Abstract
Background
Emerging evidence suggests that circular RNAs (circRNAs) were aberrantly expressed in the patients of non-small cell lung cancer (NSCLC). This study aims to evaluate the diagnostic value of potential serum biomarker in circRNAs.
Methods
Serum circRNAs were extracted and purified by RNA isolated kit and identified by quantitative real time-polymerase chain reaction (qRT-PCR) assay. We then performed a receiver operating characteristic (ROC) curve to estimate the diagnostic efficacy. The relationship between circRNA and clinic characteristics of patients was analyzed by SPSS 25.0. Univariate and multivariate analyses were also used to evaluate its diagnostic capability. The mechanism of circFOXP1 was further excavated by bioinformatics analysis.
Results
By performing qRT-PCR assay, we identified that circFOXP1 (hsa_circ_0008234) and conventional tumor markers (carcinoembryonic antigen (CEA) and cytokeratin fragment 21–1 (CYFRA21-1)) were all significantly overexpressed in the serum of patients with NSCLC when compared with healthy controls (P < 0.05). While the ROC curves analysis demonstrated that area under the curve of circFOXP1 was obviously superior to CEA and CYFRA21-1, which exerted more diagnostic advantage. Univariate and multivariate analyses revealed that serum circFOXP1 was an independent diagnostic molecule, and was significantly correlated with T stage and lymphatic metastasis in NSCLC (P < 0.05). Mechanistically, circFOXP1 might target hsa-miR-370-3p and hsa-miR-18a-5p, and be involved in vascular endothelial growth factor signaling pathways to regulate proliferative and metastasis processes.
Conclusion
Our results highlight the preferable diagnostic potential of serum circFOXP1 in NSCLC.
Introduction
Lung cancer is the leading cause of death from cancer. In 2018, the global incidence and mortality rates of lung cancer ranked ahead at 11.6% and 18.5%, respectively. In China, the incidence of lung cancer continually occupied first place among other tumors. 1 Among all cases, more than 80% are non-small cell lung cancer (NSCLC), which can be further subdivided into lung adenocarcinoma (LAD), lung squamous cell carcinoma (LSCC), large cell carcinoma (LCC), and other less frequently diagnosed histological types. 2 Although significant progress has been made in detection measures, the overall 5-year survival rate is still lower than 15%. Increasing the diagnostic ratio may contribute to improve this unsatisfactory diagnosis and treatment status.3,4 With the advantages of accessibility and economy, serum tumor markers gradually developed to be the critical diagnostic method for NSCLC. Traditional tumor markers, such as cytokeratin fragment 21–1 (CYFRA21-1) and carcinoembryonic antigen (CEA), were restricted in clinic utilization due to inferior sensitivity and specificity.5,6 Therefore, exploring novel and effective markers in NSCLC is a matter of urgency.
Circular RNAs (circRNAs) are covalently closed and single-stranded RNA molecules. Compared to linear RNAs, circRNAs maintains high stability when degraded by RNase.7,8 Evolutionary conserved throughout various species, together with tissue/cell types specificity, circRNAs gradually become to be the potential molecule in the field of tumor diagnosis.9–11 Tang et al. discovered that circ-KIAA1244 could serve as the novel circulating biomarker for gastric cancer. 12 In NSCLC, the studies regarding circRNAs are relatively few; only a small portion of circRNAs have been investigated. Thus, it is essential to reveal novel circRNAs and explore their efficiency for the diagnosis of NSCLC.
In the present study, we screened some candidate circRNAs in NSCLC and found that circFOXP1 exhibited high levels and is expected to be a potential diagnostic biomarker for NSCLC.
Materials and methods
Clinical samples
In the retrospective study, 106 serum samples, which included NSCLC and healthy controls between January 2019 and December 2019, were enrolled from the Cancer Center of Guangzhou Medical University (CCGMU). The clinical parameters of patients and the results of laboratory detection were acquired from the records of patients, and included age, sex, and tumor node metastasis stage. Serum samples were collected according to the following criterion: patients without chemotherapy or radiotherapy therapy before collection were diagnosed as having NSCLC by histology. For research purposes, prior patient consents and approval from the Institutional Research Ethics Committee of Guangzhou Medical University were obtained. Clinical information of the samples is described in detail in Supplementary Table 2.
RNA extraction
Total RNA was extracted and purified from the serum samples by the TRIzol reagent kit (Magen, China) according to the manufacturer's instructions and preserved at −80°C. The RNA concentration was detected by using a Nanodrop ND-1000 Spectrophotometer.
Quantitative real-time-polymerase chain reaction
First-strand cDNA was generated from total serum circRNA using MMLV transcriptase (Promega, WI). Quantitative real time-polymerase chain reaction (qRT-PCR) was performed on a CFX96 qRT-PCR detection system (Bio-Rad, Richmond, CA) according to a previous study. 13 The expression levels were measured using the 2−ΔΔCt (Ct is threshold cycle) formula. For cell transfection, cells were transfected with Lipofectamine 3000 (Invitrogen, USA) with indicated vectors. qRT-PCR was then performed.
The sequences of the primers are listed below:
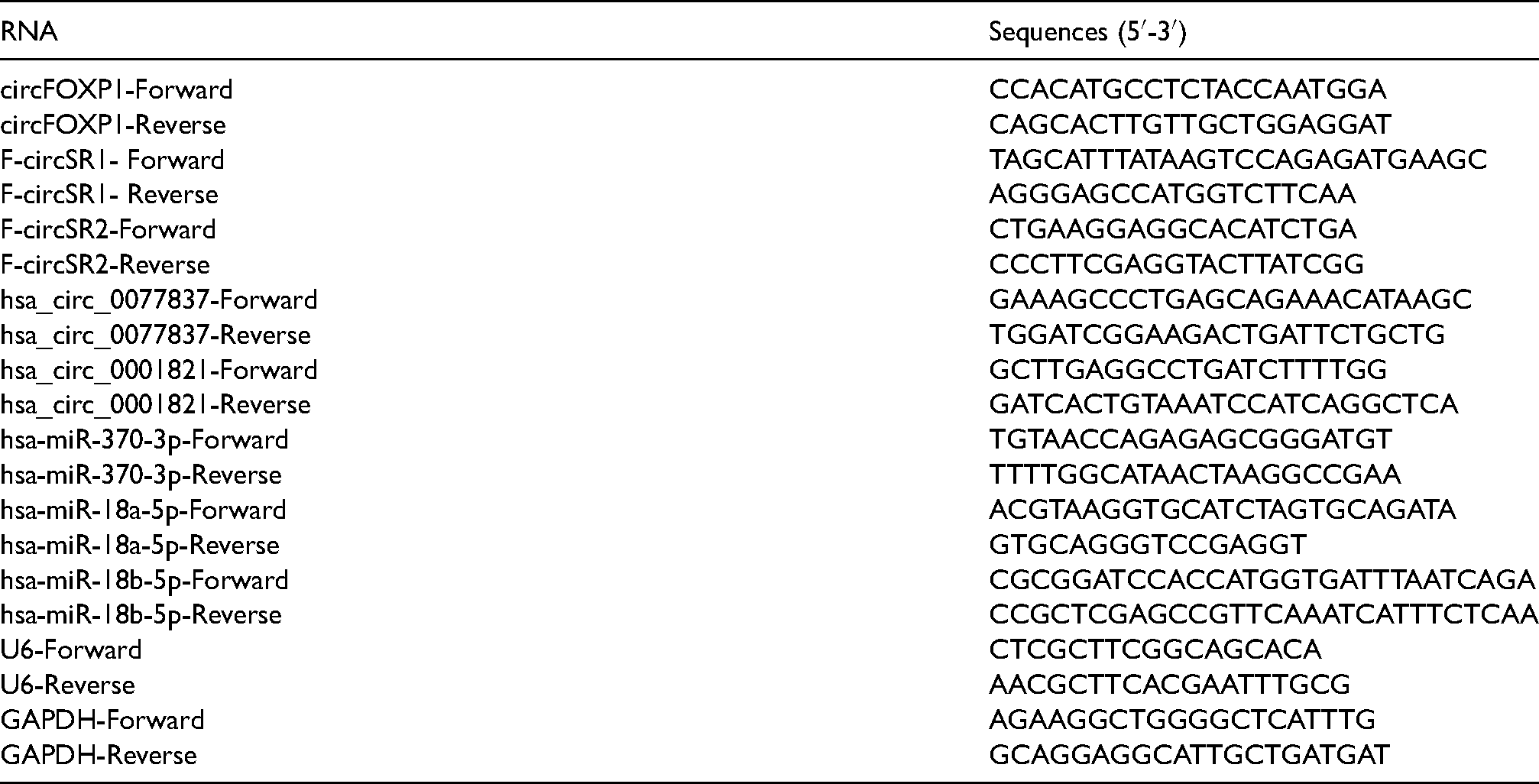
CEA and CYFRA21-1 detection
The serum CEA and CYFRA21-1 levels were tested using an Elecsys-electrochemical immune assay (Roche, USA) and were detected in a cobas 6000 modular analyzer (Roche).
Construction of lncRNA-miRNA-mRNA ceRNA network
The long non-coding RNA-microRNA-messenger RNA-competing endogenous RNA (lncRNA-miRNA-mRNA ceRNA) network was built based on the principle that lncRNAs can interact with miRNA directly by acting as sponges to regulate mRNA activity. 14 FunRich tool (version 3.1.3) package in R software was used to establish ceRNA network. 15 The ceRNA network was plotted with Cytoscape v3.6.0. 16
Statistical analysis
All statistical analyses were performed using the SPSS 25.0 statistical software package. Receiver operating characteristic curves (ROC) was used to determine diagnostic metrics that were calculated using the Delong method. 17 The diagnostic value of circFOXP1 was evaluated using ROC curve and the area under the curve (AUC). Comparisons between groups were performed using Student's t-test. All error bars represent the mean ± SD derived from three independent experiments. P values < 0.05 were considered statistically significant.
Results
Discovery of dysregulated circRNAs in the serum of NSCLC
Candidate circRNAs that were predicted by circBase (http://circrna.org/) and reported by previous studies were selected as potential serum markers in NSCLC.18–20 As shown in Figure 1(a) to (e), candidate circRNAs were validated in 30 cases of patients with NSCLC and 20 cases of healthy controls. circFOXP1 was significantly upregulated in NSCLC compared with controls, while other circRNAs did not exhibit obvious differences. Accordingly, we chose circFOXP1 as the potential molecule for further study (Figure 1(a) to (e)).
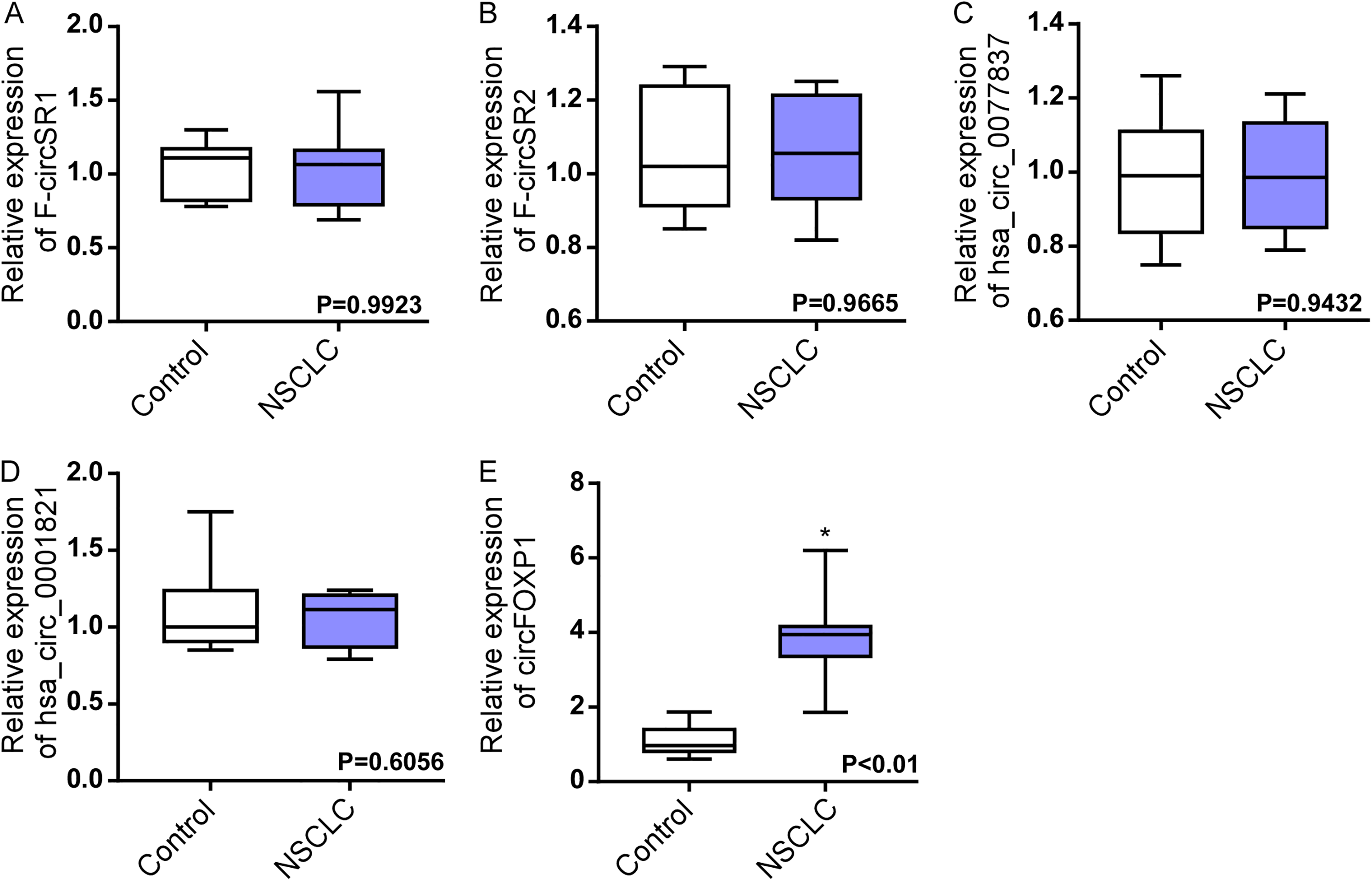
The expression of candidate circRNAs in the serum. (a) to (e) qRT-PCR analysis of circRNA F-circSR1 (a), F-circSR2 (b), hsa_circ_0077837 (c), hsa_circ_0001821 (d) and circFOXP1 (e) in the serum of healthy controls and patients with NSCLC. * P < 0.05.
Verification of circFOXP1 in the validation datasets
CircFOXP1 (hsa_circ_0008234), which is located on chromosome 3 and contains four exons, displays high sequence conservation across mammals (Figure 2(a)). We next validated its expression in 106 serum samples (82 NSCLC patients and 24 healthy controls) by qRT-PCR assay. Traditional serum tumor markers CEA and CYFRA21-1 were concurrently validated in these datasets. As shown in Figure 2(b) to (d), circFOXP1 was significantly overexpressed in NSCLC when compared to healthy controls; this was the same as the overexpression of CEA and CYFRA21-1 (Figure 2(b) to (d)).
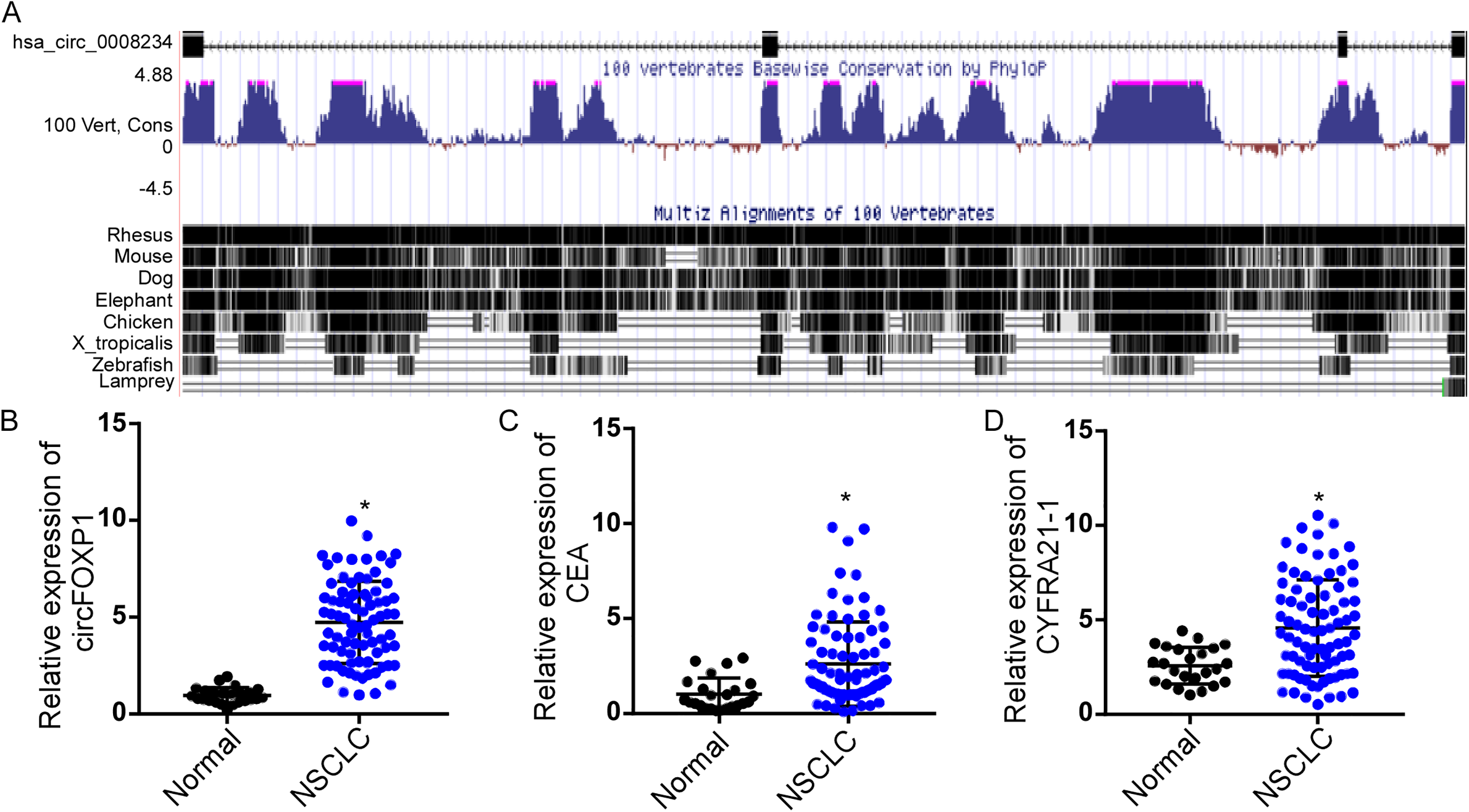
circFOXP1 exhibited high expression levels in patients with NSCLC. (a) Basic information of circFOXP1 on chromosomes and conservative prediction in mammals based on UCSC website (https://genome.ucsc.edu/). (b) to (d) qRT-PCR assay analysis of circFOXP1 (b) CEA (c) and CYFRA21-1 (d) in the serum of healthy controls and patients with NSCLC. * P < 0.05.
ROC curves analysis of circFOXP1 in NSCLC
Based on the qRT-PCR assay, ROC curve was further used to evaluate the diagnostic efficacy of circFOXP1 and two traditional serum tumor markers. The AUC of circFOXP1 was 0.88 (95% confidence interval (CI) = 0.77–0.99) when comparing patients with NSCLC to healthy controls, while the AUCs of CEA and CYFRA21-1 were 0.79 (95% CI = 0.63–0.96) and 0.77 (95% CI = 0.64–0.90), respectively (Supplementary Table 1). The results indicated that circFOXP1 was superior in distinguishing NSCLC from healthy controls (Figure 3(a) to (c)).
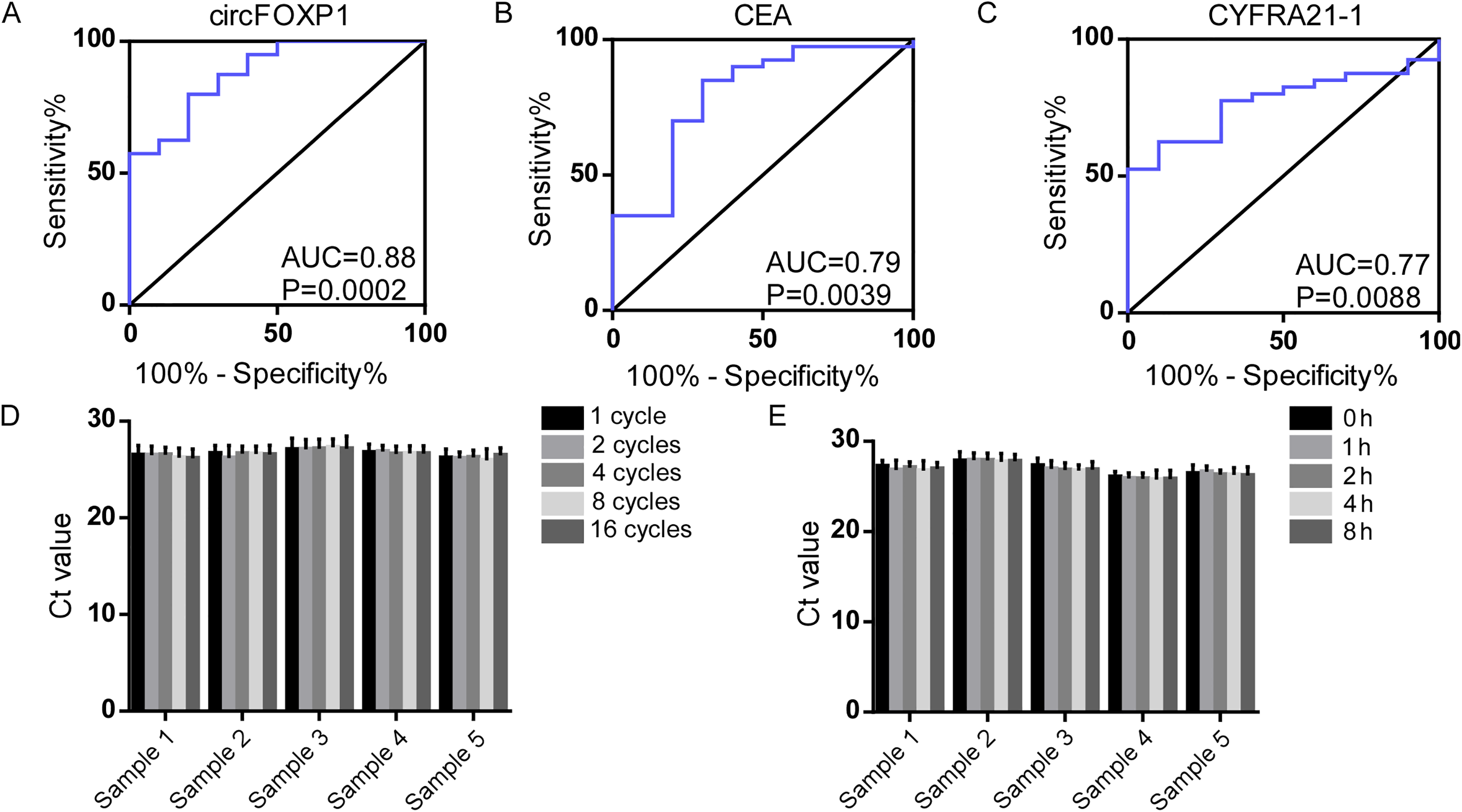
ROC curves analysis of circFOXP1 and traditional markers in NSCLC. (a) to (c) ROC curves analysis of circFOXP1 (a) CEA (b) and CYFRA21-1 (c) in distinguishing patients with NSCLC from healthy controls. (d) and (e) qRT-PCR assay analysis of cicrFOXP1 stability in the serum when subjected to 1, 2, 4, 8, and 16 cycles of freeze-thaw (d) and 0, 1, 2, 4, and 8 h incubation (e) at room temperature.
Validation of cicrFOXP1 stability
The stability of cicrFOXP1 is crucial as a potential tumor marker, and we next verified this property in serum. As shown in Figure 4(a) and (b), the Ct value of circFOXP1 in serum showed no significant changes when subjected to 1, 2, 4, 8, and 16 cycles of freeze-thaw and 0, 1, 2, 4, and 8 h incubation at room temperature (Figure 3(d) and (e)). The results reflected the proper stability of circFOXP1 in serum.
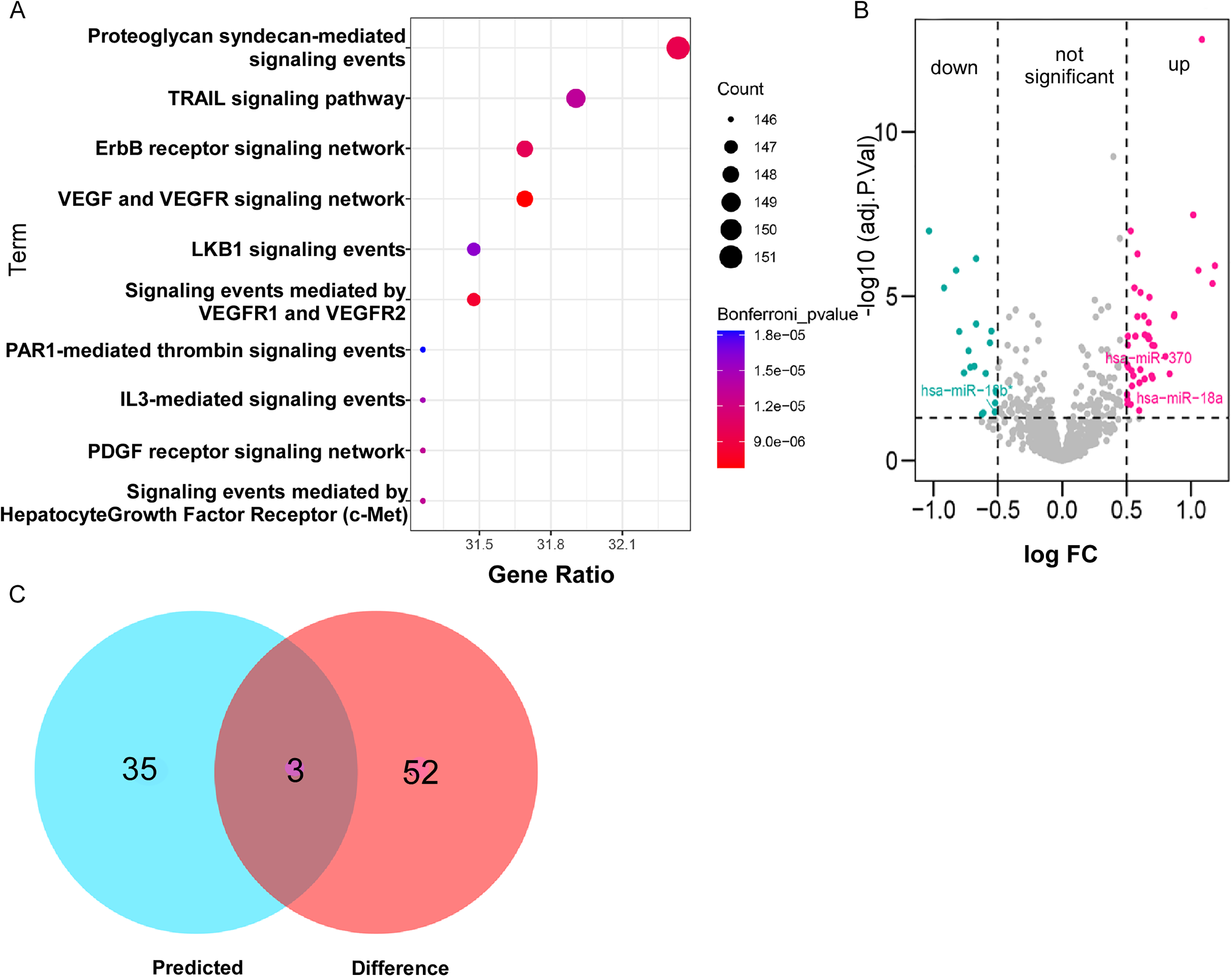
Analysis of circFOXP1-miRNA-mRNA interaction. (a) StarBase (http://starbase.sysu.edu.cn/starbase2/) and FunRich tool (version 3.1.3) prediction of circFOXP1 related miRNAs and pathways. (b) Gene expression microarray (GSE36681) (https://www.ncbi.nlm.nih.gov/gds/) was analyzed to screen differential miRNAs in NSCLC. (c) Venn diagram was conducted to show overlapping miRNAs.
The association of circFOXP1 expression and clinical characteristics in NSCLC
We further analyzed the correlation between circFOXP1 expression and the clinical characteristics of NSCLC. Clinical information of the patients was collected and is summarized in Supplementary Tables 2 and 3. These data show that circFOXP1 was significantly correlated with T stage and lymphatic metastasis, instead of sex, age and distant metastasis (Supplementary Tables 2 and 3). Further study was performed to detect the predictive value of circFOXP1 on the risk of NSCLC. Univariate analysis showed that circFOXP1 was the risk factor for NSCLC. Multivariate analysis indicated that circFOXP1 level was the independent risk factor for NSCLC when adjusting for age, gender, and other clinic factors (Supplementary Table 4). Collectively, these results identified that circFOXP1 was a potential biomarker for diagnosis of NSCLC.
circFOXP1 targeted miRNA-mRNA interaction
Against the backdrop that our aforementioned data suggested positive correlation between serum circFOXP1 and T stage with the addition of lymphatic metastasis in NSCLC. As reported in the previous study, Li et al. found that circFOXP1 promoted cell proliferation and metastasis in cells and tissues of lung adenocarcinoma, which was consistent with our research. 20 To explore possible molecular mechanisms of circFOXP1, we utilized the starBase database to predict the miRNAs that might bind to circFOXP1, and 38 miRNAs were identified. FunRich tool (version 3.1.3) was then applied to conduct enrichment analysis and to detect the possible biological pathways. Based on analysis, ErbB and vascular endothelial growth factor (VEGF) signaling pathways were supposed to be more relevant with circFOXP1 and significantly related to cell proliferation and metastasis (Figure 4(a)). To further explore differential miRNAs, data of the gene expression microarray (GSE36681) from the GEO database were utilized, and differentially expressed miRNAs in normal and NSCLC were obtained through GEO2R (adj. P value <0.05, absolute value of logFC >0.5) (Figure 4(b)). By integrating the miRNAs of the starBase prediction with GEO database analysis, three overlapping miRNAs, including hsa-miR-370-3p, hsa-miR-18a-5p, and hsa-miR-18b-5p, were ultimately identified to serve as the target molecules of circFOXP1 (Figure 4(c)). The target mRNAs of hsa-miR-370-3p, hsa-miR-18a-5p, and hsa-miR-18b-5p were obtained by overlapping the prediction results of the miRmap, microT, miRanda, PicTar, TargetScan and PITA databases, respectively. Cytoscape software (version 3.8.0) was used to visualize the circRNA-microRNA-mRNA interaction (Figure 5). Then, the expression of hsa-miR-370-3p, hsa-miR-18a-5p, and hsa-miR-18b-5p were validated in PC9 cells. As shown in supplementary Figure 1(a) to (f), after transfection with circRNA or si-circRNA, the level of hsa-miR-370-3p and hsa-miR-18a-5p was negatively affected (P < 0.05), while no change was found in the expression of hsa-miR-18b-5p (Supplementary Figure 1(a) to (f)). Interestingly, miR-370 and miR-18 were also reported to interact with the VEGF signaling pathway,21,22 which further verified our bioinformatics prediction and established the foundation for the following study. This bioinformatics analysis suggested that circFOXP1 might bind to hsa-miR-370-3p and hsa-miR-18a-5p, and be involved in VEGF signaling pathways that regulated proliferative and metastasis processes.
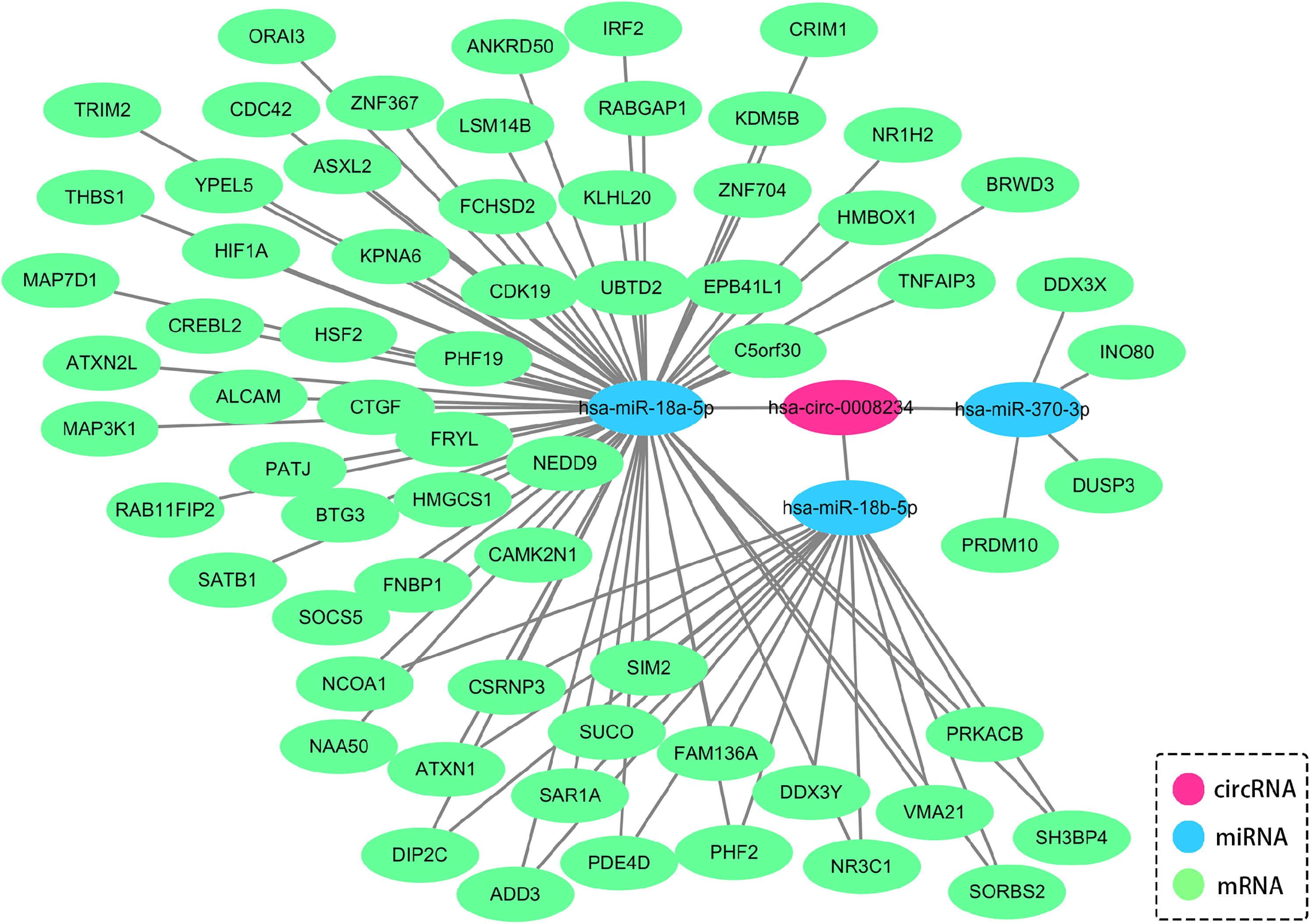
Cytoscape software analysis of circRNA-microRNA–mRNA interaction (version 3.8.0).
Discussion
In this study we screened five candidates with different circRNAs in NSCLC patients compared with healthy controls. The qRT-PCR assay showed that serum circFOXP1 exhibited differential expression, while other circRNAs did not reveal this distinction. Based on ROC curves analysis, we found that serum circFOXP1 discriminated NSCLC patients from controls with superior sensitivity and specificity when compared with conventional tumor markers. Molecular mechanism prediction suggested that circFOXP1 might bind to two critical miRNAs and be involved in VEGF signaling pathways that regulated proliferative and metastasis processes. Collectively, our research shows that circFOXP1 may serve as a potential biomarker for the diagnosis of NSCLC.
NSCLC is the leading cause of malignancy-related mortality in the world, so increasing the ratio of diagnosis is the significant method to ameliorate this mortality. 23 Imaging detection, such as low-dose spiral computed tomography (CT), is the common technique to access to tiny nodules in NSCLC, while exorbitant expense and a high false positive rate still restrict its application. 24 Semi-automated quantification methods with radiomics could reduce the false positive results significantly.25–27 Spanu et al. found that TF SPECT was a highly accurate diagnostic method in the detection of intrathoracic malignant lesions. 28 Bianconi et al. discovered that deep learning is a viable approach for semi-automated segmentation of pulmonary nodules on CT scans. 26 However, these approaches presented some challenges. For semi-automated quantification methods, much larger datasets were needed to complete the calculation. Image acquisition had a lack of standardization, strong heterogeneity and variability in scanner models, acquisition protocols and reconstruction settings. Moreover, the cost is relatively expensive and the application is not very extensive in small-scale hospitals. 25 With the advantages of rapid detection, minimal invasion, low risk, and easy access, the potential of liquid biopsy technology in tumor diagnosis and detection is gradually emerging. 29 The sensitivity and specificity of conventional diagnostic markers for NSCLC, such as CYFRA21-1 and CEA, are not so satisfactory. According to the previous reports, the AUCs of CYFRA 21-1 and CEA were under 0.80.30,31 In this experiment, serum CEA and CYFRA21-1 served as control markers, and we found that the AUCs of NSCLC patients and the control group was 0.79 and 0.77, respectively, which was consistent with the previous study.
circRNAs are circular non-coding RNA (ncRNA) molecules that were formed by the covalent binding of 3′ terminal end and 5′ terminal end after reverse splicing, which was characterized by stable structure, conservative sequence, and excellent specificity. Qin et al. found a dramatic discrepancy of hsa_circ_0001649 in hepatocellular cells (HCC) and adjacent liver tissues, indicating its potential as the diagnostic biomarker. 32 Yao et al. reported that circRNA 100876 was closely correlated with lymph node metastasis and tumor stage, which served as potential biomarker for NSCLC. 33 These studies provided new insights into the diagnosis of NSCLC and supported circRNA to be the novel biomarker for the diagnosis of NSCLC. In this study, we screened dysregulated circRNAs in NSCLC and found that circFOXP1 was the candidate circRNA for diagnosis. CircFOXP1, as the recently discovered RNA, was reported to be upregulated in several tumors, such as lung cancer. Li et al. found that circFOXP1 promoted cell proliferation of lung cancer by binding with miR-185-5p/Wnt1. 20 Nevertheless, this research is the first time its diagnostic potential in NSCLC has been detected.
In our analysis of patients with NSCLC and controls, we have demonstrated that serum circFOXP1 was significantly increased in NSCLC and exerted superior diagnostic efficacy when compared with the healthy controls. Through univariate and multivariate analyses, circFOXP1 was found to be the independent detection molecule for NSCLC when adjusting for other factors. Analysis of circFOXP1 expression and its clinical characteristics. circFOXP1 was significantly correlated with T stage and lymphatic metastasis, instead of distant metastasis, which may be due to the limited sample size. In addition, the stability analysis showed that the Ct value of circFOXP1 remained stable under extreme conditions. Furthermore, the possible molecular mechanism was preliminarily predicted and explored. This evidence supported that circFOXP1 has great potential to be the diagnostic biomarker. However, this study still needs in-depth research from the following aspects: (a) The number of included cases was limited—a larger sample size is needed to confirm the clinical value of circFOXP1 in the later study; (b) multi-centric and prospective research needs further implementation to prevent potential selection bias.
In summary, circulating circFOXP1 was significantly overexpressed in NSCLC and closely correlated with T stage. ROC curves showed that the AUC of circFOXP1 was 0.8825 in distinguishing NSCLC and controls, which was preferable to CEA and CYFRA21-1. circFOXP1 was supposed to be the independent biomarker for diagnosis with reliable stability. Overall, serum circFOXP1 is expected to be a new clinical biomarker for NSCLC, and our study provides an important aspect for the diagnosis and prospective therapeutic target for NSCLC.
Supplemental Material
sj-docx-1-jbm-10.1177_17246008211073151 - Supplemental material for CircFOXP1: A novel serum diagnostic biomarker for non-small cell lung cancer
Supplemental material, sj-docx-1-jbm-10.1177_17246008211073151 for CircFOXP1: A novel serum diagnostic biomarker for non-small cell lung cancer by Yirong Luo, Qichao Zhang, Bo Lv, Yanyan Shang, Juan Li, Lina Yang, Zhiwu Yu, Kai Luo, Xiaoyan Deng, Ling Min and Ting Zhu in The International Journal of Biological Markers
Supplemental Material
sj-tif-2-jbm-10.1177_17246008211073151 - Supplemental material for CircFOXP1: A novel serum diagnostic biomarker for non-small cell lung cancer
Supplemental material, sj-tif-2-jbm-10.1177_17246008211073151 for CircFOXP1: A novel serum diagnostic biomarker for non-small cell lung cancer by Yirong Luo, Qichao Zhang, Bo Lv, Yanyan Shang, Juan Li, Lina Yang, Zhiwu Yu, Kai Luo, Xiaoyan Deng, Ling Min and Ting Zhu in The International Journal of Biological Markers
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author contributions
Yirong Luo, Qichao Zhang and Bo Lv contributed equally to this study.
Funding
The authors disclosed receipt of the following financial supportfor the research, authorship, and/or publication of this article: This study was supported by Youth Program of National Natural Science Foundation of China (No. 81902991); Guangzhou Science and Technology Plan Project (No. 202102021056); Medical Scientific Research Foundation of Guangdong Province, China (No. 2020112314313344); Guangzhou Health Science and Technology Project (No. 20221A010061).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
