Abstract
Prolyl 4-hydroxylase, beta polypeptide (P4HB) protein has been found to be associated with tumorigenesis in many types of tumor, However, the relationship between P4HB and clear cell renal cell carcinoma (ccRCC) has not been clarified. In this study, we focus on the correlation between P4HB expression and ccRCC. Through the Cancer Genome Atlas (TCGA) database, Gene Expression Omnibus (GEO) database, our database and immunohistochemical (IHC) staining. Compared with adjacent normal tissues, both the mRNA and protein levels of P4HB in ccRCC tissues were enhanced. The Kaplan-Meier survival analysis showed that high expression of P4HB is correlated with poor prognosis in both TCGA database and our own database. Multivariate survival analysis and Univariate analysis showed that P4HB expression and age are significantly correlative with poor prognose. All the results indicated that P4HB is correlated with poor prognosis in human clear cell renal cell carcinoma.
Introduction
Renal cell carcinoma is the most common malignancy originating in the kidney and one of the most lethal genitourinary cancer [1]. At present, clear cell renal cell carcinoma is the major histological subtype of renal cell carcinoma, accounting for 80%–90% of cases [2, 3]. Although plenty of key genes found in many studies may contribute to tumorigenesis and tumor progression of ccRCC, few genes could be the target for diagnosis or treatment. Earlier detection of RCC and rising life expectancy of cancer survivors places a greater emphasis on preservation of renal function after cancer resection and during systemic therapy [4, 5]. P4HB are related to some physiological processes such as RNA binding, protein disulfide isomerase activity, integrin binding and so on [6, 7, 8]. Moreover, P4HB plays an important role in endoplasmic reticulum protein processing pathway which have been linked to solid pseudopapillary tumor of the pancreas [9]. It is also correlated with malignant melanomas, endometrial carcinoma and gastric cancer [10, 11, 12]. Hence, P4HB has been showed to play an important role in different cancers, However, the correlation between P4HB and ccRCC is unknown. In this study, we systematically investigate the expression level and prognostic value in TCGA database, GEO database and our own database. We found that P4HB is correlated with poor prognosis in human clear cell renal cell carcinoma.
Materials and methods
Extraction of Clinical and gene expression data from ccRCC databases
The ccRCC datasets of this study were extracted from GEO database (
Gene set enrichment analysis (GSEA)
GSEA software was downloaded from Broad Institute (
Patients and specimens
The study consisted of 318 samples from 159 patients diagnosed with ccRCC. All the patients had a renal resection at the Peking University First Hospital between June 2008 and January 2011. The histological diagnosis was made on sections stained with hematoxylin and eosin, according to the WTO classification guidelines. Clinical and histopathological data including histopathological diagnosis and tumor grade were extracted from medical records. The study was performed with the approval of the Ethics Committee of Peking University First Hospital. Written informed consent was given by the patients for their information and specimens were stored in the hospital database and used for research.
Immunohistochemistry
Tissue sections were prepared from the paraffin-embedded tissue samples. Immunostaining was performed using the two-step detection kit (Zsbio PV-9000, China). The sections were deparaffinized in xylene, rehydrated in a graded alcohols series, and then boiled in citrate buffer (PH 6.0) for 30 minutes in an autoclave. Endogenous peroxidase was blocked by incubation in 3% H
Evaluation of immunostaining intensity
Two independent experienced investigators examined all tumor slides. Each slide was examined at five random views and 100 cells were observed per view at
Cell culture and transfection
HK-2, 786-O, 769-P, Caki-1, ACHN, A498, OSCR-2 and KETR3 were obtained from American Type Culture Collection (Manassas, VA, USA). Cells were cultured in DMEM (Invitrogen, Carlsbad, CA, USA) or 1640 (Invitrogen, Carlsbad, CA, USA) containing 10% fetal calf serum (Invitrogen). Cells were cultured in 10 mm culture dishes and were passaged every two days with 0.25 trypsin (sigma).
Quantitative real-time PCR
Total RNAs of 20 paired clinical samples were obtain from 20 patients diagnosed by Peking University First Hospital, and used according to the manufacturer’s protocol. Briefly, quantitation of all gene transcripts was done by qPCR using SYBR Premix ExTaq kit (Takara) and used GAPDH as a normalizing control. The primer pairs used were as follows: P4HB forward primer: 5’-AGGCTGATGACATCGTGAACT-3’, P4HB reverse primer: 5’-GGTATTTGGAGAACAC GTCACTG-3’. Each reaction was performed four times, and 2
The mRNA level of P4HB in ccRCC patients based on GEO database. The mRNA expression of P4HB in ccRCC tissues and adjacent normal renal tissues were compared. Nine mRNA datasets were employed including: GSE4282, GSE46699, GSE53757, GSE15641, GSE68417, GSE14994, GSE40435, GSE71963, and GSE76351.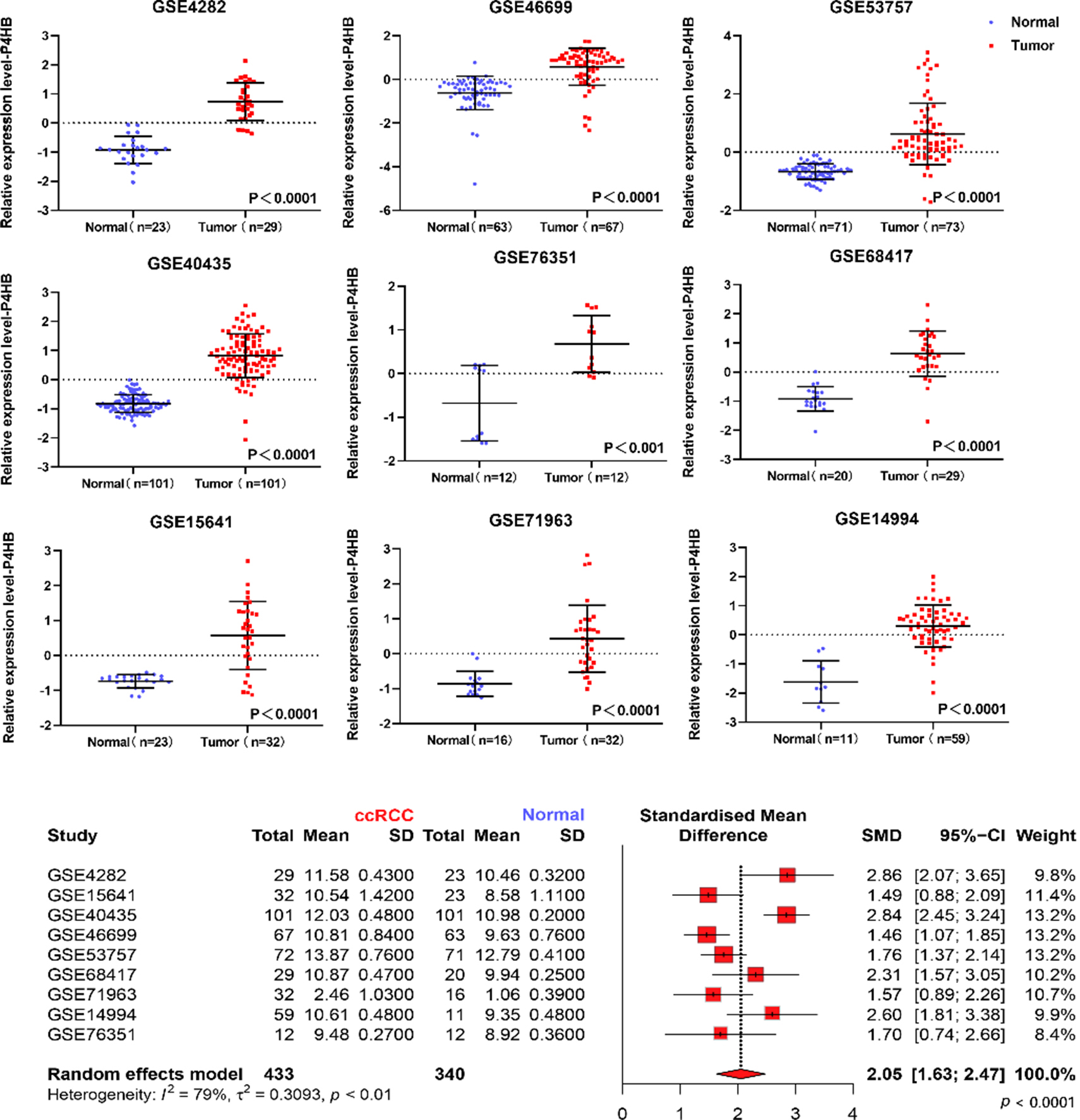
The mRNA level of P4HB in ccRCC based on our own database and TCGA database. The mRNA level of 20 paired clinical samples was shown (a). The mRNA expression of P4HB in ccRCC tissues and adjacent normal renal tissues were compared in 20 paired samples (b) and TCGA database (c).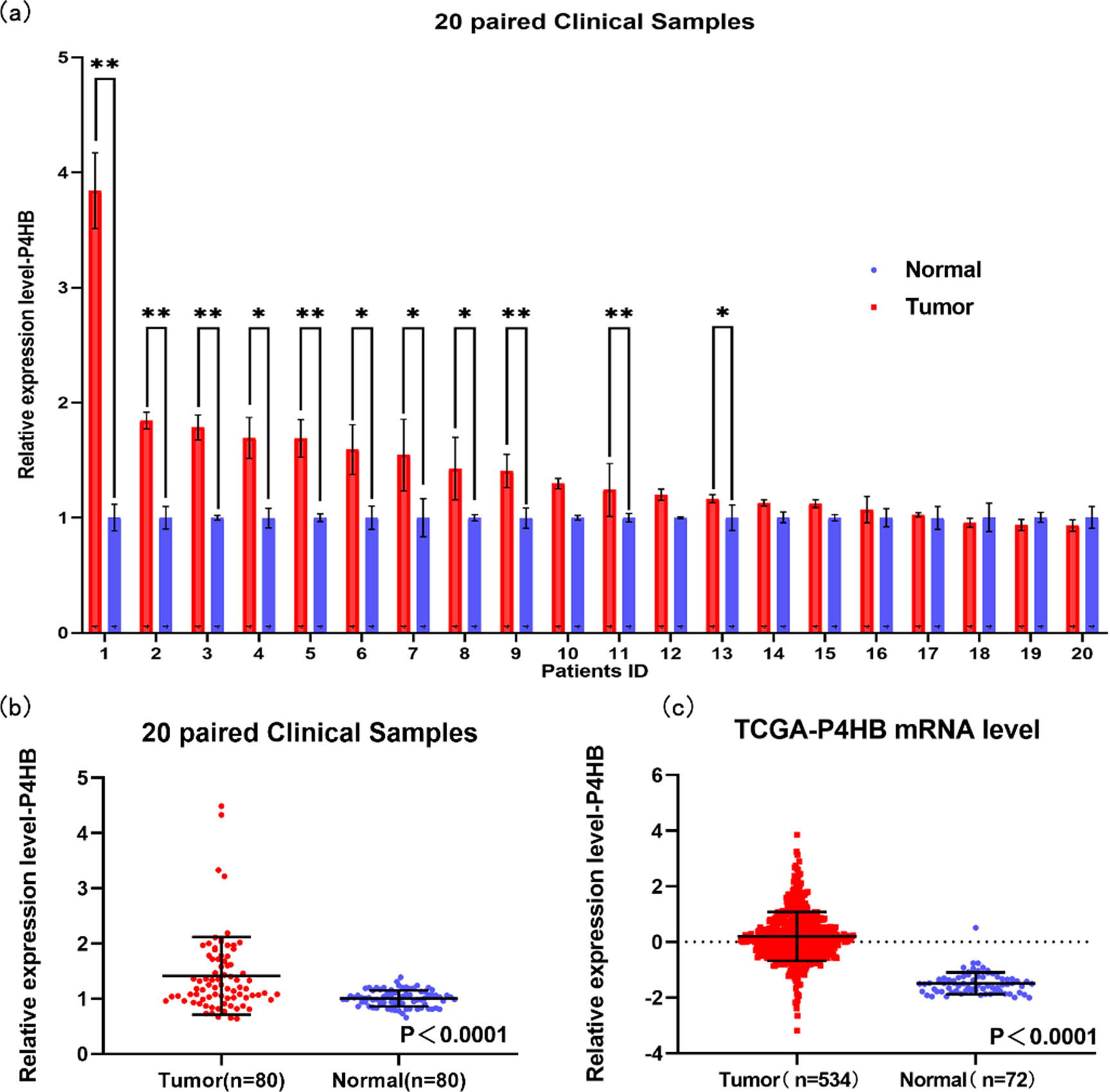
Total proteins from cells were extracted in NP-40 Lysis Buffer and quantified using the BCA method. Of the protein, 50
Statistical analysis
All data were analyzed by using SPSS 24.0. In this study, the student’s t-test was used to calculate the mRNA expression level in ccRCC tissues and adjacent normal renal tissues. Moreover, the Mann-Whitney U test was applied to compare the expression of the P4HB in terms of different clinicopathological characteristics including pathologic N and pathologic M. The one-way ANOVA was used to compare the expression of the P4HB in terms of different clinicopathological characteristics including pathologic stage and pathologic T. Kaplan-Meier method was used to calculate the survival analysis. Multivariate survival analysis and Univariate analysis were performed by using Cox model. All p values are based on a two-sided statistical analysis, and
Univariate analysis and multivariate analysis overall survival in TCGA database
Univariate analysis and multivariate analysis overall survival in TCGA database
Representative images of P4HB expression interpreted by immunohistochemistry as Negative (a), Weak (b), Moderate (c) and Strong (d).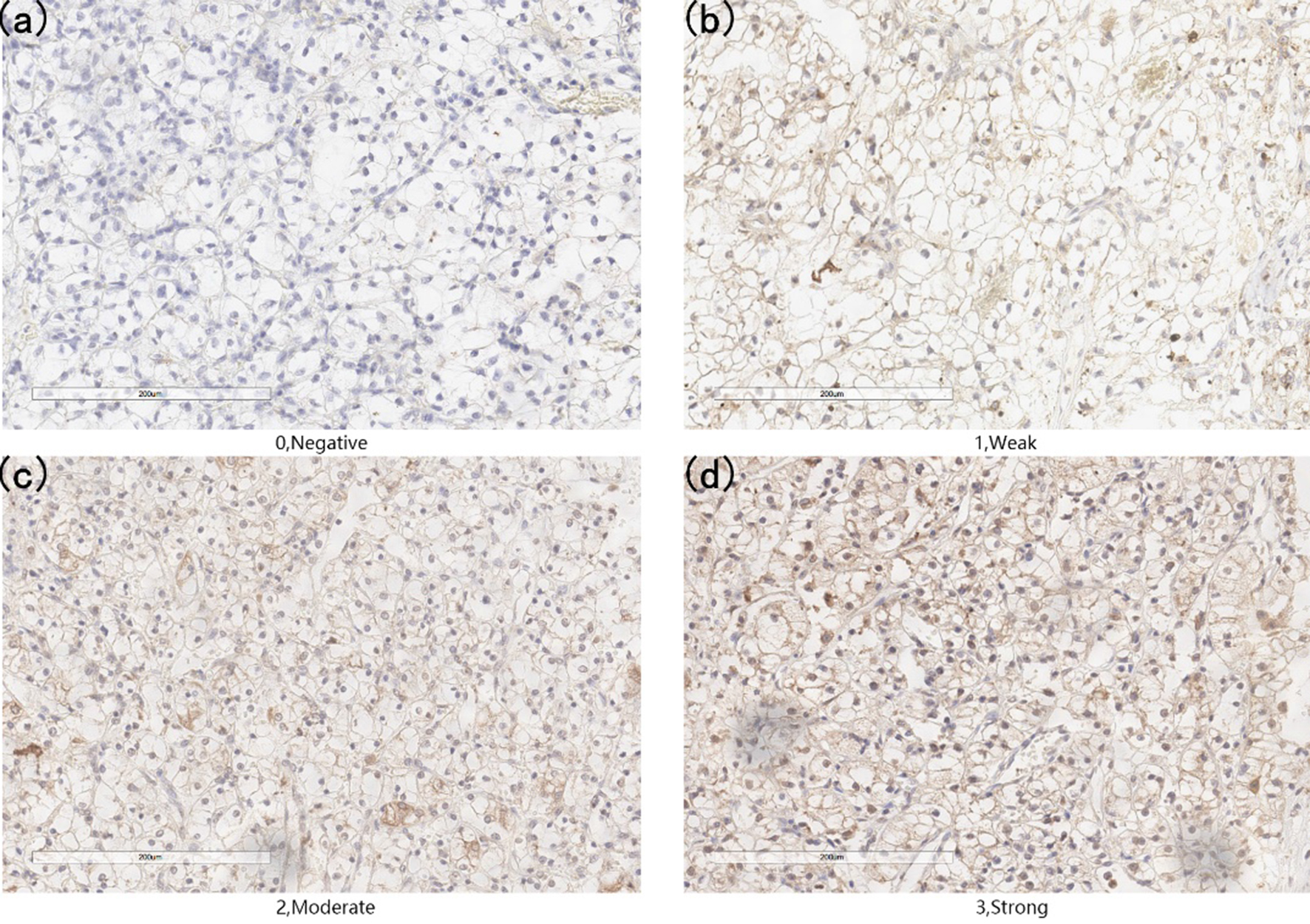
Representative adjacent normal renal tissues staining (A), ccRCC tissues staining (B) and the result of western bolt in RCC cell lines were shown (C).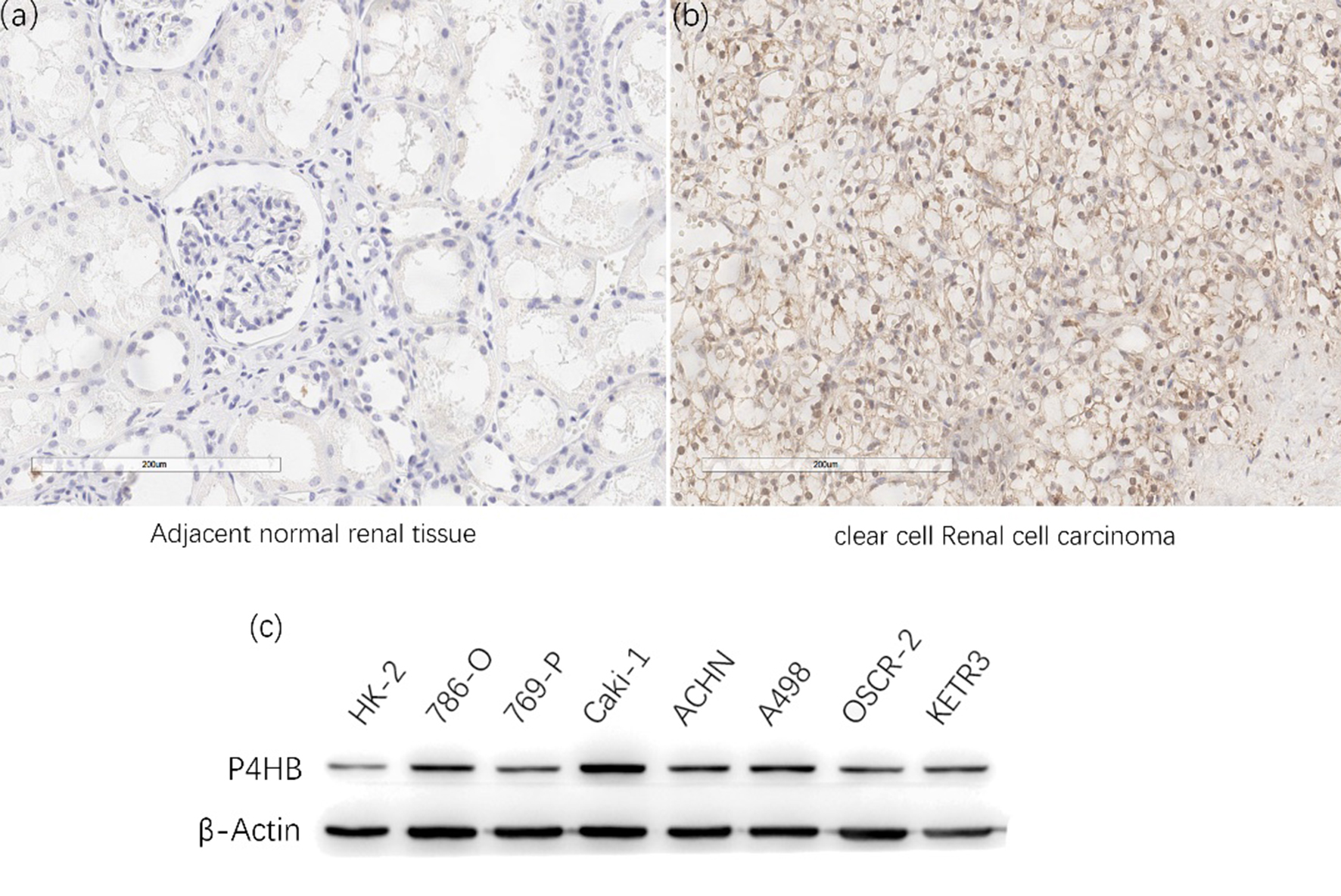
Univariate analysis and multivariate analysis of overall survival in our clinical database
The Kaplan–Meier survival curves for OS of P4HB in our database was shown (a). The protein expression of P4HB in ccRCC tissues and their adjacent normal renal tissues were compared using immunohistochemical staining. The average score of immunohistochemical staining (b) and the frequency distributions of proteins expression across the cohort were shown (c).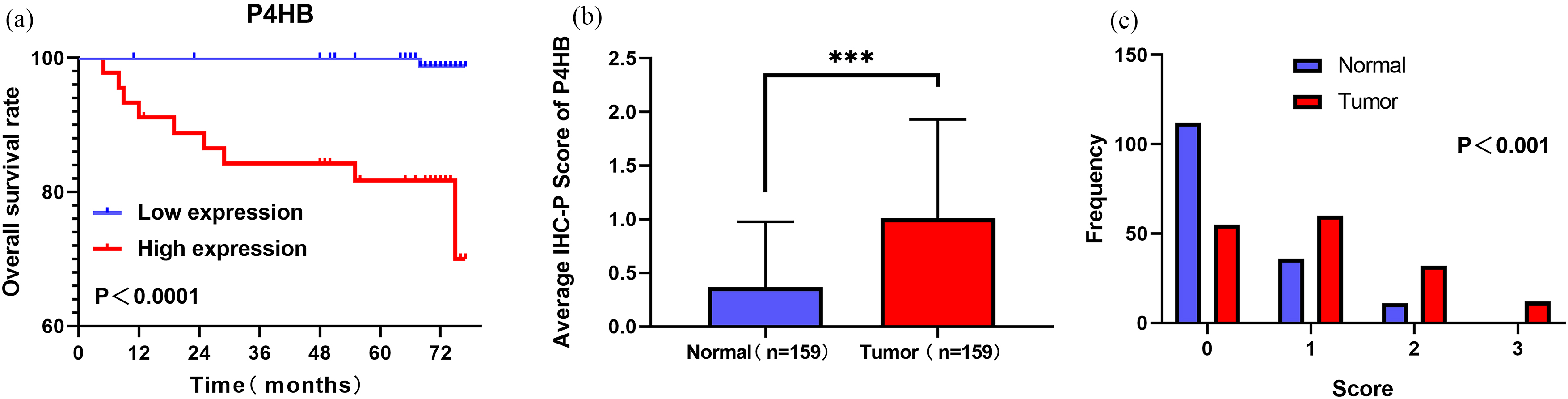
The mRNA level of P4HB is up-regulated in ccRCC
Based on 9 datasets from GEO database (GSE4282, GSE46699, GSE53757, GSE15641, GSE68417, GSE 14994, GSE40435, GSE71963, GSE76351), compared with normal tissues, P4HB was over-expressed in ccRCC tissues (Fig. 1). Meanwhile, compared with normal tissues, P4HB was over-expressed in ccRCC tissues in TCGA database (Fig. 2c). In addition, the relative RNA expression level of P4HB was also up-regulated in 20 paired clinical samples in our database (Fig. 2a and b).
The protein level of P4HB is up-regulated in ccRCC
Representative staining of these scores were presented (Fig. 3a–d). Meanwhile, representative staining of ccRCC tissue and adjacent normal renal tissue were showed (Fig. 4a and b). The IHC scores of 159 paired ccRCC tissues and their adjacent normal tissues showed that protein level of P4HB is up-regulated in ccRCC patients. The mean scores of P4HB proteins in ccRCC tissues and adjacent normal renal tissues were 1.06 and 0.36 (Fig. 5b), And the frequency distributions of these scores were presented (Fig. 5c). The outcomes of western bolt in renal cell lines showed that the P4HB protein level is up-regulated in cell lines of renal carcinoma, especially in ccRCC cell lines (Fig. 4c).
Kaplan-Meier survival curves for OS (a) and RFS (b) of P4HB in TCGA database were shown. The analysis of pathologic stage (c), pathologic T (d), pathologic M (e) and pathologic N (f) were shown.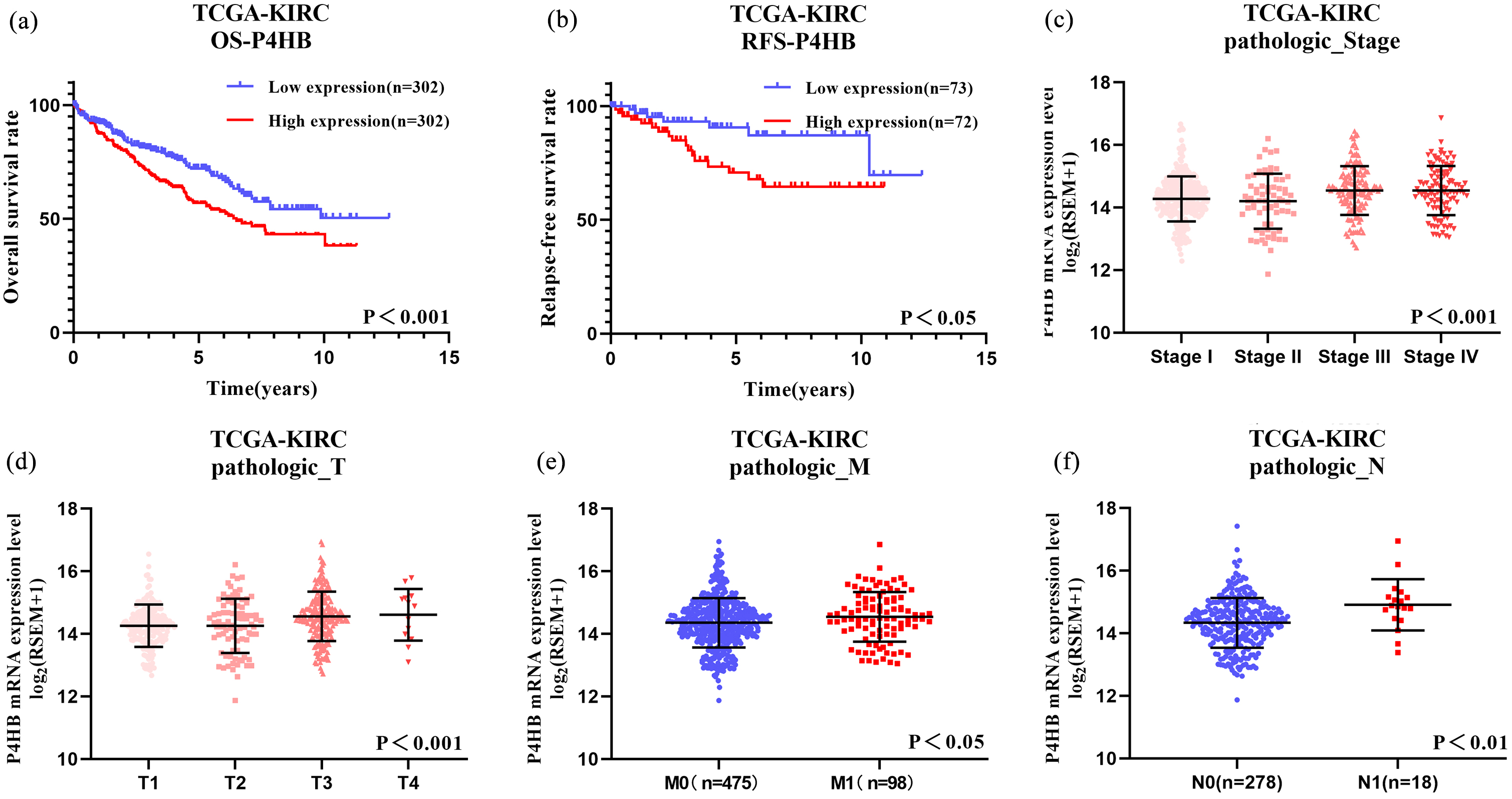
GSEA analysis was performed using 545 samples in TCGA database with the high expression and low expression of P4HB. The signature was defined by genes with significant expression changes.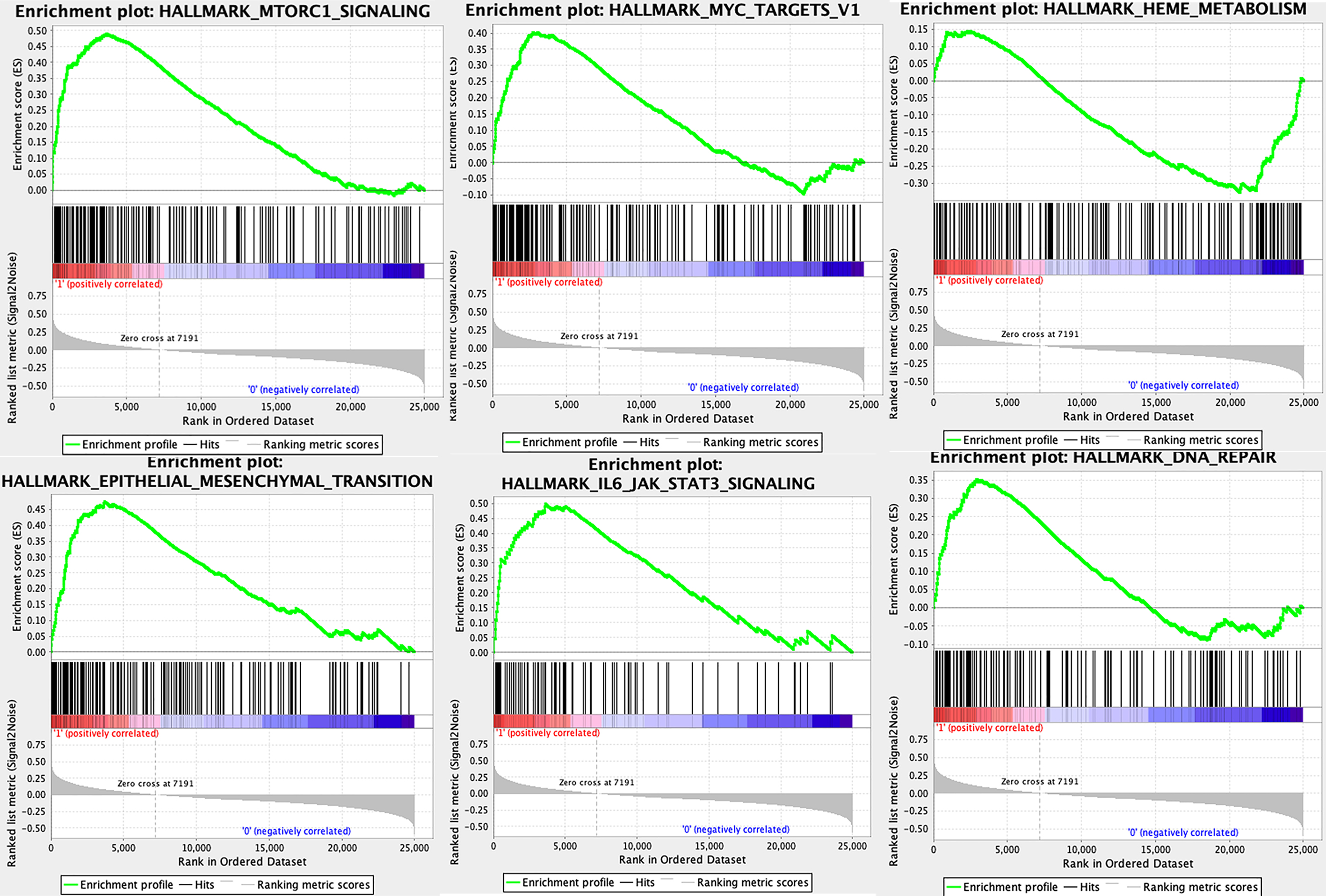
According to the median of P4HB, the patient’s OS in both TCGA database and our own database were analyzed via Kaplan-Meier analysis. High P4HB expression was identified to be correlated with worse OS and RFS in ccRCC patients (Fig. 6a and b). Besides, OS in high P4HB expression group was also poorer than the low P4HB expression group in our own database (Fig. 5a). Furthermore, the associations between P4HB and clinicopathological characteristics in ccRCC patients were explored. P4HB was significantly different in pathologic stage pathologic stage (
To evaluate whether P4HB could be the independent prognostic factors for ccRCC patients, the univariate and multivariate Cox regression analysis was conducted in both TCGA database and our own database, including age at initial pathologic diagnosis, gender, size (longest dimension), pathologic T, pathologic N, pathologic M and P4HB expression levels in TCGA database and initial pathologic diagnosis, gender, size (longest dimension) fuhrman scores and P4HB expression levels in our database. In the univariate Cox regression analysis, high P4HB expression, age, tumor size, Pathologic T and Pathologic M were correlated with worse OS in ccRCC patients in TCGA database (Table 1). Meanwhile, high P4HB expression, tumor size, Pathologic T and Fuhrman scores were correlated with worse OS in ccRCC patients in our database (Table 2). Multivariate COX regression analysis indicated that high P4HB, age, Pathologic T and Pathologic M were independent prognostic factors for OS in TCGA database (Table 1). At the same time, high P4HB, age and Fuhrman Scores were independent prognostic factors for OS in our database (Table 2).
P4HB may influence several vital processes in tumorigenesis
To explore the potential significance of P4HB in ccRCC, we used GSEA to compare the high expression and low expression of P4HB in the TCGA database. The result indicated that several vital regulatory genes involved in DNA repair, EMT, metabolism, JAK-STAT3 signaling, MTORC1 signaling and MYC target were enriched in cells with high P4HB expression (Fig. 7).
Discussion
P4HB, which encodes the beta subunit of prolyl 4-hydroxylase, is a highly abundant multifunctional enzyme that belongs to the protein disulfide isomerase family [14]. According to the previous studies, the activation of cellular stress responses mediated by the endoplasmic reticulum (ER) allowed cancer cells to survive outside their normal environment and P4HB was associated with enhancement of PDI activity [15]. Moreover, Gumireddy K found that P4HB promotion was related with tumor metastasis in vivo by using a mouse model in 2007 [16]. Other previous studies showed that P4HB was related to the invasive properties of malignant glioma [17], and P4HB attenuated temozolomide resistance in malignant glioma via ERSR pathways [18]. In addition, other studies have reported that high expression of P4HB is correlated with poor prognosis in both ovarian cancer and gastric cancer [12, 19]. GSEA also proved that P4HB played many important roles in tumorigenesis, several of which have been proved by previous studies. Theoretically, overexpression of P4HB in clear cell renal cell carcinoma may have several important roles in tumorigenesis.
As one of the most lethal malignant tumors, ccRCC has the worst prognosis among the common epithelial tumors of the kidney [20]. Although wide ranging clinical outcomes can be attributed to tumour heterogeneity in RCC, opportunities to further improve clinical outcomes on the basis of individual tumour characteristics (so called precision medicine) is an emerging field and genetic biomarkers are also beginning to be studied for associations with treatment outcome in various metastatic settings [21]. In this study, we examined the correlation between P4HB and ccRCC both in mRNA and protein level. There are more than 1000 samples from GEO database, TCGA database and our clinical database. According to our results, compared with adjacent normal renal tissue, the mRNA expression of P4HB in ccRCC tissue is higher in both GEO database and TCGA database. At the same time, a higher protein expression level of P4HB was observed in the 159 ccRCC tissues compared with paired adjacent normal renal tissue by using IHC method. In addition, the increased P4HB expression was significantly associated with poor prognosis of ccRCC patients, especially age and T stage related, which suggested that P4HB may be as a biomarker for diagnosis in ccRCC.
There are several limitations in the present study. Firstly, ccRCC cancer cell lines and in animal models need to be verified in vitro. Moreover, the mechanisms by which P4HB influences ccRCC remain unclear and required to be explored in the furture.
In conclusion, this study firstly demonstrated that P4HB was increased in ccRCC sample and cell lines. Moreover, overexpression of P4HB is an unfavorable prognostic factor. More studies are needed to verify the mechanism of P4HB as an oncogene and an unfavorable clinical biomarker for ccRCC.
Footnotes
Acknowledgments
This study was supported by the National Natural Science Foundation of China (No. 81670617).
Conflict of interest
The authors have read and approved this manuscript. All authors declare no conflicts of interest regarding the publication of this paper.
