarray analysis
This study was approved by The Ethics Committee of Ningbo First Hospital, and First Affiliated Hospital, Soochow University. All experiments were performed in accordance with the provisions of the Declaration of Helsinki. The experiments were undertaken with the understanding and written consent of each subject. A total of 118 clear cell RCC cases were collected from Ningbo First Hospital and First Affiliated Hospital, Soochow University. These tissues were freshly frozen in liquid nitrogen immediately after surgery and subsequently stored at
Cell culture
Human kidney proximal tubular cell line (HK-2) and RCC cell lines 786-O, OSRC-2 were purchased from the Chinese Academy of Sciences (Shanghai, China). HK-2 and 786-O cells were cultured in DMEM (Life Technologies, Carlsbad, CA, USA) supplemented with 10% fetal bovine serum (FBS) (Life Technologies, Carlsbad, CA, USA). OSRC-2cells were cultured in RPMI-1640 medium (Life Technologies, Carlsbad, CA, USA) supplemented with 10% FBS.
RNA extraction and qRT-PCR
Total RNAs were extracted from tissues and cells using TRIzol reagent (Invitrogen, Karlsruhe, Germany) in accordance with the manufacturer’s protocol. CDNA was synthesized by reverse transcription (RT) using random primers and the GoScript RT system (Promega, Madison, WI, USA). The real-time quantitative reverse transcription-polymerase chain reaction (qRT-PCR) was conducted using the GoTaq qPCR Master Mix system (Promega, Madison, WI, USA) on ABI Stepone Plus real-time PCR system (ABI, MA, USA) following the manufacturer’s instructions. The primer for lncRNA was synthesized by Xiangyin Biological Co. Ltd. (Hangzhou, China) as listed in Table 1.
The primer sequence
The primer sequence
Tissues and cells were collected and lysed with cell lysis buffer (Beyotime, Haimen China) for western blotting. The proteins (30
Virus construction and cell transfection
Short hairpin RNA (shRNA) lentivirus was designed by Genechem Co. Ltd. (Shanghai, China) to target human MIR4435-1HG. The vector used for MIR4435-1HG knockdown was hU6-MCS-Ubiquitin-EGFP-IRES-puromycin and it’s named Lv-sh MIR4435-1HG. Negative control was named Lv-sh NC. To constitutively overexpress MIR4435-1HG in cells, recombinant lentivirus vector containing the full-length sequence of MIR4435-1HG was constructed by Genechem. The vector used for MIR4435-1HG overexpression was Ubi-MCS-SV40-EGFP-IRES-puromycin. The empty virus was used as control. Cells were then infected with lentivirus following the manufacturer’s instructions, and PCR was used to validate the efficiency of MIR4435-1HG knockdown and overexpression.
Cell viability assay and colony formation assay
Cell proliferation was assessed using Cell Counting Kit-8 (CCK-8) (Dojindo Molecular Technologies, Kyushu, Japan), according to the manufacturer’s instructions. Cells were seeded into 96-well culture plates at a density of 6
Apoptosis and cell cycle analysis
For cell cycle analysis, cells were harvested and fixed with 75% ethanol at 4
Cell migration and invasion assays
For the in vitro migration and invasion assays, 5
Fluorescent in situ hybridization (FISH)
To detect the subcellular localization of MIR4435-1HG, FISH was performed by Genecreate Biological Engineering Co. Ltd. (Wuhan, China). Briefly, oligonucleotide probes targeting MIR4435-1HG, 6u, and 18s were designed. OSRC-2 and 786-0 cells were treated in hybridisation buffer with probes at 42
RNA pull-down and mass spectrometry
RNA pull-down and mass spectrometry was performed by Genecreate biological engineering Co. Ltd. (Wuhan, China). Briefly, T7 promoter tagged oligos were produced by PCR. Biotin-labeled MIR4435-1HG RNA (GFP RNA as control) was in vitro transcribed with the Biotin RNA labeling mix (Roche, Branford, CT, USA) and T7 RNA polymerase (Roche, Basel, Switzerland) and purified with RNeasy MiniKit (Qiagen, Germantown, MD, USA). RNAs were incubated with cell extract from 786-0 cells in the presence of anti-RNase, protease inhibitor cocktail. Proteins pulled down by biotinylated RNA were subjected to mass spectrometry. The peptides were analyzed by LC-MS/MS (AB Sciex TripleTOF 5600-plus) at a scan range of m/z 400–1500. The files were analyzed using the Proteinpilot platform. MS/MS spectra of protein identified were retrieved from the protein database (huma-refseq-20140303-71465s.fasta, National Center of Biotechnology Information).
Animal studies
All procedures for animal experiments were approved by the Animal Use and Care Committee of Soochow University. SPF-class BALB/c nude mice were purchased from Shanghai SLAC Company. 786-0 cells transfected with Lv-sh MIR4435-1HG or Lv-sh NC were injected at 2
MIR4435-1HG was upregulated in renal carcinoma tissues and cell lines. a: LncRNAs microarray data of ccRCC samples compared with that of adjacent non-tumour samples is presented in a heat map. b: QRT-PCR analysis of MIR4435-1HG between ccRCC samples and paired adjacent non-tumour samples. c: QRT-PCR analysis of MIR4435-1HG in HK-2, 786-0, and OSRC-2 cell lines. d: Overall survival (OS) analysis. High MIR4435-1HG expression was associated with reduced OS. E: Recurrence free survival (RFS) analysis. High MIR4435-1HG expression was associated with reduced RFS. f: ROC curve analysis of the diagnostic performance of MIR4435-1HG. AUC 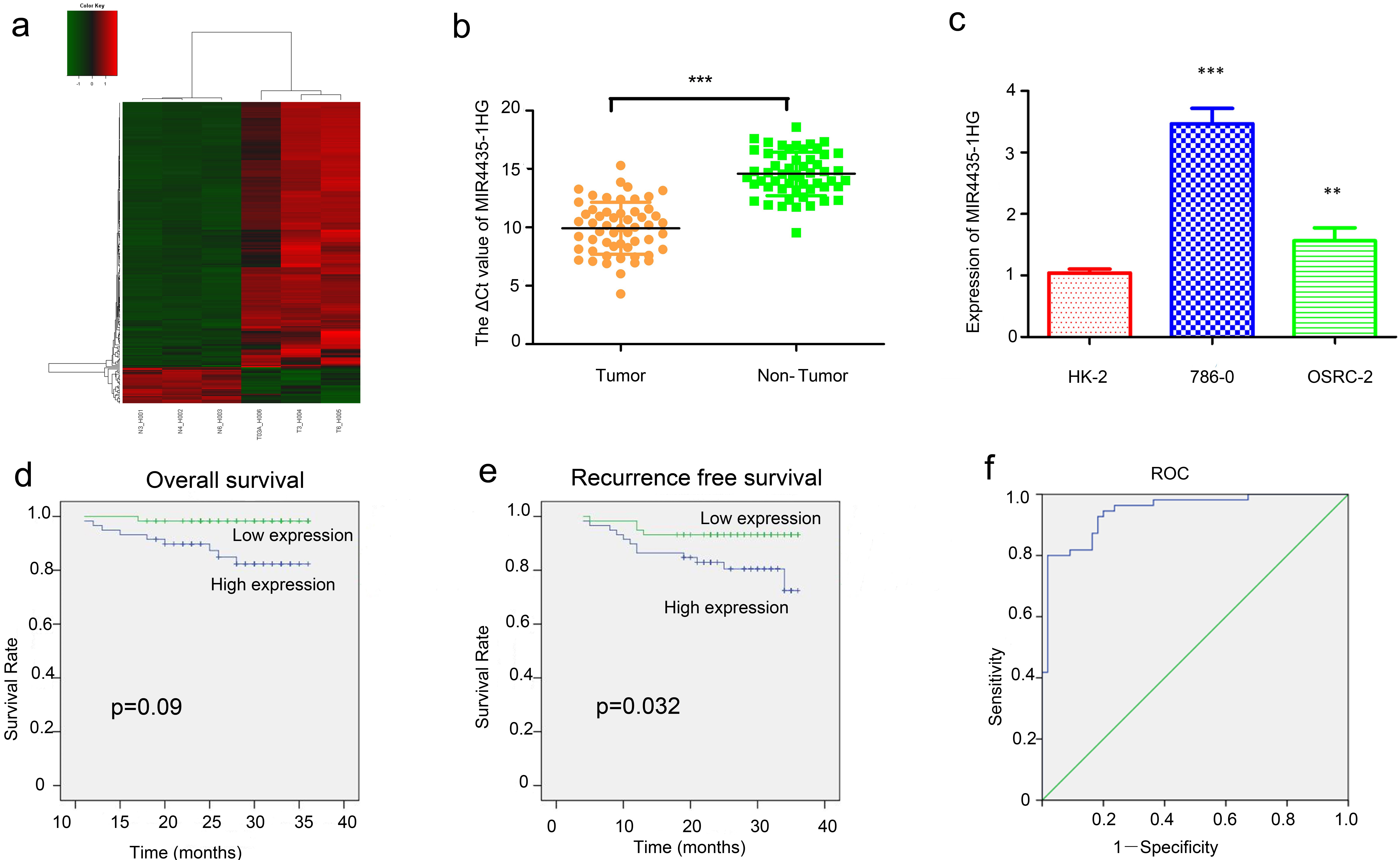
Data were analysed using SPSS version 16.0 (SPSS Inc., Chicago, IL, USA). Differences in lncRNA expression levels between renal cancer tissues and paracancerous tissues were calculated using the paired sample
Results
The expression level of MIR4435-1HG in renal cell carcinoma tissues and cell lines
A total of 118 cases of clear cell renal carcinoma cases were enrolled in this study, including 55 cases of radical nephrectomy and 63 cases of partial nephrectomy.
To identify lncRNAs essential for ccRCC oncogenesis, three ccRCC tissues and their adjacent nontumor tissues were subjected to a lncRNA expression microarray. Cluster analysis showed a clear distinction between ccRCC tissues and nontumor tissues (Fig. 1a). Among the deregulated lncRNAs, the expression of MIR4435-1HG (ENST00000439362) was found to be significantly higher in the three ccRCC tissue samples than in the adjacent nontomor tissue samples (
To further verify the findings of the microarray, we next examined the expression of lncRNA in paired ccRCC tissues and their adjacent nontumor tissues from 55 radical nephrectomy cases by qRT-PCR. The expression level of MIR4435-1HG in renal cancer tissues was much higher than that in adjacent tissues (
The association between MIR4435-1HG expression levels in tumor tissues and the clinicopathological features of renal cell carcinoma patients
The association between MIR4435-1HG expression levels in tumor tissues and the clinicopathological features of renal cell carcinoma patients
The median age of the 118 patients at diagnosis was 58.0 years (56.8
The effect of MIR4435-1HG on proliferation
To investigate the roles of MIR4435-1HG in renal carcinoma cells, we knocked down or overexpressed MIR4435-1HG by lentivirus infection. After transfection, the expression of MIR4435-1HG in the overexpression group was remarkably elevated compared with that in the control group (Fig. 2a). Two independent Lv-sh MIR4435-1HG were designed and transfected into cells to reduce MIR4435-1HG expression. The lentivirus (Lv-sh-2) that achieved better knockdown efficacy was chosen for the following experiments (Fig. 2b).
MIR4435-1HG knockdown inhibited the proliferation of renal cancer cells. a: Relative expression levels of MIR4435-1HG in virus transfected cells. MIR4435-1HG overexpression was achieved in virus transfected cells. b: Relative expression levels of MIR4435-1HG in virus transfected cells. Lv-sh-2 achieved better reduction efficacy than Lv-sh-1. c: Growth curves of cells transfected with virus. d: Colony formation assays of cells transfected with virus. *, 
The cell proliferation ability of MIR4435-1HG was detected by the CCK-8 method. Compared with the negative control (NC) group, the cell proliferation rate of the knockdown (KD) group was decreased. Compared with the NC group, the cell proliferation rate of the overexpression (OE) group increased (Fig. 2c). The results of colony formation experiments were consistent with the CCK-8 assay. After lentivirus infection, the number of cell colony formation in the KD group was significantly reduced as compared with that in the NC group, and the difference was statistically significant; compared with that in the NC group, the number of cell colony formation in the OE group was significantly increased, and the difference was statistically significant (Fig. 2d).
The results of cell cycle assay showed that the proportion of cells in G0/G1 phase of the KD group was higher than that of the NC group, and the proportion of cells in S phase was lower (
Cell cycle analysis and cell apoptosis in HK-2, 786-0 and OSRC-2 cells. a, b, c: Cell cycle analysis of HK-2, 786-0 and OSRC-2 cells. d, e, f: Cell apoptosis in HK-2, 786-0 and OSRC-2 cells. *, 
Transwell assay was performed to investigate the cell migration and invasive capacity. Regarding invasion, the number of invasive cells in the OE group was larger than that in the NC group; the number of invasive cells in the KD group was smaller than that in the NC group (
Cell invasion and migration. a and b: MIR4435-1HG inhibition inhibited tumor invasion and MIR4435-1HG overexpression promoted tumor invasion. c and d: MIR4435-1HG inhibition inhibited tumor migration and MIR4435-1HG overexpression promoted tumor migration. Scale bar 
The FISH results showed that 18S and U6 localized to the cytoplasm and nucleus respectively. MIR4435-1HG was expressed mainly in the cytoplasm (Fig. 5a and b). RNA Pulldown was further performed. In the silver staining analysis, the protein band did not show a significant difference in terms of protein amount. Thus, mass spectrometry was conducted and an interaction between MIR4435-1HG and proteins was revealed (Table 3). By western blotting, we explored whether MIR4435-1HG may influence the expression of those proteins. We found that there was a significant difference in the PC protein level in ccRCC cells with MIR4435-1HG depletion and those with overexpression (Fig. 5d).
Intracellular localization of MIR4435-1HG and RNA-protein interaction. a and b: Fluorescence in situ hybridization showed that 18S and U6 localized in cytoplasm and nucleus respectively. MIR4435-1HG was expressed mainly in cytoplasm. c: Identification of pyruvate carboxylase. Representative MS/MS spectra of the tryptic peptides from pyruvate carboxylase: TSTAPAASPNVR. The b and y ions were indicated with green and red colors, respectively. d: Expression of pyruvate carboxylase by western blotting. The level of pyruvate carboxylase was lower in MIR4435-1HG knockdown cells and higher in MIR4435-1HG overexpression cells as compared with negative control. NC, negative control; OE, overexpression; KD, knockdown.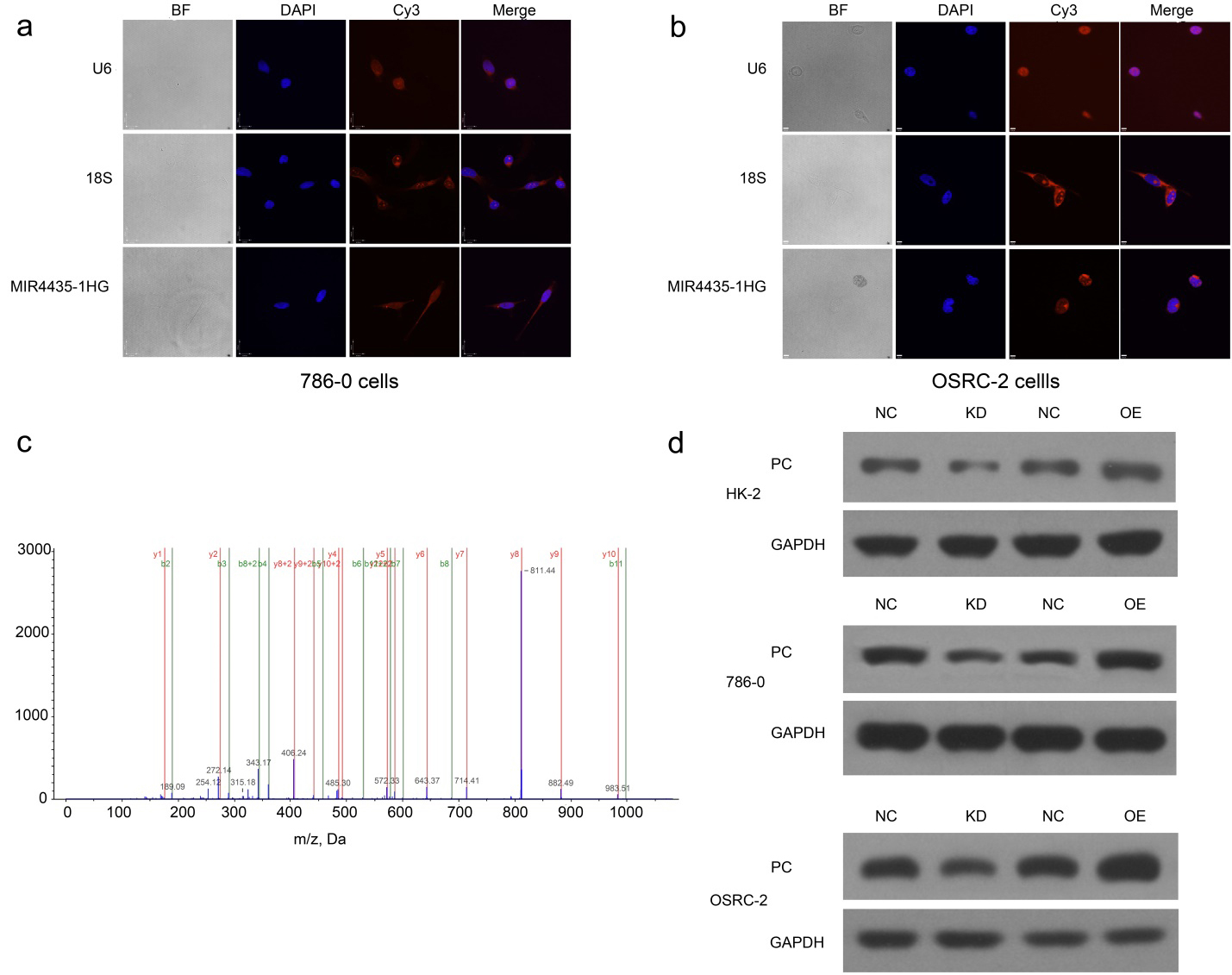
To examine the effect of MIR4435-1HG on RCC growth in vivo, we established a subcutaneous xenograft tumor model. The tumor of the KD group was smaller than that of the NC group on the second, third and fourth weeks (
Identification of proteins by mass spectrometry
Identification of proteins by mass spectrometry
Knockdown of MIR4435-1HG suppressed the tumor growth of RCC cells in vivo. a: The tumor size of the KD group was smaller than the tumor size of the NC group at the second, third and fourth weeks. b: Weight of nude mice was of no different between NC group and KD group statistically. c: The appearance of subcutaneous xenograft tumor before taken out of mice. d: The appearance of subcutaneous xenograft tumor after taken out of mice. e. The tumor weight of the KD group was lower than that of the NC group. f: The expression level of pyruvate carboxylase was lower in the KD group as compared with the NC group by western blotting. ***, 
RCC is the second leading cause of death associated with urological malignant neoplasms [11]. Late-stage ccRCC patients, especially metastatic ccRCC patients, have a poor prognosis and limited clinical therapeutic options at present [12]. Hence, it is of urgent demand to investigate the biological basis of ccRCC and identify novel targets for systemic therapy.
Long non-coding RNAs are transcribed by RNA polymerase II and play an important role in cell regulation, proliferation and apoptosis [13]. LncRNAs have been recently discovered and identified to have a significant impact on tumor biology [14, 15, 16]. In renal carcinoma, a group of lncRNAs were detected to be associated with tumor progression and metastasis [17, 18, 19]. However, the molecular mechanism by which lncRNAs influence renal carcinoma cells remains unclear.
In the present study, we found a lncRNA transcript, MIR4435-1HG, which had not been previously reported in renal cell carcinoma according to our knowledge. MIR4435-1HG was upregulated in ccRCC tissues through the lncRNA expression microarray. The survival analysis revealed that higher MIR4435-1HG expression was correlated with shorter survival time of patients. Additionally, analysis showed that higher expression of MIR4435-1HG was correlated with larger tumor size, later TNM stage, and higher Fuhrman grade. The ROC curve showed the potential value of MIR4435-1HG as a biomarker for diagnosis. These results suggested that MIR4435-1HG may be involved in the oncogenesis of RCC. MIR4435-1HG may serve as a novel biomarker for RCC prognosis.
We next explored the biological functions of MIR4435-1HG in renal cancer cells by using lentivirus-mediated gene silencing and overexpression. The results of cell counting and colony formation assays revealed that MIR4435-1HG knockdown led to a significant inhibition of the proliferation of cells, which was later confirmed by a subcutaneous xenograft tumor model. Apoptosis and cell cycle analysis partially explained the effects of MIR4435-1HG on cell proliferation. Yang et al. reported that MIR4435-1HG knockdown in lung cancer cells inhibited the formation of colonies, which might be associated with the induction of cell cycle arrest at G1 phase [20]. Min Fu et al. Reported that MIR4435-1HG knockdown resulted in G1 phase arrest and inhibited the migration and invasion of gastric cancer cells [21]. Our result is consistent with those reports and suggests that MIR4435-1HG may play an oncogenic role in renal carcinoma by regulating the cell cycle.
Protein – RNA interactions are important in the involvement of lncRNA in cellular processes [22]. In our study, FISH showed that MIR4435-1HG was primarily located in the cytoplasm, which indicates that MIR4435-1HG may function in the cytoplasm. RNA pull-down followed by MS showed that MIR4435-1HG may interact with PC. PC, which is located in mitochondria, is an anaplerotic enzyme that plays an essential role in various cellular metabolic pathways [23]. More recent studies also show that PC is strongly involved in tumorigenesis in several cancers including renal carcinoma, and that pyruvate carboxylation acts as a metabolic hub that feeds carbon skeletons of downstream metabolites of oxaloacetate into the biosynthesis of various cellular components [24]. In addition, the Krebs cycle is a source of energy that requires the replenishment of intermediates, which is partly accomplished via the carboxylation of pyruvate to oxaloacetate by PC [25]. Our experiments show that MIR4435-1HG positively regulates PC expression. This study suggests that MIR4435-1HG may perform its role in oncogenesis through PC-related metabolism regulation in renal carcinoma cells.
In summary, our study reveals that MIR4435-1HG is highly expressed in renal cell carcinoma. MIR4435-1HG plays a critical role in the oncogenesis of ccRCC by positively regulating PC expression and may represent a potential therapeutic target for ccRCC.
Footnotes
Acknowledgments
This work was supported by the Natural Science Foundation of Zhejiang Province (LY19H160013), the Natural Science Foundation of Ningbo City (2018A6 10295), as well as the Medicine and Hygiene Program of Zhejiang Province (2014KYB231, 2018KY686, 2018KY154). We thank Professor Junmin Guo and Guangbo Zhang for their kind guidance in study design. We thank Cheng Yang, Guang Yan, Cheng Zhou, Zhaohui Jiang, and Qi Ma for thier assistance in sample collection and instruction on the experiments. We also thank Yiwei Lin, Kai Yang and Xiao Wang for their previous support in tumorigenesis knowledge.
Conflict of interest
The authors have no conflict of interest to declare.
