Abstract
Ovarian carcinoma ranks fifth in the leading causes of cancer-relevant deaths among the female, with the highest fatality rate in all gynecological malignant tumors and the rising incidence worldwide. Mounting evidence has unveiled that lncRNAs are implicated in the tumorigenesis and cancer development. Several studies have proven the carcinogenic role of SNHG8 in various malignancies, but the physiological functions of SNHG8 in ovarian carcinoma need more detailed explanations. The present study certified that inhibition of SNHG8 executed suppressive activities in ovarian carcinoma by obstructing cell proliferation, migration, EMT process and stemness as well as driving cell apoptosis. Moreover, SNHG8 bound with CAPRIN1 and positively modulated the expression of CAPRIN1. Further experiments manifested that CTNNB1 and Axin1 displayed a binding affinity with CAPRIN1. Knockdown of CAPRIN1 promoted the mRNA degradation of CTNNB1 and Axin1. Finally, we corroborated that CTNNB1 (or Axin1) ectopic expression or activation of Wnt/
Introduction
Ovarian carcinoma ranks fifth in the leading causes of cancer-relevant deaths among the female, with the highest fatality rate in all gynecological malignant tumors and the rising incidence worldwide [1, 2]. For ovarian carcinoma patients, late diagnosis generally occurs due to insufficient specific symptoms and screening methods [3, 4]. Once diagnosed, patients with ovarian carcinoma often miss out on the surgery [5]. Despite the continual improvement in ovarian carcinoma therapies, the survival remains limited in advanced ovarian carcinoma patients, and the survival rate within five years of diagnosis of sufferers with distant metastasis is only 28.9% [6]. In consequence, it is imperative to conduct more research to understand the latent mechanisms of ovarian carcinoma.
Long noncoding RNAs (lncRNAs) are defined as a subset of noncoding RNAs which are longer than 200 nucleotides and featured with limited or no protein-coding capacity [7]. Accumulating paper has proved that lncRNAs are involved in the tumorigenesis and cancer development by regulating biological activities, such as cell proliferation, invasion, apoptosis and differentiation [8, 9]. For example, lncRNA NEAT1 facilitates ovarian carcinoma cell metastasis through targeting miR-382-3p/ROCK1 axial [10]. Novel lncRNA OTUD6B-AS1 correlates with unfavorable prognosis and retards the proliferation of clear cell renal cell carcinoma [11]. LncRNA SNHG20 facilitates cell proliferation and apoptosis of glioma by regulation of PTEN/PI3K/AKT signaling pathway [12]. According to recent investigations, the carcinogenesis of SNHG8 has been certified in multiple malignancies, including non-small-cell lung cancer [13], gastric carcinoma [14], pancreatic adenocarcinoma [15]. Nevertheless, the physiological functions of SNHG8 in ovarian carcinoma need more detailed explanations.
Wnt signaling plays a pivotal role during organ development and tissue homeostasis in several diseases, including tumors [16, 17]. It is well accepted that Wnt signaling is diversified into two pathways, namely the canonical and noncanonical pathway [18]. Besides, LEF-1 and
Therefore, the purpose of current work was to identify the functions of SNHG8 on the cellular process of ovarian carcinoma cells and the mechanism through which SNHG8 might regulate Wnt/
Materials and methods
Cell culture and transfection
Human normal fallopian tube epithelial cell line FTE187 and ovarian carcinoma cell lines (SKOV3, ES2, CaOV3 and HG-SOC) were offered by American Type Culture Collection (ATCC, USA). All cells were cultivated in DMEM (Gibco, Carlsbad, CA, USA) or RPMI-1640 medium (Invitrogen, Thermo Fisher Scientific, USA) complemented with 10% fetal bovine serum (Invitrogen, USA), 100
Cell counting Kit-8 (CCK-8) assay
Cell viability was estimated by Cell Counting Kit-8 (CCK-8; Beyotime Institute of Biotechnology, Shanghai, China). Transfected cells were cultured on a 96-well plate at a density of 2
5-ethynyl-2’-deoxyuridine (EdU) assay
An EdU incorporation assay kit (Ribobio, China) was employed for evaluation of cell proliferation. In brief, 2
Quantitative real-time polymerase chain reaction (qRT-PCR)
Total RNA extraction from cells was carried out with TRIzol (Invitrogen) under the recommendations of the manufacturer. The PrimeScript RT reagent kit (Promega, USA) was applied to synthesize cDNA. Then, PCR was conducted with a SYBR Green PCR kit (Takara, Japan) in an ABI 7500 real-time PCR system (Applied Biosystems, USA). The expression of genes was quantified by the 2
Wound-healing assay
Forty-eight h after transfection, cells were seeded into 6-well plate and grown to approximately 85% confluence. Then, the sterile pipette was applied to scratch the bottom of the culture plate. After washing by PBS buffer, cells were added with serum-free DMEM and maintained at 37
Cell migration
Cell migration was detected by transwell assay employing transwell chambers (8
Sphere formation assay
As previously described [24], cell self-renewal capacity was evaluated by this assay. The single cell suspension was prepared by enzymatic disassociation, filtrated using a 40
Flow cytometry analysis
Stem cell marker CD133 was measured in transfected cells by flow cytometry analysis as previously described [25].
Caspase-3 activity assay
The caspase-3 activity kit (Beyotime Institute of Biotechnology) was adopted for the analysis of caspase activity. Total protein from cells was treated with caspase-3 substrate Ac-DEVD-pNA at 37
Western blot
RIPA lysis buffer (Beyotime, China) supplemented with protease inhibitors was utilized to extract proteins from cells, and the BCA protein assay (Pierce, USA) was performed to determine protein concentrations. Equal amounts (40
RNA precipitation
To figure out the possible proteins that bind with SNHG8, biotin-labeled SNHG8 RNA probe was employed in RNA precipitation assay. Then PierceTM Silver Stain Kit (Fisher Scientific) was utilized to conduct silver staining or western blot as previously described [26].
RNA immunoprecipitation (RIP)
The Magna RIP RNA-Binding Protein Immunoprecipitation Kit (Millipore, Bedford, MA, USA) was applied to conduct RIP assay in light of the suggestion of the manufacturer. Cell lysates were incubated in RIP buffer supplemented with magnetic beads coated with anti-CAPRIN1 (Abcam) antibody or negative control IgG (Millipore). The immunoprecipitated RNA was obtained with proteinase K and purified RNA was analyzed by qRT-PCR.
Silencing of SNHG8 impedes cell proliferation and contributes to cell apoptosis. (A) SNHG8 expression was analyzed by qRT-PCR in ovarian carcinoma cells (SKOV3, ES2, CaOV3 and HG-SOC) and normal cells FTE187. (B) Transfection efficiency was verified by qRT-PCR. (C–D) Cell proliferation of SKOV3 and CaOV3 cells was tested byCCK-8 and EdU assays. (E) Caspase-3 activity assay was applied to analyze cell apoptosis. 
SNHG8 suppression restrains the migration, EMT and stemness of ovarian carcinoma. (A) The migrated ability of SKOV3 and CaOV3 cells was assessed by wound healing assay. (B) Transwell assay for cell migration in transfected cell lines. (C) Western blot was applied to estimate the expression of EMT process-related proteins. (D–E) The sphere formation ability was detected by sphere formation assay and western blot. (F) Percentage of CD133 in SNHG8-downregulated cells was measured. 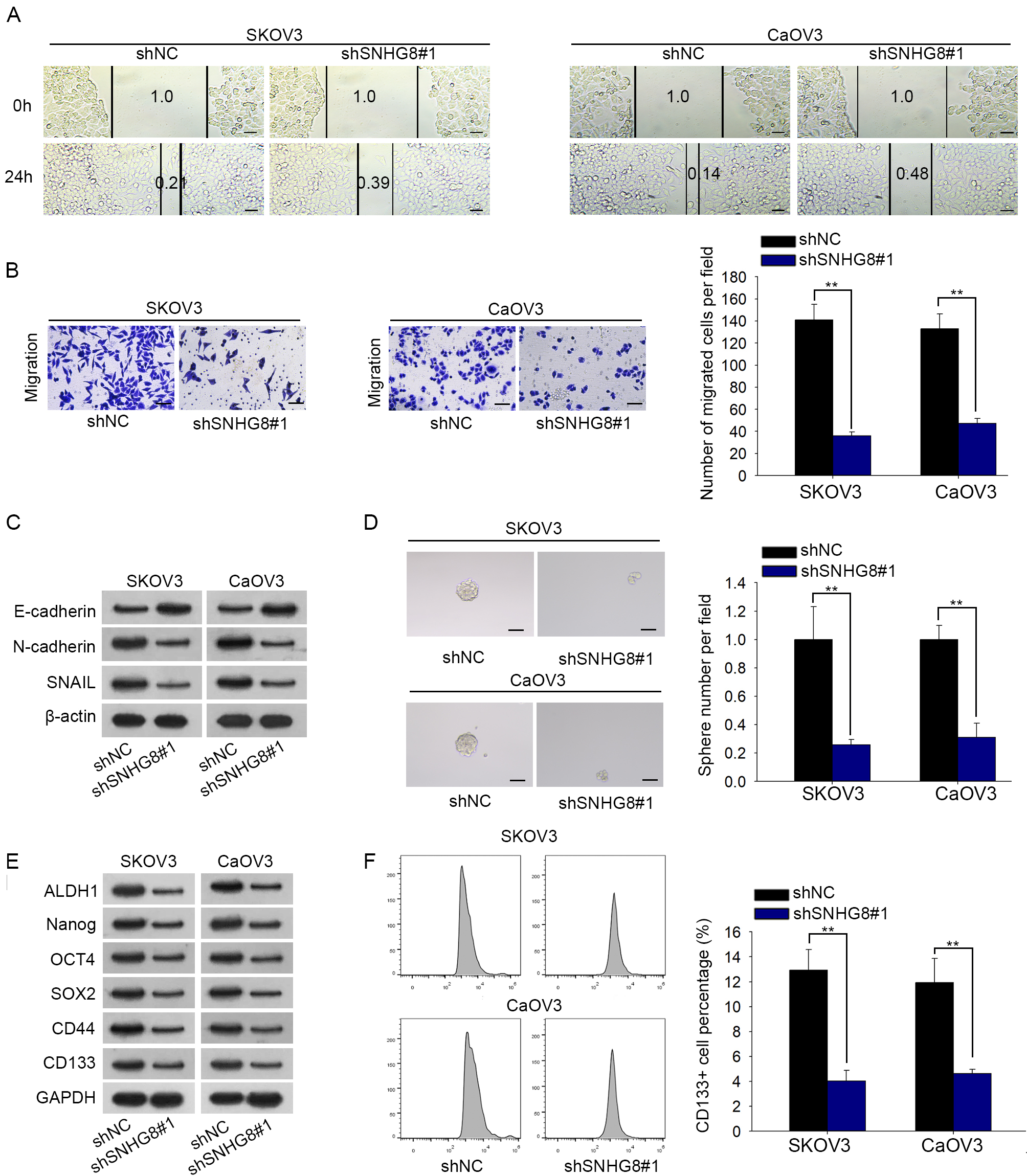
Full-length sense and antisense of SNHG8 and CTNNB1 RNA were transcribed using MEGAscript
Analysis of mRNA stability
After transfection, cells were collected and supplemented with the transcriptional inhibitor 5
Statistical analysis
All results were shown as the mean
Results
Silencing of SNHG8 impedes cell proliferation and contributes to cell apoptosis
In order to estimate the physiological function of SNHG8 in ovarian carcinoma, we first detected the expression of SNHG8 in normal cells and ovarian carcinoma cells. qRT-PCR analysis demonstrated that SNHG8 was pronouncedly elevated in ovarian carcinoma cells compared to normal FTE187 cells (Fig. 1A). Thereafter, SKOV3 and CaOV3 cells were transfected with shRNAs targeting SNHG8, leading to the overt downregulation of SNHG8 expression (Fig. 1B). CCK-8 and EdU assays were adopted for evaluation of cell proliferation in response to SNHG8 inhibition and knockdown of SNHG8 resulted in a remarkable decrease of cell proliferation (Fig. 1C and D). In concert with these findings, cell apoptosis rate was notably enhanced in the face of SNHG8 depletion (Fig. 1E). In a word, our results indicate that inhibition of SNHG8 exerts inhibitive functions in ovarian carcinoma by retarding cell proliferation and inducing cell apoptosis.
SNHG8 regulates the expression of CAPRIN1 by binding to it. (A) RNA-pull down assay confirmed the binding between SNHG8 and CAPRIN1. (B) The level of CAPRIN1 in cell lines was examined by qRT-PCR assay. (C) The association between SNHG8 and CAPRIN1 was confirmed by RIP assay. (D–E) The mRNA and protein expression of CAPRIN1 was measured by qRT-PCR and western blot respectively. 
We subsequently investigated the impacts of SNHG8 on cell migration. The wound healing assay manifested that SNHG8 downregulation caused slower scratch wound closure in SKOV3 and CaOV3 cells, suggesting that depletion of SNHG8 inhibited the migration of ovarian carcinoma cells (Fig. 2A). This result was further confirmed by the transwell assay (Fig. 2B). Considering that epithelial-mesenchymal transition (EMT) process plays a vital role in cell metastasis progression of malignancy [27], we evaluated the EMT process in transfected cells. In western blot assay, an evident increase in the expression of epithelial marker E-cadherin and an obvious diminution in the expression of mesenchymal markers (N-cadherin and SNAIL) was observed when SNHG8 was silenced (Fig. 2C). Furthermore, given that cancer cells undergoing EMT process exhibit stem cell properties [28], sphere formation assay was conducted. Results presented that inhibition of SNHG8 repressed the sphere formation ability (Fig. 2D). Concordant with these data above, we discovered that the expression levels of stem factors (ALDH1, Nanog, SOX2, OCT4, CD44, CD133) were dropped due to SNHG8 knockdown (Fig. 2E). Additionally, stem cell marker CD133 was evaluated in SNHG8-downregulated cells through flow cytometry analysis. CD133 positivity was obviously decreased after depletion of SNHG8 (Fig. 2F). On the whole, we draw a conclusion that downregulation of SNHG8 impairs ovarian carcinoma cell migration, EMT process and stemness.
SNHG8 regulates the expression of CAPRIN1 by binding to it
Pull-down silver staining was carried out to explore the proteins that CAPRIN1 could interact with SNHG8 (Fig. S1A). By the assistance of starBase database, we found that CAPRIN1 may possess the potential binding sites with SNHG8. Then western blot assay elucidated that CAPRIN1 was pulled down by SNHG8 and had no response to antisense SNHG8 (Fig. 3A). As CAPRIN1 expression in ovarian carcinoma cells was upregulated (Fig. 3B), CAPRIN1 was chosen for in-depth study on the mechanism of SNHG8. Besides, RIP assay showed that SNHG8 was predominantly abundant in CAPRIN1 immunoprecipitate compared with IgG group (Fig. 3C). Given that CAPRIN1 bound with SNHG8, we explored the association between SNHG8 and CAPRIN1. Results of qRT-RCR and western blot proofed that SNHG8 suppression led to a significant attenuation of CAPRIN1 mRNA and protein levels in SKOV3 and CaOV3 cells (Fig. 3D and E). Jointly, SNHG8 binds with CAPRIN1 and positively modulates the expression of CAPRIN1.
CAPRIN1 promotes the stability of CTNNB1. (A) qRT-PCR detected CTNNB1 level in ovarian carcinoma cells and normal fallopian tube epithelial cells. (B–C) RIP and RNA-pull down assays examined whether CTNNB1 bound with CAPRIN1. (D–E) The expression of CAPRIN1 and CTNNB1 was quantified by qRT-PCR. 
SNHG8 facilitates ovarian carcinoma via activation of Wnt/
To further understand the molecular mechanism of SNHG8, the proteins that were pulled down by SNHG8 were submitted to GO analysis. The results suggested that SNHG8 was highly possible to influence Wnt signaling pathway and mRNA stability regulation (Fig. S1B). Given that CTNNB1 and Axin1 are Wnt signaling pathway-related genes and were predicted by starBase to be regulated by CAPRIN1 (Fig. S1C), therefore, CTNNB1 and Axin1 were chosen to be studied. Further, the expression of CTNNB1 and Axin1 in ovarian carcinoma cells was upregulated in comparison with that in normal FTE187 cells (Fig. 4A, Fig. S1D). RIP and RNA-pull down experiments were then carried out to verify the relationship between CAPRIN1 and CTNNB1 or Axin1. Our findings revealed that CTNNB1 and Axin1 were enriched by anti-CAPRIN1 antibody (Fig. 4B, Fig. S1E). Afterwards, CAPRIN1 and Axin1 can be only detected in the pull-down complex of CTNNB1 but not in the antisense CTNNB1 group, implying that CTNNB1 displayed a binding affinity with CAPRIN1 (Fig. 4C, Fig. S1F). Considering that CAPRIN1 is known as a RNA-binding protein which is participated in the regulation of target genes [29], we assessed the influences of CAPRIN1 on CTNNB1 and Axin1 expression. CAPRIN1 expression was knocked down and qRT-PCR confirmed the efficiency of transfection (Fig. 4D). Subsequently, ovarian carcinoma cells were treated with RNA synthesis inhibitor Actinomycin D and results illuminated that the mRNA degradation of CTNNB1 and Axin1 were quicker owing to CAPRIN1 depletion, implying that absence of CAPRIN1 restrained the CTNNB1 mRNA stability (Fig. 4E, Fig. S1G). Furthermore, CAPRIN1 was overexpressed in SKOV3 and CaOV3 cells by the transfection of pcDNA3.1/CAPRIN1 (Fig. S1H). Finally, it was validated that the suppressed mRNA stability, mRNA and protein level of CTNNB1 and Axin1 in sh-SNHG8#1-transfected cells were reversed by CAPRIN1 overexpression (Fig. S1I–J). Taken together, CAPRIN1 inhibition enhances the mRNA degradation of CTNNB1 and Axin1.
SNHG8 facilitates ovarian carcinoma via activation of Wnt/
-catenin pathway
Given that CTNNB1 is a protein that related with Wnt/
Discussion
Ovarian carcinoma ranks as the eighth most prevalent malignancy and becomes most fatal gynecological cancer in women [30]. Ovarian carcinoma is disproportionately lethal in view of lacking sophisticated early diagnosis approaches [31]. Although there are advances in treating ovarian carcinoma, the 5-year survival rate of patients is still unsatisfactory [32]. Hence, our study intended to explore the potential mechanism of ovarian carcinoma so as to find out a potent target for better therapy.
It is evident from increasing investigations that lncRNAs function as core participants in regulation of biological processes in various tumors, including ovarian carcinoma [33, 34, 35]. Mounting evidence has implied the oncogenic functions of SNHG8 on several malignancies. For instance, lncRNA SNHG8 enahnces the malignancy of hepatocellular carcinoma by sponging miR-149-5p [36]. LncRNA SNHG8/miR-152/c-MET axis facilitates the development of endometrial carcinoma [37]. lncRNA SNHG8 acts as an oncogene in Epstein-Barr virus-associated gastric carcinoma [38]. Nonetheless, the impacts of SNHG8 on ovarian carcinoma remain to be probed. Our work illustrated the upregulated SNHG8 in ovarian carcinoma cells and silencing of SNHG8 elicited suppressive influences by hampering cell proliferation, migration, EMT process and stemness whereas promoting cell apoptosis.
Cell cycle associated protein 1 (CAPRIN1), a 709 amino acid protein located in the cytoplasm, is a member of the conserved protein family observed throughout vertebrates [39, 40]. And it was detected to be the most enriched protein to be pulled down by biotinylated SNHG8. Previous paper has delineated the critical role of CAPRIN1 in the development of diverse cancers. For example, enhanced caprin1 level could predict unfavorable prognosis in hepatocellular carcinoma [41]. Caprin-1 exerted its oncogenic functions on osteosarcoma tumor [42]. microRNA-223 suppresses the malignancy of breast cancer via targeting Caprin-1 [43]. Results from experiments disclosed that CAPRIN1 was upregulated in ovarian carcinoma and bound with SNHG8. Furthermore, SNHG8 positively modulated the expression of CAPRIN1. More importantly, GO analysis of the proteins that were pulled down by SNHG8 implied that SNHG8 might regulate Wnt signaling pathway and mRNA stability. Besides, CAPRIN1 has been reported to act as an RNA-binding protein [44]. Considering that CTNNB1 and Axin1 are Wnt signaling pathway-related genes and were probed by starBase to be regulated by CAPRIN1, therefore, the interaction between CAPRIN1 and CTNNB1 or Axin1 was further explored. Our findings manifested that CAPRIN1 displayed a binding affinity with CTNNB1 and Axin1 as well as promoted the mRNA stability of CTNNB1 and Axin1.
Wnt/
In conclusion, this study clarified the carcinogenic role of SNHG8 in ovarian carcinoma and that SNHG8 enhances the malignancy of ovarian carcinoma via activation of Wnt/
Footnotes
Acknowledgments
Authors deeply appreciated all lab members.
Conflict of interest
None.
Supplementary data
(A) Pull-down silver staining was carried out to explore the proteins that could interact with SNHG8. (B) GO analysis of the proteins that were pulled down by SNHG8. (C) The combination between CAPRIN1 and CTNNB1 (or Axin1) was predicted by starBase website. (D) qRT-PCR analyzed Axin1 expression in ovarian carcinoma cells and normal fallopian tube epithelial cells. (E–F) RIP and RNA-pull down assays disclosed the affinity between Axin1 and CAPRIN1. (G) The expression of Axin1 after treating with ActD was quantified by qRT-PCR. (H) The overexpression efficiency of CAPRIN1 was verified by qRT-PCR. (I–L) The mRNA stability, mRNA and protein level of CTNNB1 or Axin1 in different transfected cells were investigated determined by qRT-PCR and western blot assay. 
(A) qRT-PCR analysis evaluated overexpression efficiency. (B) Western blot determined the protein levels of key target genes in Wnt/
