Abstract
Lung cancer accounts for a large proportion of cancer-related deaths worldwide. Personalized therapeutic medicine based on the genetic characteristics of non-small cell lung cancer (NSCLC) is a promising field, and discovering clinically applicable biomarkers of NSCLC is required. LINC00472 is a long non-coding RNA and has been recently suggested to be a biomarker of NSCLC, but little is known of its mechanism in NSCLC. Thus, the current study was performed to document changes in gene expression after LINC00472 overexpression in NSCLC cells. As a result of cell viability and migration assay, LINC00472 downregulated cell survival, proliferation, and motility. Transcriptome sequencing analysis showed 3,782 genes expression were changed in LINC00472 overexpressing cells. Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis showed most genes were associated with intracellular metabolism. The PPP1R12B, RGS5, RBM5, RBL2, LDLR and PTPRM genes were upregulated by LINC00472 overexpression and these genes functioned as tumor suppressors in several cancers. In contrast, SPSB1, PCNA, CD24, CDK5, CDC25A, and EIF4EBP1 were downregulated by LINC00472, and they functioned as oncogenes in various cancers. Consequently, the function of LINC00472 in tumorigenesis might be related to changes in the expressions of other oncogenes and tumor suppressors.
Introduction
Lung cancer is a common cause of cancer death because during the early stage the disease is asymptomatic, and thus, it is usually detected at an advanced stage [1]. Lung cancer is traditionally classified as non-small cell lung cancer (NSCLC) or small cell lung cancer (SCLC), and NSCLC accounts for about 80% of patients [2]. Although most cases of SCLC are treated non-surgically, NSCLC is treated using surgery and adjuvant therapy [2]. The most common cause of lung cancer is smoking [3], and thus, preventing tobacco use is the most important factor in terms of overcoming NSCLC. Nevertheless, more aggressive, sophisticated treatment regimens are needed to completely eradicate the disease. The treatment of NSCLC has undergone many changes, including advances in targeted therapies and immunotherapy. Furthermore, personalized therapeutic medicine shows great promise for the treatment of NSCLC [4]. Therefore, it is necessary to discover the applicable biomarkers and to profile gene expression changes in NSCLC.
LncRNAs are generally non-coding RNAs longer than 200 nucleotides [5]. In general, although the intracellular expressions of lncRNAs are low [6], lncRNA is importantly involved in various cellular processes including transcription, splicing, translation, cell cycle, and heat shock response [7, 8]. In addition, lncRNAs are known to be involved in cancer progression [9, 10] and in the pathogeneses of several other diseases [11]. Abnormal lncRNA expression is associated with tumor formation and can be used as a biomarker to predict cancer prognosis [12]. Interestingly, some lncRNAs function as oncogenes, whereas others function as tumor suppressors [13]. For example, the oncogenic lncRNA, lncRNA PTAR directly targets mir-101 to promote NSCLC cell growth [14], but lncRNA FENDRR acts as a tumor suppressor by binding to oncogenic mir-761 and inhibiting NSCLC aggressiveness [15].
One type of lncRNA, LINC00472, is a long intergenic non-protein coding RNA 472 located on human chromosome 6q13 and has been recently reported to act as a tumor suppressor in various cancers. Meta-analyses showed LINC00472 lowered the risk of tumor recurrence in breast cancer patients with a grade 2 tumor [16] and negatively controlled the cell cycle, metastasis, and proliferation in colorectal cancer [17]. The tumor inhibitory effect of LINC00472 has also been reported in NSCLC. LINC00472 inhibited cell invasion and migration of NSCLC cells and reduced tumor growth in nude mice [18]. However, despite evidence that LINC00472 acts as a tumor suppressor, few studies have investigated LINC00472-related genes and signaling pathways in NSCLC.
In this study, after overexpressing LINC00472 in the NCI-H838 cells (an NSCLC cell line), Transcriptome Sequencing analysis (total RNA-seq) was used to profile LINC00472-induced gene expressional changes to provide important data for future investigations on the LINC00472 signaling pathway in NSCLC.
Materials and methods
Cell culture and transfection
Cell culture and transfection were performed as previously described [19]. NCI-H838 cells (a human NSCLC cell line) were obtained from the American Type Culture Collection (ATCC, Manassas, VA, USA), authenticated, and then maintained in early passages for no more than 6 months after receipt. Cells were grown in RPMI-1640 (HyClone, Marlborough, MA, USA) supplemented with 10% FBS (Hyclone), 100 U/ml penicillin and 100
Cells were transiently transfected with a plasmid containing a full-length LINC00472 construct (pCDH-LINC00472) and a control plasmid (pCDH empty vector) using Lipofectamine 2000 (Invitrogen, Waltham, MA, USA), incubated for 24 h, and subjected to cell survival and metastasis assays.
Total RNA preparations
After transient transfection, total RNA was isolated from cells using TRIzol (Invitrogen) [20]. RNA qualities and concentrations were assessed using OD 260/280 ratios using a Nanodrop 2000 (Thermo Scientific, Waltham, MA, USA). Extracted RNAs were used for reverse transcription PCR and forwarded to Macrogen (Seoul, Republic of Korea) for transcriptome sequencing.
Reverse transcription PCR
A Total RNA isolated from cells was synthesized as cDNA using the MMLV Reverse Transcriptase system (Bioneer, Daejeon, Republic of Korea). The temperature conditions for reverse transcription (RT) were 60 min at 37
Cell viability assay
A Cell Counting Kit-8 (Dojindo, Kumamoto, Japan) was used to measure cell survival and proliferation. A cell suspension of NCI-H838 cells (5,000 cells/100
In addition, trypan blue solution (Gibco) was used to measure cell viability [20]. Briefly, cells (1
Colony-forming assay
For colony-forming assays [21], NCI-H838 cells were plated at a density of 600 cells per 35 mm dish. After 24 h, cells were transfected with pCDH-LINC00472 or pCDH empty vector, incubated at 37
Wound healing assay
For wound healing assays [22], cells were cultured until 70% confluent in growth medium. Cell monolayers were then scratched with a 200
Cell migration assay
The Radius™ 96-well Cell Migration Assay (Cell Biolabs, Inc., San Diego, CA, USA) was used to measure cell migration ability. After wells had been pretreated with Radius™ Pretreatment Solution and Radius™ Wash Solution, 3.0
Transcriptome sequencing
Total RNA concentrations were calculated using Quant-IT RiboGreen (Invitrogen). Samples were run on the TapeStation RNA screentape (Agilent, Santa Clara, CA, USA) to assess total RNA integrities. Only high-quality RNA preparations with RIN values of
LINC00472 acts as a tumor suppressor in various cancers
LINC00472 acts as a tumor suppressor in various cancers
Raw reads were preprocessed from the sequencer to remove low quality and adapter sequences before analysis and processed reads were aligned to Homo sapiens (hg19) using HISAT v2.1.0 [23]. HISAT utilizes two index types for alignment, that is, a global, whole-genome index and tens of thousands of small local indexes. These two index types’ are constructed using the same BWT (Burrows-Wheeler transform)/a graph FM index (GFM) as Bowtie2. Because of the use of these efficient data structures and algorithms, HISAT generates spliced alignments several times faster than the widely used Bowtie and BWA. The reference genome sequence of Homo sapiens (hg19) and annotation data were downloaded from the NCBI, and transcript assembly of known transcripts, novel transcripts, and alternative splicing transcripts was processed using StringTie v1.3.4d [24, 25]. Base on the results obtained, expressional abundances of transcripts and genes were calculated as read counts or FPKM values (Fragments Per Kilobase of exon per Million fragments mapped) per sample. Expression profiles were for additional analysis, for example, to Differentially Expressed Gene (DEG). In samples obtained using different conditions, differentially expressed genes or transcripts were filtered by statistical hypothesis testing.
Statistical analysis was performed to identify differentially expressed genes using estimates of gene abundances in samples. Relative gene abundances were measured in FPKM using StringTie. Genes with one more that zeroed FPKM values in samples were excluded. To facilitate log2 transformation, 1 was added to the FPKM values of filtered genes. Filtered data were log2-transformed and subjected to quantile normalization. The statistical significances of differential expression data were determined using fold changes. The sequencing results have been deposited in the Gene Expression Omnibus database (GEO Series accession number GSE164230). For DEG sets, hierarchical clustering analysis was performed using complete linkage and Euclidean distance as measures of similarity. Gene-enrichment analysis and KEGG pathway analysis of DEGs were performed using Gene Ontology (
Statistical analysis
Results are presented as the mean
Effects of LINC00472 overexpression on NSCLC cell viability and metastasis. (A) Expression of LINC00472 in NCI-H838 after transient transfection. Relative intensity 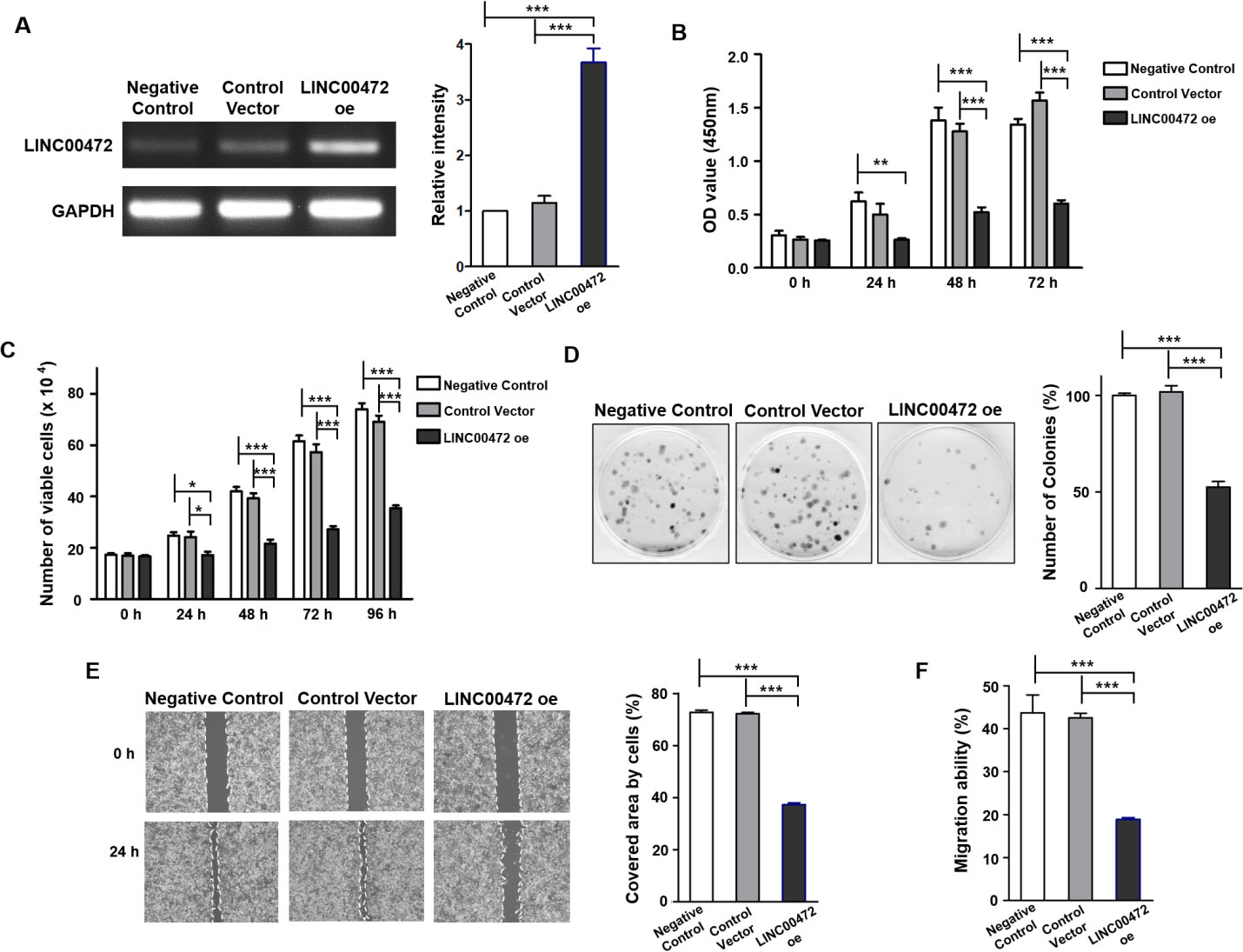
Effects of LINC00472 on the survival and invasiveness of NSCLC cells
LINC00472 has been demonstrated to act as a tumor suppressor in several cancers including breast cancer, colorectal cancer, and lung cancer (Table 1). In bladder carcinoma, LINC00472 functioned as a tumor suppressor by upregulating the expression of programmed cell death protein 4 (PDCD4) [26], and in colorectal cancer cells, LINC00472 directly targeted mir-196a to increase the expression of PDCD4, and thereby, inhibited proliferation and tumor growth [27]. In hepatocellular carcinoma, LINC00472 inhibited cell proliferation, migration, and invasion by upregulating PDCD4 expression by sponging mir93-5p [28]. In a study of osteosarcoma tumorigenesis, LINC00472 inhibited cancer cell proliferation by upregulating FOXO1 via the downregulation of mir-300 [29]. In breast cancer, LINC00472 was highly expressed in estrogen receptor (ER)-positive cells, and the presence of these cells was associated positively with survival and prognosis as compared with ER-negative cells, and LINC00472 was found to act as a tumor suppressor by inhibiting phosphorylation of nuclear factor
Sequence data quality check results
Sequence data quality check results
Total read bases: Total number of sequenced bases after trimming Total reads: Total number of reads after trimming Q20 (%): The proportion of bases with a quality of Phred quality score 20 or higher Q30 (%): The proportion of bases with a quality of Phred quality score 30 or higher.
To confirm that LINC00472 acts as a tumor suppressor in NSCLC cells, CCK-8, trypan blue, cell migration, wound healing, and colony-forming assays were performed on LINC00472-overexpressing NSCLC cells before proceeding with total RNA-seq. Successful transfection of LINC00472 in NCI-H838 was confirmed by PCR (Fig. 1A). CCK-8 assays (Fig. 1B) showed the OD of cells overexpressing LINC00472 (LINC00472-oe) was lower than that of the negative or vector controls (
After transfecting NCI-H838 cells with pCDH-LINC00472 or pCDH empty vector, total RNA-seq was performed to investigate mRNA profile differences. The results of the quality check of sequencing data are presented in Table 2. Cells transfected with control vector had 132,325,002 total reads whereas LINC00472-oe cells had 117,742,244 total reads. Since Phred quality scores (Q20 and Q30), which represent the accuracy of nucleotide sequence analysis, were over 90%, the sequencing data was deemed reliable and used for further analysis. The sequencing data was deposited in the Gene Expression Omnibus database (GEO Series accession number GSE164230).
After performing total RNA-seq, only genes whose expressional values differed by more than 2-fold between LINC00472-oe cells and controls (
Differentially expressed genes as determined by transcriptome sequencing and the results of GO and KEGG enrichment analyses. The sequencing results were registered in the GEO (GSE164230). (A) 1,922 genes were upregulated by 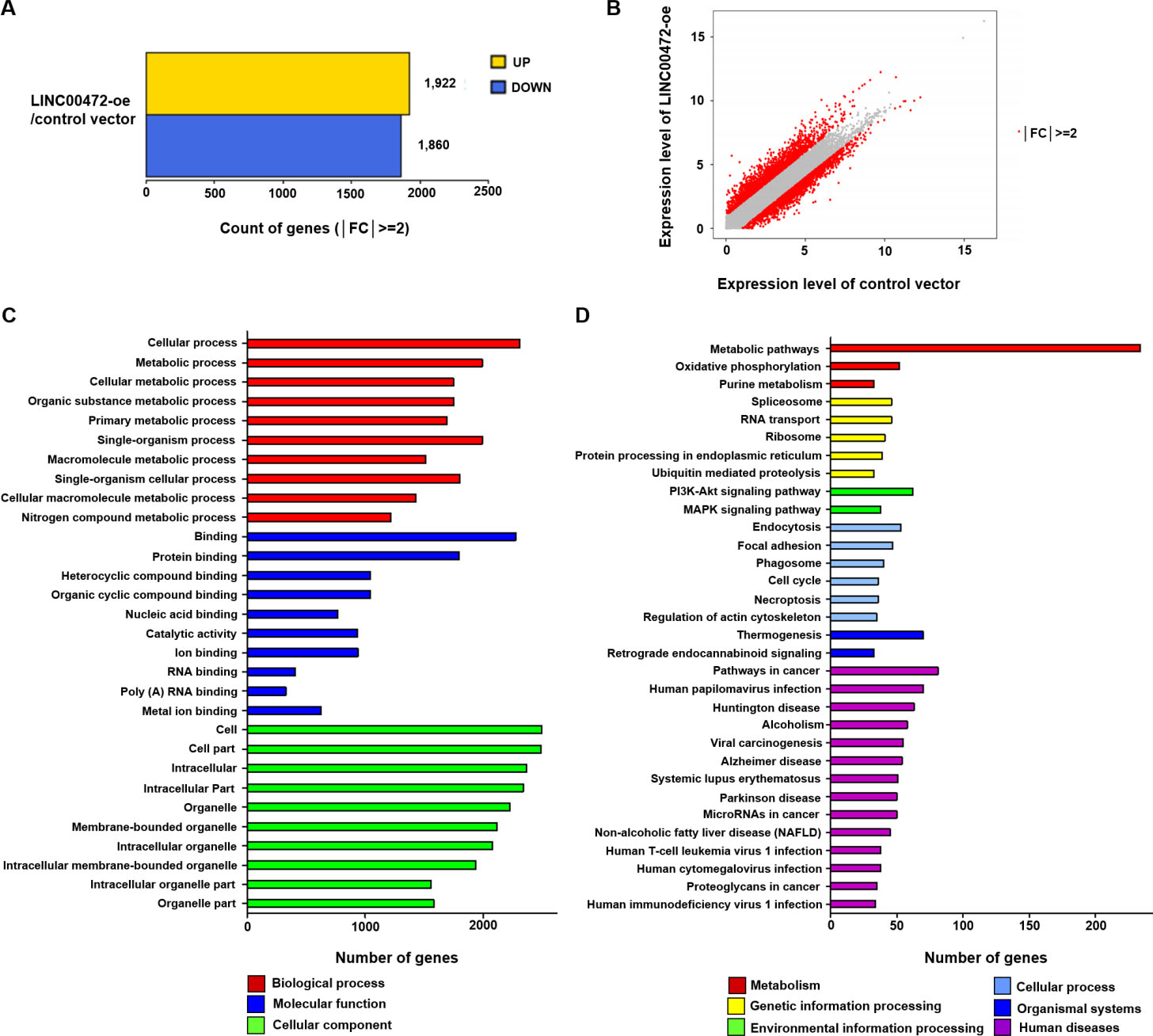
GO terms and KEGG pathways were analyzed to evaluate DEG functions and associated signaling pathways. GO and KEGG enrichment analyses were used to identify the top 10 terms of GO category and the top 32 KEGG pathways. In the Biological Process GO category, enriched DEGs were associated with the following processes; “cellular process” (2,308 genes,
KEGG pathway analysis revealed significant gene enrichment in the 32 top pathways (Fig. 2D). Of the 32 top-ranked pathways, “metabolic pathways” (234 genes,
Selected genes among DEG when LINC00472 is overexpressed
To identify genes that interact with LINC00472, 12 genes were selected from among DEGs identified by total RNA-seq (each
Total-RNA seq showed PPP1R12B, RGS5, RBM5, RBL2, LDLR, and PTPRM were upregulated in LINC00472-oe cells, and that SPSB1, PCNA, CD24, CDK5, CDC25A, EIF4EBP1 were downregulated (each
The expression levels of 12 selected DEGs in the LINC00472 overexpressing NCI-H838 cells as compared with those in cells treated with control vector. Six genes (PPP1R12B, RGS5, RBM5, RBL2, LDLR, and PTPRM) were upregulated, and the other six genes (SPSB1, PCNA, CD24, CDC5, CDC25A, EIF4EBP1) were downregulated (all 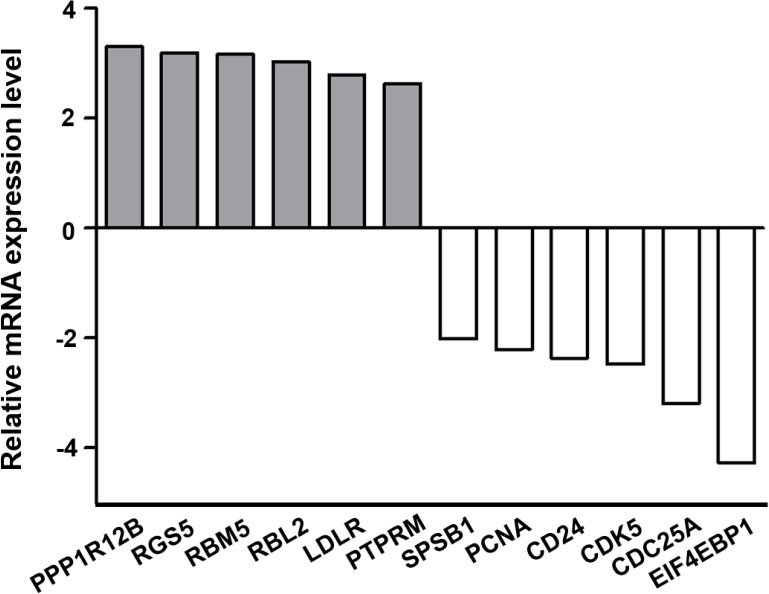
The expression patterns of 12 selected DEGs in NSCLC tissues based on the Oncomine database. (A) The Landi dataset showed PPP1R12B is downregulated in lung adenocarcinoma (fold change 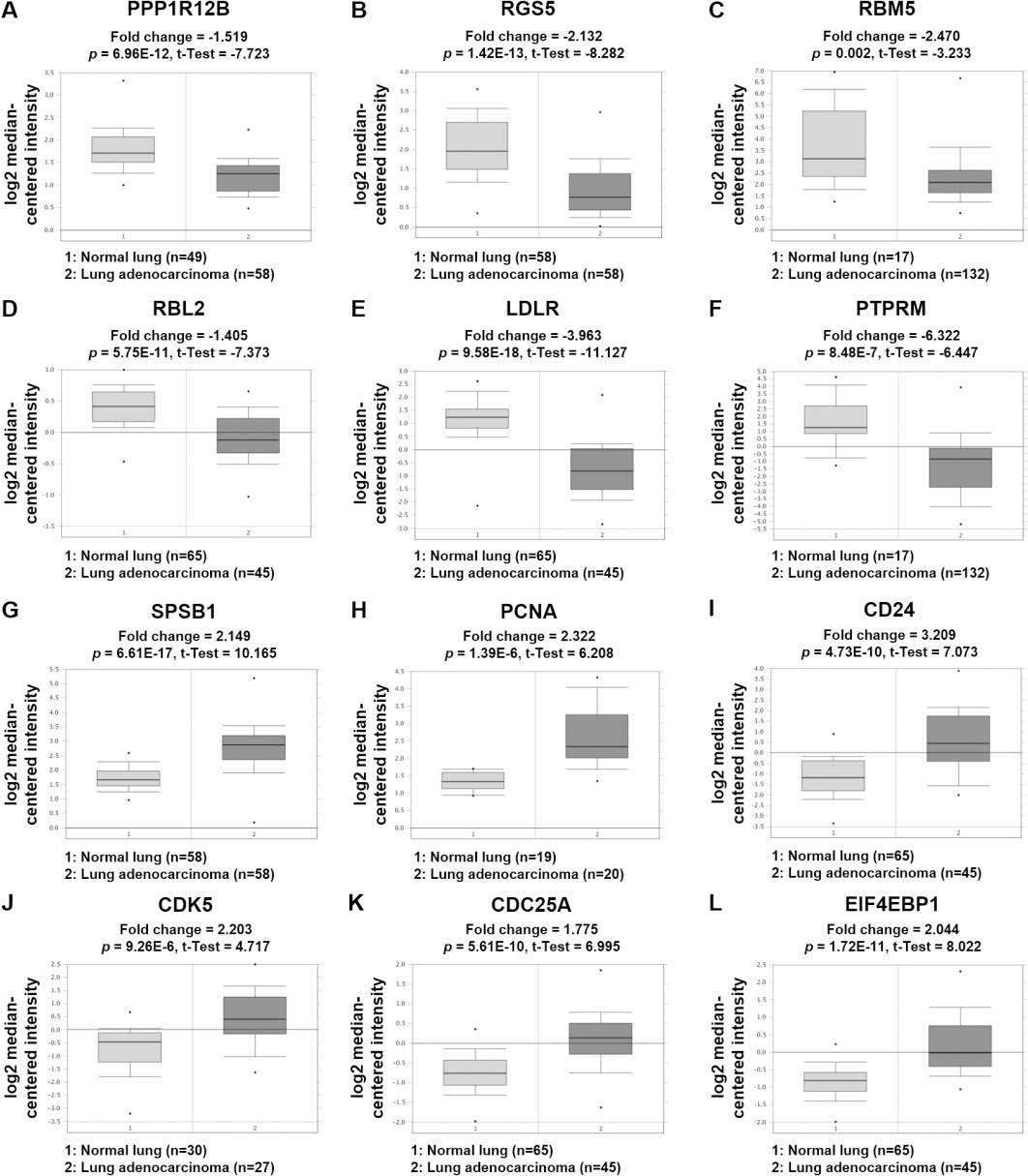
According to recent cancer studies, LINC00472 acts as a tumor suppressor. However, despite strong evidence supporting this role, few studies have investigated LINC00472-related mechanisms in NSCLC. In this study, to identify genes that interact with LINC00472 in NSCLC, mRNA expressions were profiled by transcriptome sequencing analysis of a LINC00472 overexpressing NSCLC cell line.
We prepared an NSCLC cell line overexpressing LINC00472 by transient transfection. As a result of measuring the cell viability, proliferative capacity, and migration capacity, it was confirmed that LINC00472 has a tumor-suppressive function in NSCLC cells. Transcriptome sequencing of empty vector controls and LINC00472-oe cells resulted in the screening of 3,782 genes as DEGs. Many genes were found to be related to metabolism by GO and KEGG pathway analyses. The gene regulations of intracellular metabolism have been linked to many cancers. For example, spermidine/ spermine N1-acetyltransferase 1 (SAT1), which was not included in the list of 12 selected DEGs but was upregulated by 3.64 fold in LINC00472-oe cells, functions as a rate-limiting enzyme in the catabolic pathway of polyamine metabolism and acetylates spermidine and spermine [54], and disruption of polyamine metabolism by abnormal SAT1 expression is associated with cancer [55]. In a study of the mammalian cell line 293T, it was demonstrated that SAT1 overexpression rapidly depleted intracellular spermidine and spermine, and thus, inhibited cell growth and the induction of mitochondrial apoptosis [56]. In addition, it has been reported that p53 activates the expression of SAT1, and that activated SAT1 induces lipid peroxidation resulting in ferroptosis caused by ROS-induced oxidative stress [57]. SAT1 was included in KEGG “metabolic pathways” and in GO “metabolic process”. Adenylosuccinate lyase (ADSL) provides another example. ADSL is involved in the synthesis of de novo purine, and ADSL deficiency leads to defects in the metabolism of purines [58], which are the basic building blocks of nucleotides. In actively dividing cancer cells, the de novo purine biosynthetic pathway is required to supplement purines [59], and thus, the expression of ADSL may be related to tumor growth. In fact, ADSL has been shown to be upregulated in several cancers [60, 61, 62]. In this study, ADSL was included in KEGG “metabolic pathways” and GO “metabolic process”. Also, ADSL was downregulated
Twelve DEGs (PPP1R12B, RGS5, RBM5, RBL2, LDLR, PTPRM, SPSB1, PCNA, CD24, CDK5, CDC25A, EIF4EBP1) selected based on cell cycle, survival, and immunity were not included in KEGG “metabolic pathways”, but were included in GO “metabolic process”. In LINC00472-oe cells, genes PPP1R12B, RGS5, RBM5, RBL2, LDLR and PTPRM were upregulated, but are downregulated in NSCLC tissues. These genes are expected to function as biomarkers as tumor suppressors in lung cancer. In fact, it has been reported that tumor cells exhibiting high RGS expression in NSCLC have a low risk of lymph node metastasis and vascular invasion [63]. RBM5 is a tumor suppressor gene in lung adenocarcinoma cells and induces apoptosis and autophagy, inhibits tumor growth, and improves chemosensitivity [64]. On the other hand, reduced RBM5 expression in an in vivo study on RBM5 loss-of-function mice was associated with more aggressive lung cancer in terms of cell numbers and tumor size [65]. RBL2 has also been reported to suppress tumor cell proliferation in vitro [66]. A study on lung cancer and mesothelioma suggested that RBL2/p130 is a direct substrate of AKT kinase and that it is required for AKT inhibition-induced apoptosis [67]. In lung cancer, PTPRM dephosphorylated STAT3 and its overexpression inhibited cell growth [68]. Although PPP1R12B and LDLR have not been sufficiently studied in lung cancer, there is evidence that they function as tumor suppressors in other cancers. PPP1R12B inhibited tumor growth and metastatic capacity by deregulating Grb2/PI3K/Akt signaling in colorectal cancer [69], and LDLR overexpression reduced cell proliferation in cervical cancer [70]. Consequently, it appears PPP1R12B, RGS5, RBM5, RBL2, LDLR, and PTPRM act as tumor suppressors in several cancers, and that LINC00472 might act as a tumor suppressor by upregulating the expressions of these genes.
In contrast, SPSB1, PCNA, CD24, CDK5, CDC25A, and EIF4EBP1 were downregulated in LINC00472-oe cells but upregulated in NSCLC tissues. These genes are expected to function as oncogenic biomarkers in lung cancer. For example, in NSCLC, PCNA promoted cell proliferation and tumorigenesis and inhibited cell death by phosphorylating STAT3 [71], which contrasts to the activities of PTPRM, which dephosphorylates STAT3 in lung cancer cells. As regards CD24, its downregulation in lung cancer inhibited metastasis of lung cancer cells to bone in vivo [72]. High CDK5 expression in lung cancer was reported to be responsible for poorer prognosis, and CDK5 was also found to promote proliferation and metastasis of lung cancer cells [73], and to upregulate a transcriptional coactivator with a PDZ-binding motif (TAZ) to activate the Hippo signaling pathway, and thus, increasing radiation resistance of lung cancer cells [74]. On the other hand, reduced expression of CDC25A targeted by mir-365 improved the radiation sensitivity of NSCLC cells [75]. Finally, the expression of phosphorylated EIF4EBP1 was reported to be significantly correlated with poor prognosis in SCLC [76]. In contrast to EIF4EBP1 in the nucleus responsible for translation inhibition, EIF4EBP1 overexpressed in the cytoplasm has been reported to promote tumor growth by activating a hypoxic response to convert cap-dependent translation into cap-independent translation [47]. Studies on SPSB1 in lung cancer are lacking, but it has been reported to function as an oncogene in several cancers. SPSB1 has been associated with breast cancer recurrence [77]. and the survival of cancer cells in ovarian cancer [78]. Taken together, SPSB1, PCNA, CD24, CDK5, CDC25A, and EIF4EBP1 act as oncogenes in various cancers, and LINC00472 is expected to downregulate the expressions of these genes, and thereby, inhibit tumor growth.
Our mRNA profiling data indicate that LINC00472 regulates the expressions of genes that are important in various signaling pathways, such as metabolic pathways, involved in metabolism, cell cycle, and tumorigenesis. Analysis of selected DEGs confirmed that several genes expressionally regulated by LINC00472 are involved in various cancer-related signaling pathways. Consequently, our findings indicate LINC00472 acts as a tumor suppressor by increasing the expressions of tumor suppressor genes and decreasing the expressions of oncogenes.
The selected DEGs regulated by LINC00472 are expected to interact closely with LINC00472, but, the molecular mechanisms of these interactions, including intracellular interactions, need to be demonstrated experimentally in vitro and in vivo. Nevertheless, the profiling of mRNA expressions in LINC00472 overexpressing NSCLC cells provides a blueprint for the function and role of LINC00472 in NSCLC cells. We believe this will considerably aid understanding of the function of LINC00472 in NSCLC cells and provide guidance for biochemical studies aimed at elucidating signaling pathways in cancer involving LINC00472.
Footnotes
Acknowledgments
We thank Herbert Yu (University of Hawaii Cancer Center, Honolulu, HI, USA) for their valuable supports with providing the plasmid constructs (pCDH-LINC00472 and pCDH).
Funding
This work was supported by the National Research Foundation of Korea (NRF) funded by the Ministry of Science and ICT (2019R1C1C1009423).
