Abstract
BACKGROUND:
Non-small cell lung cancer (NSCLC) is the most common cancer worldwide. Circular RNAs (circRNAs) are recently identified as important gene regulators with critical roles in cancer biology. In this study, we focus on the effect of circ_0000376 targeting miR-384 on malignant phenotypes of NSCLC cells.
METHODS:
Circ_0000376 and miR-384 expression in NSCLC tissue samples were measured using qRT-PCR. The association between pathological parameters and the circ_0000376 expression was analyzed as well. Human NSCLC cell lines A549 and NCI-H460 were used as cell models. CCK-8 and BrdU assay were used to assess the effect of circ_0000376 on NSCLC cell line proliferation and drug sensitivity. Transwell assay was conducted to detect the effect of circ_0000376 on migration and invasion. Further, luciferase reporter assay was employed to validate the targeting of miR-384 by circ_0000376.
RESULTS:
Circ_0000376 expression in NSCLC clinical samples was up-regulated and this was linked to unfavorable pathological parameters. Circ_0000376 markedly accelerated the proliferation and metastasis, and enhanced chemoresistance of NSCLC cells. Mechanically, circ_0000376 overexpression could bind with miR-384 and repress its expression.
CONCLUSIONS:
Circ_0000376 is a newly discovered oncogenic circRNA in NSCLC, and can be potentially regarded as a diagnostic biomarker and therapy target.
Introduction
Non-small cell lung cancer (NSCLC) is the most common malignant tumor with the highest morbidity and mortality worldwide [1]. In accordance with a report issued by the World Health Organization in 2018, there are 18.1 million newly diagnosed cancer cases and 1.8 million lung cancer-related deaths every year [2]. Unfortunately, most patients are at advanced stage when they are diagnosed. Current chemotherapy has been applied to more than 90% of NSCLC patients, and platinum-based chemotherapy is the main treatment strategy [3].
Although circular RNAs (circRNAs) have been previously considered as a by-product of splicing, it has now been regarded as a key regulator of a wide range of biological processes. Recently, circRNA has been proved to participate in the occurrence and progression of many human diseases including tumors [4, 5]. For instance, circRHOT1 accelerates liver cancer progression via inducing NR2F6 expression [6]. CircRNA_069718 enhances the proliferation and invasion of triple-negative breast cancer cells via activating Wnt/
MicroRNAs (miRNAs) are small non-coding RNAs (ncRNA) containing 19–25 nucleotides, which are characterized by the binding of miRNAs to the complementary sequence of the 3’ untranslated region (3’ UTR) of the mRNA, resulting in mRNA degradation, thereby effectively repressing its target gene [8]. Increasing researches have discovered that miRNAs are not only engaged in the differentiation, proliferation, apoptosis, and metabolism of normal cells, but also in the tumorigenesis and metastasis of tumors [9]. For instance, miR-134 blocks the progression of esophageal squamous cell carcinoma through regulating PLXNA1-mediated MAPK signaling pathway [10]. miR-3653 impedes the epithelial-mesenchymal transition (EMT) of colon cancer cells by targeting ZEB2 [11]. It is confirmed that miR-384 is remarkably down-regulated in NSCLC tissues, reflecting that miR-384 exerts a tumor suppressor effect in NSCLC progression [12]. Nonetheless, the upstream mechanism that causes miR-384 low expression remains largely undefined.
It should be noted that bioinformatics analysis suggested that there is a potential binding site between circ_0000376 and miR-384, and it is predicted that circ_0000376 may be a molecular sponge for miR-384. Herein, we demonstrated that circ_0000376 was highly expressed in NSCLC, and in vitro results implied that circ_0000376 enhanced the proliferation, metastasis, and chemotherapy resistance of NSCLC cells. Circ_0000376, as the sponge of miR-384, attenuated the inhibitory effect of miR-384 on the malignant phenotypes of NSCLC cells, thereby accelerating NSCLC progression. Our study helped clarify the molecular regulatory mechanism in the development of NSCLC, and could provide new potential therapeutic targets for this disease.
Materials and methods
Tissue specimens
The cancer tissues and their corresponding adjacent normal tissues were collected during surgery and immediately frozen in liquid nitrogen for further study and analysis. The histology of NSCLC patients was identified, and none of the patients received pre-operative radiotherapy or chemotherapy. All patients involved gave informed consent to the study and signed a written consent form. The cancer tissues and matched adjacent tissues were collected under the approval of Ethics Review Committee of Jiangsu Province Hospital.
Cell culture
Normal bronchial epithelial cells 16HBE and NSCLC cell lines (A549, NCI-H520, LK-2, NCI-H2291, and NCI-H460 cells) were purchased from the Cell Center of Chinese Academy of Sciences (Shanghai, China). Cells were cultured in RPMI-1640 medium containing 10% fetal bovine serum (FBS) (Thermo Fisher Scientific, MA, USA) and 1% penicillin/streptomycin (Invitrogen, Carlsbad, CA, USA), and cultured in an incubator at 37
Real-time quantitative PCR
After the total RNA was extracted from each group using TRIzol Reagent (Thermo Fisher, Shanghai, China) and the concentration was measured, total RNA was reversely transcribed into cDNA with the PrimeScript RT Reagent Kit (Invitrogen, Shanghai, China) according to the manufacturer’s protocol. SYBR Green Premix Ex Taq II (Takara, Dalian, China) was adopted for RT-PCR. The primer sequences were as shown in Table 1. Reaction conditions: pre-denaturation at 95
Primer sequences used for RT-PCR in this study
Primer sequences used for RT-PCR in this study
Abbreviation: F stands for forward; R stands for reverse; RT stands for reverse transcription.
The association between the expression level of circ_0000376 and pathological indexes
Hsa-miR-384 mimics and inhibitors, and negative controls (NC) were designed and constructed by GenePharma (Shanghai, China). The circular transcript expression vector possesses two elements termed as the front circular and the back circular frame which were specially designed and containing inverted repeat sequences flank. The full-length cDNA of circ_0000376 was amplified in NSCLC cells, and was cloned into the specific vector between two frames. To knockdown circ_0000376, three small interfering RNAs (siRNAs) against circ_0000376 and siRNA-NC (si-NC) were synthesized. In brief, cells were seeded at a density of 5
Cell proliferation assay
NSCLC cells in the logarithmic growth phase were trypsinized with trypsin, adjusted to a cell density of 2
BrdU assay
5
Circ_0000376 upregulated in NSCLC tissues and cell lines. (A) In 60 pairs of NSCLC tissues and adjacent normal bronchial tissues, qRT-PCR results for circ_0000376 showed that circ_0000376 was significantly up-regulated in NSCLC tissues compared with normal bronchial tissues. (B) qRT-PCR was performed on circ_0000376 in normal bronchial cells and NSCLC cell lines, and the results showed that the level of circ_0000376 was significantly up-regulated in the NSCLC cell lines compared with normal bronchial cells.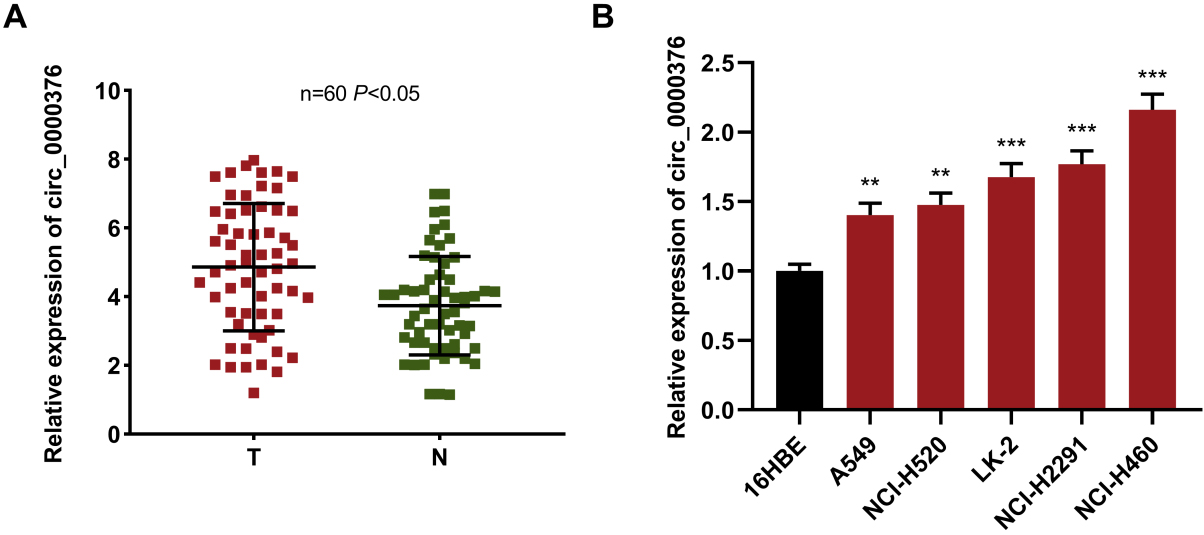
Circ_0000376 promoted the proliferation and drug resistance of NSCLC cells in vitro. (A) Expression of circ_0000376 was detected by qRT-PCR in A549 cells transfected with empty plasmid or overexpressing circ_0000376 plasmid, and in NCI-H460 cells transfected with three siRNAs targeting circ_0000376 or negative control siRNA. (B) CCK-8 assay showed that overexpression of circ_0000376 promoted the proliferation of A549 cells, and knockdown of circ_0000376 inhibited the proliferation of NCI-H460 cells. (C) Determination of the IC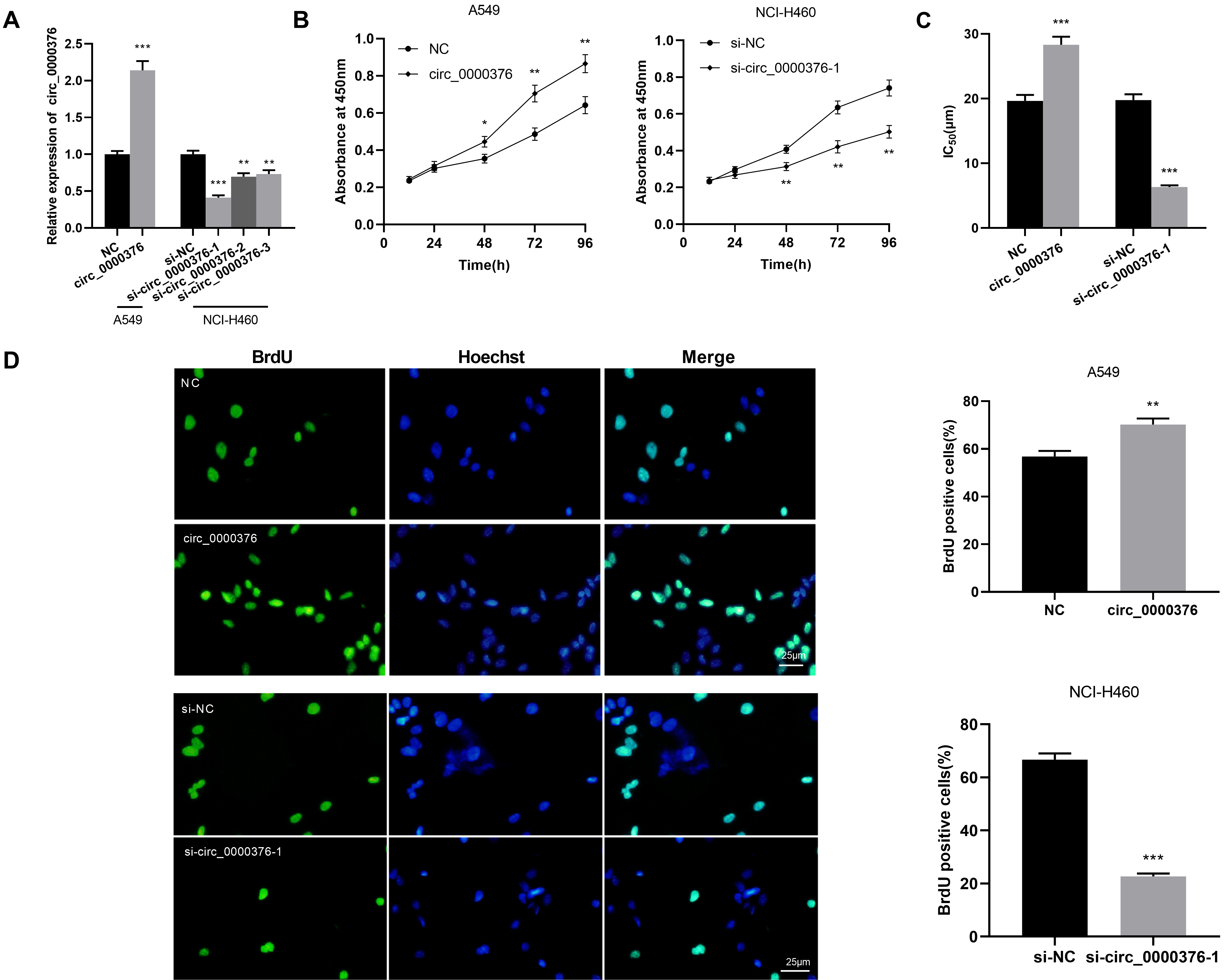
CCK-8 assays were performed to determine the sensitivity of NSCLC cells to cisplatin. In brief, first, cells were seeded into 96-well plates (3000 cells/well) and cultured overnight. The cells were then treated with different concentrations of cisplatin for 24 h. After that, the medium was discarded and 10
Cell migration and invasion assays
The experimental steps of cell migration and cell invasion were basically the same. The only difference was that the cell invasion assay used a Transwell chamber (pore size, 8
Dual-luciferase reporter assay
Together with miR-384 mimic or NC microRNAs, circ_0000376 wild-type or mutant plasmid containing the luciferase reporter were respectively transfected into the cells. After the cells were cultured for 48 h, the medium then was discarded. Subsequently, the cells were washed with PBS. Next, the cell lysates were added to the wells to lyse the cells. Afterwards, the cells were oscillated for 5 to 10 min at room temperature and centrifuged at 3000
Statistical analysis
The statistics of the data were analyzed using SPSS 17.0 statistical software (SPSS Inc., Chicago, IL, USA). All data were expressed as the average mean
Result
Circ_0000376 upregulated in NSCLC tissues and cell lines
First, qRT-PCR was employed to detect the circ_ 0000376 expression in 60 pairs of NSCLC tissues. In comparison with normal tissues adjacent to cancer, circ_0000376 was remarkably over-expressed in NSCLC tissues (Fig. 1A). Furthermore, we demonstrated that circ_0000376 expression levels were higher in NSCLC cell lines including A549 and NCI-H460 cells in comparison with the normal bronchial epithelial cells 16HBE (Fig. 1B). These results implied that circ_0000376 could take part in promoting cancer progression of NSCLC.
Circ_0000376 was linked to multiple clinicopathological indicators of NSCLC
To further clarify the role of circ_0000376 in the tumorigenesis and progression of NSCLC, the above 60 cases of NSCLC samples were employed to analyze the correlation between the circ_0000376 expression and diverse pathological indicators of NSCLC patients (Table 2). Chi-square test revealed that circ_0000376 high expression in tumor tissues was notably linked to the increased tumor T stage (
Circ_0000376 was involved in the regulation of NSCLC cell migration and invasion. (A) Transwell assay showed that circ_0000376 overexpression significantly promoted the migration and invasion of A549 cells; circ_0000376 knockdown significantly inhibited migration and invasion of NCI-H460 cells. (B) The expression levels of EMT marker proteins E-cadherin, N-cadherin, and Vimentin were determined by Western blot.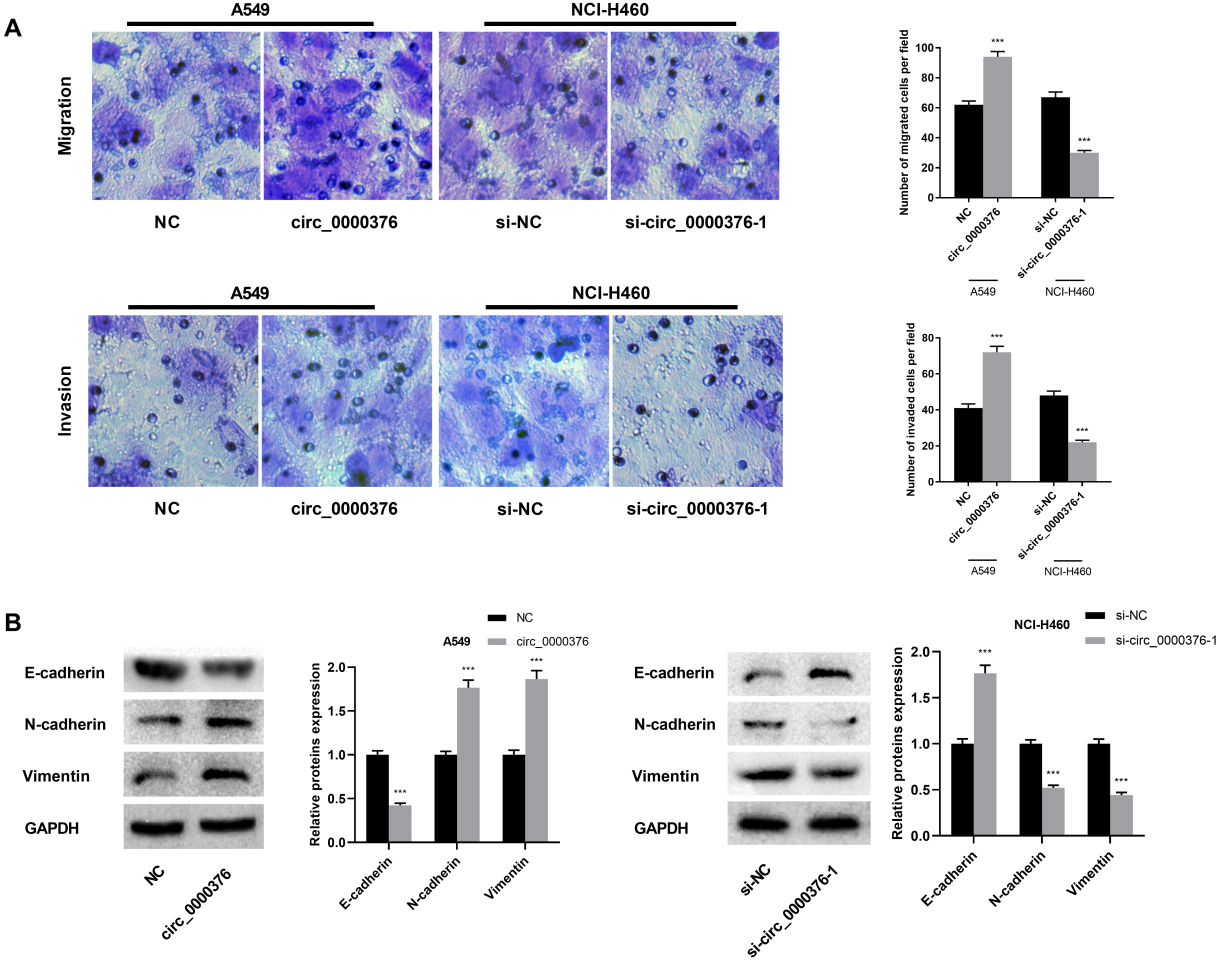
Circ_0000376 targeted miR-384. (A) Schematic representation of the predicted miR-384 binding site in the wild-type and mutant binding sites of circ_0000376. (B) The dual-luciferase reporter assay showed a significant decrease in luciferase activity of the vector containing the wild type (WT) miR-384 binding site in circ_0000376. (C) qRT-PCR results showed that the knockdown or overexpression of circ_0000376 affected the expression level of miR-101 in A549 and NCI-H460 cells. (D) qRT-PCR results showed that circ_0000376 was negatively correlated with miR-384 expression in NSCLC samples.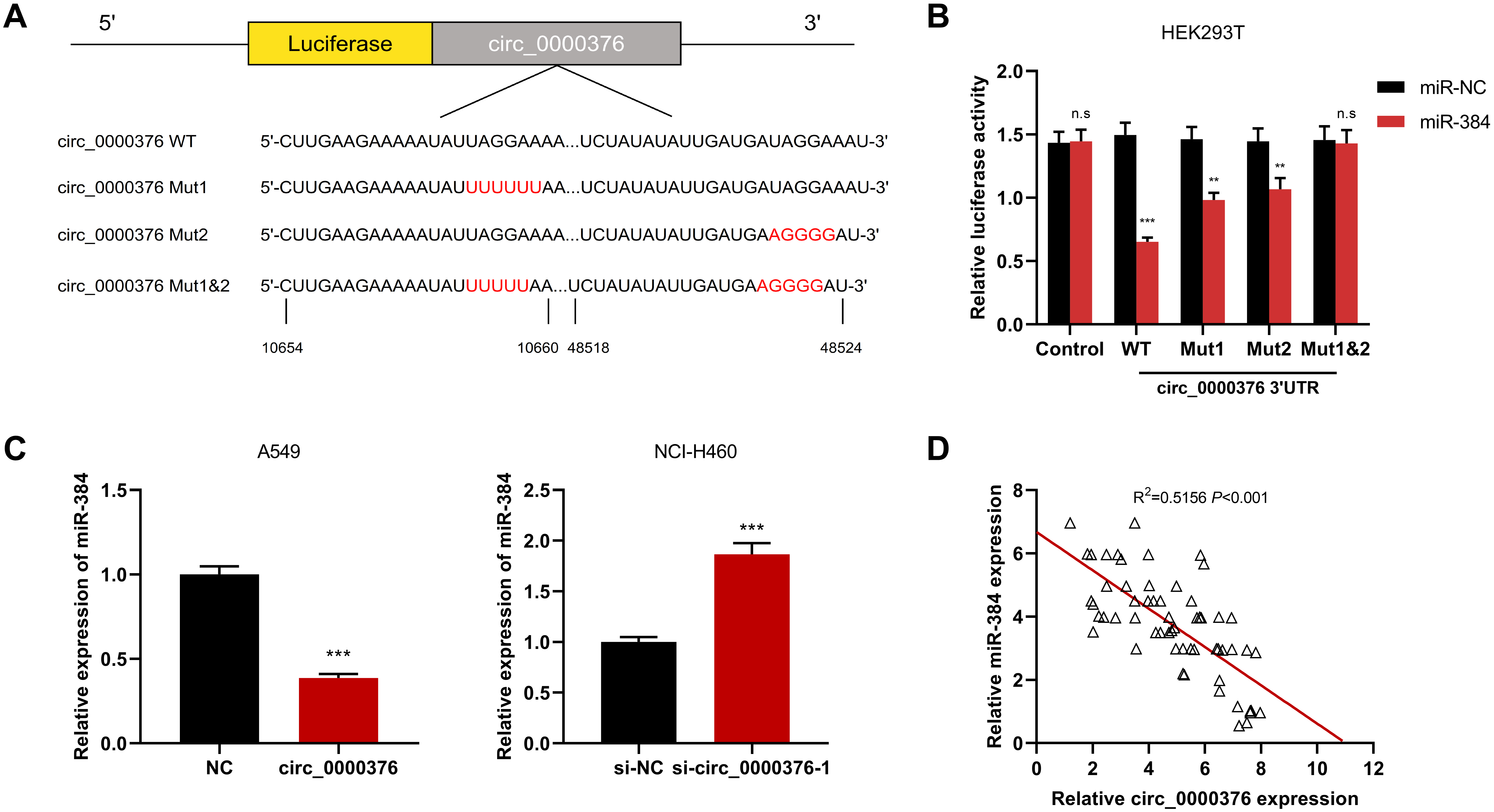
Circ_0000376 regulated the proliferation, metastasis, and drug resistance of NSCLC cells by targeting miR-384. (A) CCK-8 assay showed that overexpression of miR-384 inhibited the proliferation of A549 cells, while overexpression of circ_0000376 attenuated the inhibition of cell proliferation by miR-384; inhibition of miR-384 promoted the proliferation of NCI-H460 cells, while knocking down circ_0000376 could reverse this effect. (B) Based on the CCK-8 method, the IC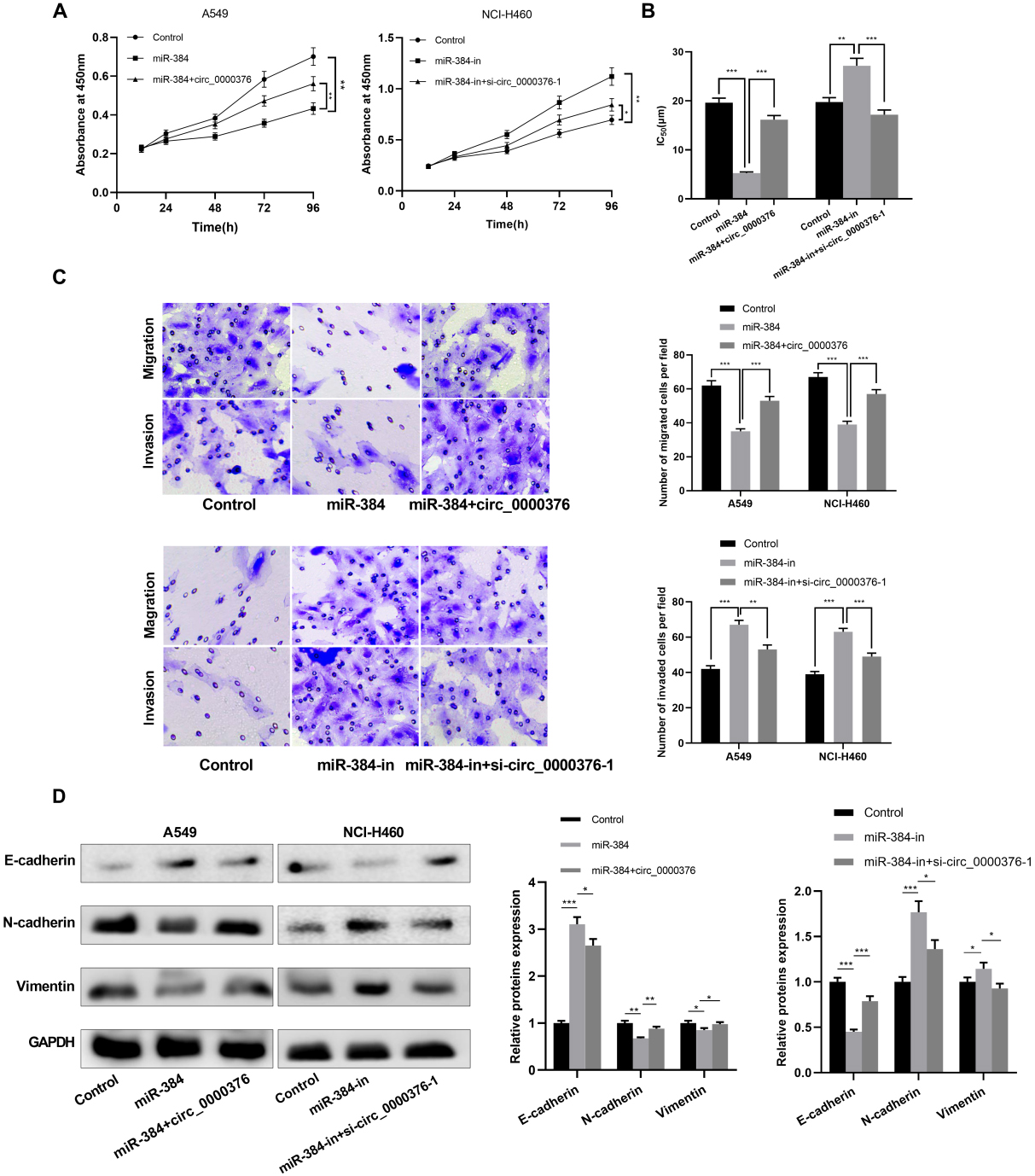
Among the five NSCLC cells, the circ_0000376 expression was the lowest in A549 cells, while the highest is in the NCI-H460 cells. Therefore, A549 cells were selected to construct an overexpressed circ_0000376 cell model, while NCI-H460 cells were selected to construct a circ_0000376 knockdown cell model, and the transfection was verified by qRT-PCR (Fig. 2A). Since the knockdown effect of si-circ_0000376-1 was the most remarkable in the three knockdown cell models, si-circ_0000376-1 was adopted for subsequent experiments. Hence, CCK-8 and BrdU assays were conducted to monitor cell proliferation and calculate IC
Circ_0000376 took part in the regulation of NSCLC cell migration and invasion
After clarifying that circ_0000376 can modulate the proliferation and chemoresistance of NSCLC cells, we were curious about whether circ_0000376 can modulate the migration and invasion. To probe the role of circ_0000376 in NSCLC cell migration and invasion, Transwell assay was performed. As is shown, in comparison with the control group, A549 cells overexpressing circ_0000376 showed a pronounced increase in the ability of migration and invasion. Conversely, the migration and invasion of NCI-H460 cells were impeded by circ_0000376 knockdown (Fig. 3A). Since our data indicated that circ_0000376 may take part in the cancer metastasis, whether circ_0000376 can modulate gene expression in tumor metastasis markers was then investigated. Three important biomarkers of EMT, E-cadherin, N-cadherin, and Vimentin were examined by Western blot. As we expected, circ_0000376 overexpression reduced the E-cadherin expression in A549 cells and increased the N-cadherin and Vimentin expression; on the contrary, knocking down circ_0000376 increased the E-cadherin expression in NCI-H460 cells and decreased N-cadherin and Vimentin expression (Fig. 3B), reflecting that circ_0000376 facilitated the migration and invasion of NSCLC cells via regulating EMT process.
Circ_0000376 targeted miR-384
Then we searched for miRNAs that could potentially bind with circ_0000376 using the online bioinformatics database (
Circ_0000376 modulated the malignant phenotypes of NSCLC cells via targeting miR-384
To further probe the role of circ_0000376/miR-384 axis in NSCLC progression, the proliferation of NSCLC cells was monitored by CCK-8 assay. In comparison with the control group, the proliferation of NCI-H460 cells was remarkably attenuated after transfection with miR-384 mimics. In addition, the sensitivity of the cells to cisplatin was enhanced. In comparison with the control group, after transfection of miR-384 inhibitors, the proliferation of A549 cells was strikingly enhanced, and the cisplatin chemotherapy resistance to the cells was enhanced. Moreover, in comparison with the overexpressed miR-384 group, the proliferation of NSCLC cells was markedly enhanced after further up-regulation of circ_0000376, and the cisplatin chemotherapy resistance to the cells was enhanced. In comparison with the low-expressed miR-384 group, circ_0000376 knockdown notably impeded the proliferation of NSCLC cells, and the cisplatin sensitivity to the cells was enhanced (Fig. 5A and B). Transwell assays reflected that the migrated and invaded NCI-H460 cells were remarkably inhibited after transfection of miR-384 mimics compared with the control group, and this inhibition was attenuated by circ_0000376. Conversely, after the miR-384 inhibitor was transfected into NCI-H460 cells, the migrated and invaded cells increased. Additionally, circ_0000376 knockdown strikingly repressed the metastasis of NSCLC cells in comparison with the miR-384 inhibition group (Fig. 5C). After discovering that circ_0000376 modulated miR-384s to regulate NSCLC cell metastasis, their effect on EMT progression was investigated. As we expected, transfection of miR-384 mimics increased the E-cadherin expression, and the N-cadherin and Vimentin expression declined, while circ_0000376 overexpression could reverse the effect. Additionally, transfection of miR-384 inhibitors attenuated E-cadherin expression while increasing the N-cadherin and Vimentin expression, but knocking down circ_0000376 reversed this effect (Fig. 5D). To sum up, we concluded that circ_0000376 can modulate the malignant phenotypes of NSCLC cells via targeting miR-384.
Discussion
CircRNA is a class of ncRNAs with a stable circular structure that acts as a competing endogenous RNA (ceRNA) or as an RNA sponge to modulate mRNA expression [15]. Previous studies have confirmed that circRNA plays an essential role in tumorigenesis and can affect tumor cell proliferation, apoptosis, and metastasis. For instance, circ_0058124 facilitates the development and invasion of papillary thyroid carcinoma through the NOTCH3/GATAD2A axis [16]; the upregulation of CircRNA-CDC45 expression is a poor prognostic indicator of gliomas [17]; F-circEA-4a, a tumor-promoting circRNA generated from the back-splicing of EML4-ALK variant 3b (v3b), promotes cancer cell migration and invasion, and is a novel liquid biopsy biomarker for NSCLC [18, 19]. Knocking down circRNA_102231 strikingly impeded the proliferation and invasion of lung cancer cells [20]; circ_100876 was highly expressed in lung cancer, and its overexpression was extremely linked to the poor prognosis of patients [21]; circ_0000064 was upregulated in lung cancer tissues and cell lines, and knockdown of it remarkably attenuated cell proliferation, blocked cell cycle progression, and induced apoptosis [22]. The role of mechanism of circ_0000376 is largely unknown in cancer biology. In this study, we demonstrated that its expression in NSCLC tumor tissues was significantly up-regulated, and its high expression level was linked to unfavorable pathological parameters of NSCLC patients, which implied that circ_0000376 was expected to be an indicator of the poor prognosis in NSCLC. Additionally, overexpression of circ_0000376 notably enhanced the malignant phenotypes of NSCLC cells in comparison with the control group. Nevertheless, knocking down circ_0000376 exerted the opposite effect. These findings suggest that circ_0000376 takes part in the progression of NSCLC.
There are increasing evidences that miRNAs are engaged in diverse physiological and pathological processes. Additionally, miRNAs can bind directly to the target mRNA and induce mRNA degradation or translational inhibition [23]. Currently, miR-384 plays a vital role in multiple tumors. For instance, miR-384 inhibition accelerates the growth and metastasis of osteosarcoma MG63 cells via regulating SLBP [24]. miR-384 is low-expressed in human prostate cancer cells and exerts anti-tumor function via acting on HOXB7 [25]. Likewise, miR-384 represses the progression of esophageal squamous cell carcinoma via blocking LIMK1/cofilin signaling pathway by binding to LIMK1 [26]. miR-384 was also a tumor suppressor in NSCLC, and its target gene includes COL10A1 and AEG-1 [13, 14]. Given that circ_0000376 and miR-384 exert opposite effects in NSCLC, we suspected that there was a targeting relationship between the two. Interestingly, through bioinformatics analysis, we found a binding site between circ_0000376 and miR-384. It was confirmed by luciferase reporter assay that circ_0000376 can adsorb miR-384. Furthermore, knocking down circ_0000376 led to an increase in miR-384 expression. Likewise, miR-384 up-regulation suppressed the proliferation and metastasis of NSCLC cells, and this inhibition can be attenuated by circ_0000376. On the other hand, miR-384 down-regulation strikingly enhanced the proliferation and metastasis in NSCLC cells, while knocking down circ_0000376 remarkably reversed this promotion. Therefore, we concluded that circ_0000376 takes part in the regulation of the proliferation, migration, and invasion of NSCLC cells by sponging miR-384.
This study has certain limitations. First of all, in vivo experiments are needed to verify our conclusion in the following studies. Secondly, other downstream targets of circ_0000376 needs to be identified. Additionally, this study is limited to single-center samples, and larger numbers of clinical samples are needed to confirm our conclusion, and it’s quite interesting to explore whether the dysregulation of circ_0000376 is associated with the overall survival time and disease-free survival time of NSCLC patients. Last but not the least, it is necessary to investigate whether the target genes of miR-384 were modulated by circ_0000376 indirectly, which can further verify the role of circ_0000376 as ceRNA.
Collectively, our findings signified the circ_0000376 high expression in NSCLC and its clinical significance. We also found that circ_0000376 enhances the proliferation, invasion, and metastasis in NSCLC cells via modulating miR-384. To our best knowledge, this is the first study to investigate the role of circ_0000376 in cancer biology and it is expected to provide new therapeutic targets for NSCLC patients.
Footnotes
Conflict of interest
The authors declare that they have no competing interest.
Supplementary data
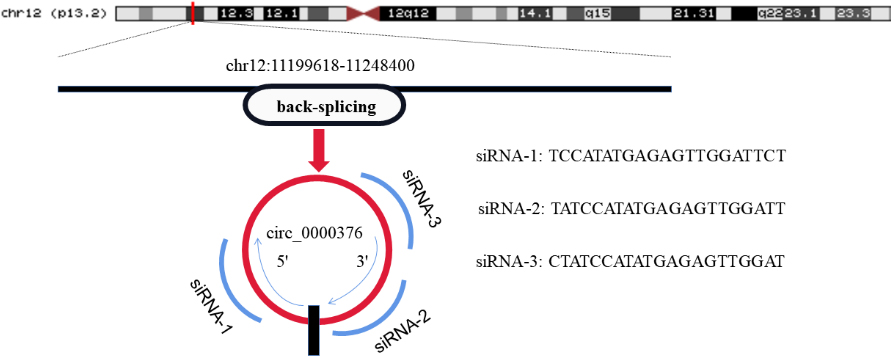
The structure of circ_0000376 and siRNAs for knockdown.