Abstract
BACKGROUND:
Circulating tumor DNA (ctDNA) has been recognized as a promising biomarker for colorectal cancer (CRC) early diagnosis and postoperative monitoring. However, we hypothesize that the clinical value of ctDNA sequencing may differ for colon cancer (CC) and rectal cancer (RC).
METHODS:
Forty-three patients with primary CRC were prospectively enrolled. Tumor tissue samples, paired preoperative plasma samples and a series of postoperative plasma samples were obtained. Mutations in each sample were identified and compared.
RESULTS:
For 73.0% patients, at least one concordant mutation was detected in both tumor tissue DNA and paired preoperative ctDNA. The mutation concordance rate were higher in CC patients compared to RC patients (92.3% vs 45.5%;
CONCLUSION:
Liquid biopsy via ctDNA sequencing has great potential for the early detection and postoperative monitoring of CRC. The DNA of CC tissues is more likely to be released into blood than the DNA of RC tissues. This should be considered when diagnosing CRC patients with ctDNA sequencing technology.
Keywords
Introduction
The early diagnosis of colorectal cancer (CRC) offers great potential for reducing mortality. CRC is the third-most commonly diagnosed malignancy and the fourth leading cause of cancer-related deaths worldwide, accounting for about 1.4 million diagnosed cases and almost 700,000 deaths in 2012 [1]. The five-year survival rate for patients with stage I or II CRC reaches up to 90%, whereas it is only 12% for patients with stage IV CRC with multiple organ metastases. Unfortunately, approximately three fifths of patients are diagnosed at the late stage. CRC relapse occurs in 30–50% of patients, even in cases where treatment is effective [2, 3]. The survival rate can be rose, if the evaluation of each patient’s risk of recurrence can be improved. Therefore, there is a need for modified diagnostic tools in CRC. Colonoscopy has been widely used for the screening of CRC and contributed strongly to a drop in the mortality rate of CRC patients [4]. However, up to 25% of all colonoscopies fail due to extraordinary pain and inadequate bowel preparation [5]. Nowadays, carcinoembryonic antigen (CEA) is widely used as a potential biomarker for cancer diagnosis; however, it does not provide sufficient sensitivity and reliability since high CEA levels may be caused by infection and other diseases [6, 7]. Furthermore, CEA is not a sufficiently good indicator of the cancer severity since it may not increase in patients with advanced CRC [6]. Positron emission tomography (PET) and computed tomography (CT) are also effective diagnostic tools for cancers; however, they are associated with radiation exposure and also suffer from a high rate of false positivity [8]. Traditional recurrence risk evaluation methods that examine CEA and/or PET/CT also have not shown promising results [9, 10]. Almost 40% of patients with recurrence are not detected by CEA and/or PET/CT [11].
Fragments of circulating cell-free DNA (cfDNA) are released by different tissues in patients as well as healthy individuals. Noninvasive liquid biopsy based on cfDNA can be a great supplement of traditional diagnostic approaches [12, 13, 14]. Tumor-specific DNA mutations can be detected in circulating tumor DNA (ctDNA) [15], released from primary tumors or metastases, which is a significant component of the cfDNA [16]. Extensive studies have demonstrated that ctDNA could be used for the early diagnosis of a variety of cancers [17, 18, 19, 20], since it features higher sensitivity and specificity than traditional biomarkers [21, 22]. In addition, ctDNA can reflect the tumor burden in real time and offers both the potential for clinical implications in monitoring patients with cancer and prediction of recurrence risk [12, 21, 23, 24, 25]. The results of this study indicate the association between ctDNA and the occurrence and recurrence of colorectal cancer.
The obtained results also show that the ctDNA sequencing can provide different clinical outcome for colon cancer (CC) and rectal cancer (RC). Although CC and RC are collectively labeled CRC, their embryonic origin, anatomy, and function are different [26]. For example, the capillary networks of the colon are multi-layered and more abundant than those of the rectum [26]. Accordingly, we hypothesized that the ctDNA pattern of CC and RC patients may also differ. To further verify this hypothesis, the somatic mutations detected in the sequencing data of tumor tissues and paired plasma from patients with CC or RC were analyzed. The results show that the ctDNA can enable different clinical outcome for CC and RC.
Materials and methods
Ethics statement and patient cohorts
This study protocol was approved by the ethics committee of Shenzhen People’s Hospital on 4/6/2016 and the methods were carried out in accordance with the approved guidelines. All patients were counseled and consented according to the Declaration of Helsinki. Written informed consent was obtained from all patients for the use of blood and tumor tissue under the approval of the Ethics Committee. All samples and medical data used in this study have been irreversibly anonymized.
From September 2016 to May 2019, 43 patients with stage I, II, III, and IV CRC were prospectively enrolled at the Shenzhen People’s Hospital in Shenzhen, China. All patients were diagnosed with primary CRC and underwent radical resection (R0 resection), including 29 patients with CC and 14 patients with RC. For each patient, a fresh frozen tissues sample was collected during surgery and immediately stored at
DNA extraction and sequencing
Tumor DNA was extracted with the EasyPure
Clinic pathological characteristics and their correlation with preoperative ctDNA
Clinic pathological characteristics and their correlation with preoperative ctDNA
About 200 ng tumor DNA fragments and 10–50 ng cfDNA were input to construct libraries by the KAPA Library Preparation Kit (KAPA Biosystems, USA) according to the manufacturer’s protocol. Target enrichment was performed with the custom SeqCap EZ Library Kit (Roche NimbleGen, Madison, WI, USA). The targeted capture probes were designed based on genomic regions of 85 genes that covered most CRC related mutations (Supplemental Table 1). Capture hybridization was conducted according to the manufacturer’s protocol. After that, the libraries were amplified for 7–10 PCR cycles using 1
Sequencing data were preprocessed by Fastp for profiling quality status, trimming sequencing adapters and filtering low quality reads [28]. The filtered rdata were than mapped and aligned to the reference human genome, hg19 genome (GRch37) using Burrows Wheeler Aligner after removal of terminal adaptor sequences and low-quality reads. Pileup files for properly paired reads with mapping quality
Patient enrolment and study overview.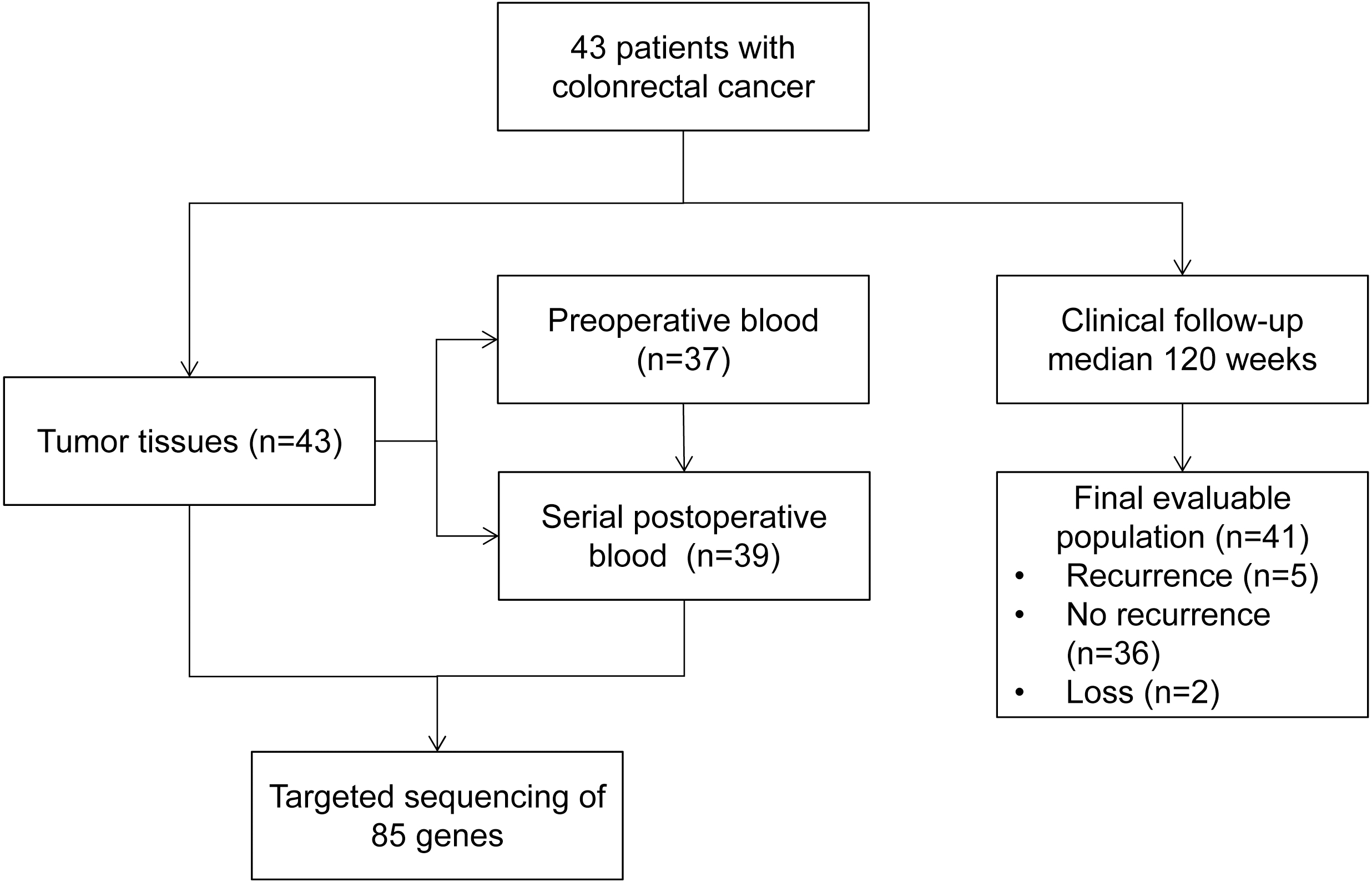
The yield of pre and post-operative cfDNA.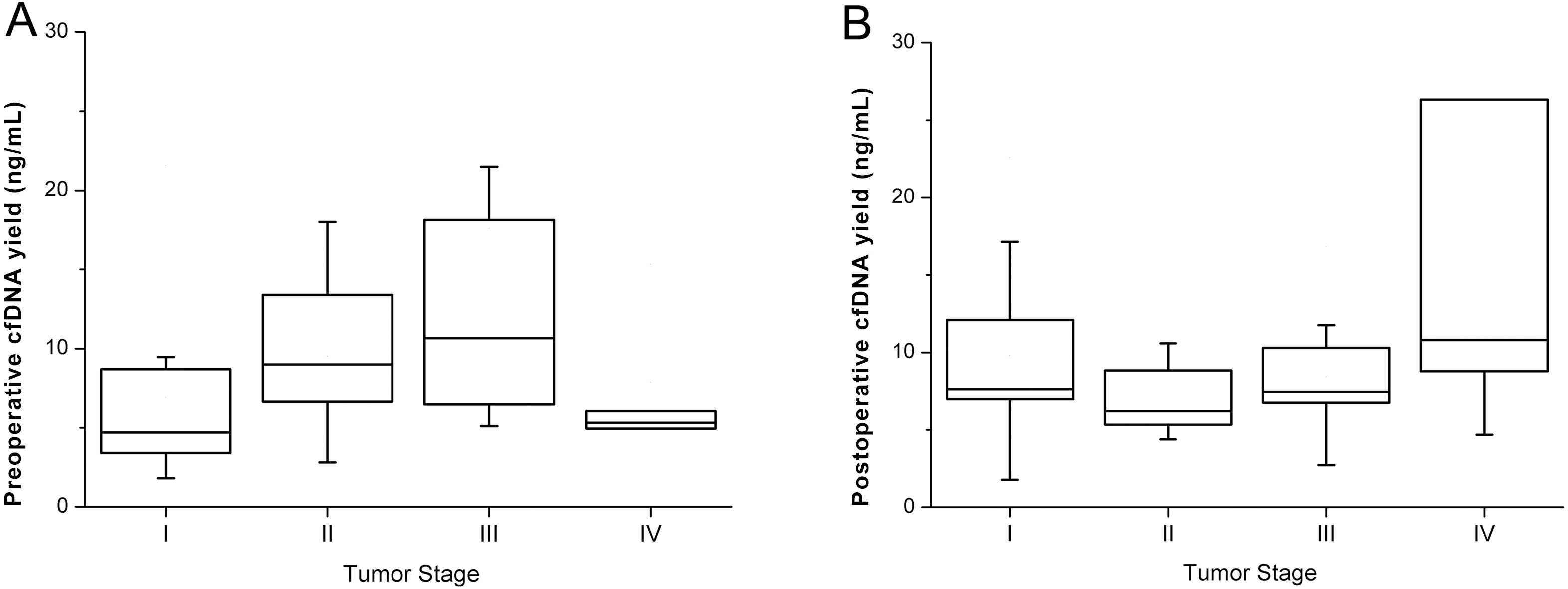
Differences in the sensitivity between ctDNA and CEA and between colon cancer and rectal cancer were assessed using the Fisher’s exact test. Correlation analyses between ctDNA status and clinical characteristics were evaluated using the Spearman’s coefficient of correlations. Hazard rations (HRs) were estimated using univariate Cox proportional hazard models. The two-tailed
Results
Patient characteristics
From September 2016 to May 2019, 43 CRC patients were prospectively enrolled at the Department of Gastrointestinal Surgery of the Shenzhen People’s Hospital in China. Patient characteristics and tumor features are summarized in Table 1 and Supplemental Table 2. The average age of patients was 58 and 53.5% patients were male. All eligible patients had histologically confirmed adenocarcinoma of the colon (67.4%) or rectum (32.6%) with stage I, II, III, and IV, and 39 (90.7%) of patients were moderately differentiated. All 43 patients underwent radical resection (R0 resection) and 29 (67.4%) patients received adjuvant chemotherapy at their clinician’s discretion. According to clinical and pathological features, 27 (62.8%) patients were identified with a high recurrence risk. By the end of 30 June 2019, the median follow-up time was 120 weeks (range, 7–146 weeks). At the end of follow-up, five patients (12.2%) experienced recurrence and of two (4.9%) died of disease progression. Patient enrolment and the study overview are presented in Fig. 1.
Mutation statistics of tumor samples and preoperative plasma from 43 patients with CRC. The colored boxes indicate where the mutations were detected in tumor tissues, and the triangles where the mutations were detected in preoperative plasma ctDNA.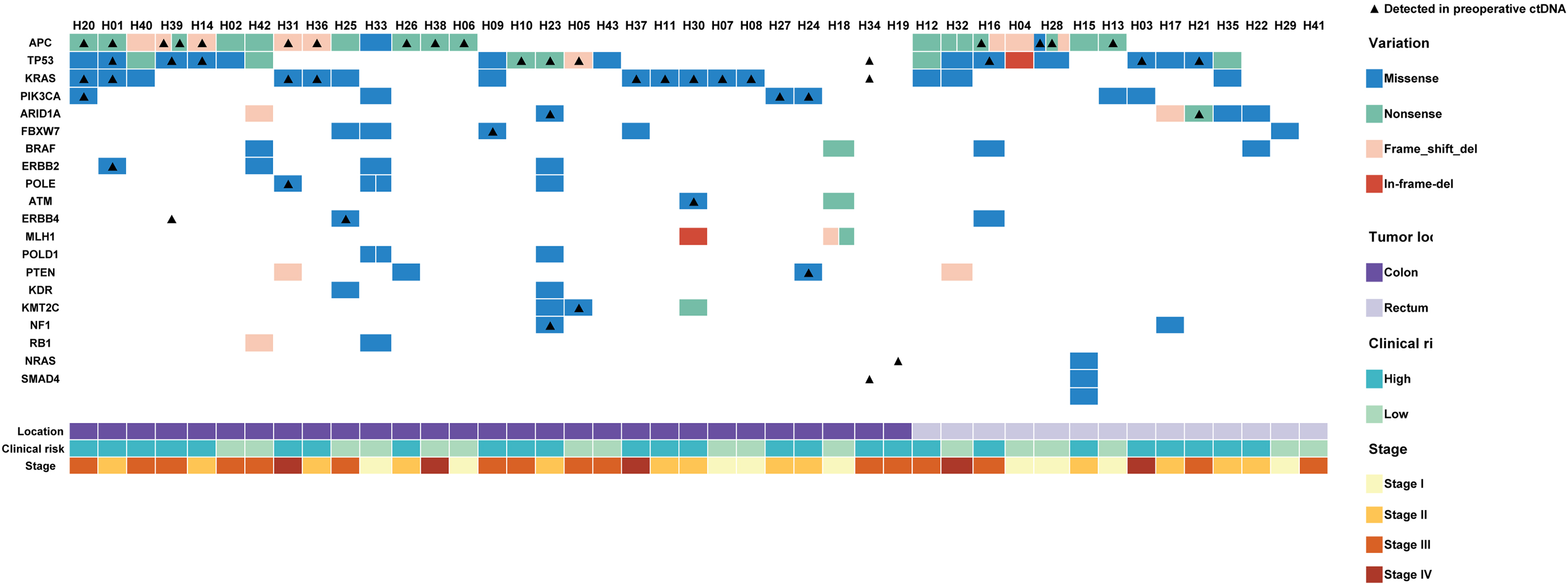
The median cfDNA yield in properative and postoperative plasma was 7.50 ng and 7.64 ng per mL plasma, range from 1.8 ng to 73.8 ng and from 1.8 ng to 117.2 ng. There were neither correlation (
Preoperative plasma status and CRC diagnosis
To identify tumor somatic single nucleotide variants (SNV) and short insertion/deletion (InDel) mutations, they were detected in tumor tissues via targeted capture sequencing. At least one somatic mutation was identified in tumor tissue of 40 of the 43 (93.0%) eligible patients. The median mutation number was 3 (range, 0–15). The most frequent mutations were gene TP53 (51.2%), APC (48.8%), KRAS (37.2%), and PIK3CA (13.95%) in the enrolled patients, which were “hotspot” mutations consistent with previous studies. No mutation of these four genes was detected in only six patients (14.0%) (Fig. 3).
Thirty-seven (86.0%) paired preoperative plasma samples were collected and sequenced using targeted capture panel. In 73.0% (27/37) of all patients, at least one mutation could be detected in preoperative plasma, which was consistent with the paired tumor tissue. No mutations were detected in tumor tissue of patient H19, H34 and H41. However, a missense mutation in exon 2 of NRAS (G37A) was found in the preoperative plasma sample of H19, and the VAFs was 0.3%. KRAS G12D, TP53 R273H and SMAD4 G419V were found in the preoperative plasma sample of H34. And there wasn’t found any mutation in the preoperative plasma sample of H41. The other seven patients did not show any mutation in the preoperative ctDNA. The most frequently identified alterations in preoperative ctDNA were also APC (32.4%), TP53 (27.0%), KRAS (27.0%), and PIK3CA (8.1%), which was consistent with the tumor tissues. Furthermore, none of these four genes were mutated in twelve 32.4%) patients (Fig. 3), among 8 patients were negtive.
The sensitivity of ctDNA for diagnosis CRC reached up to 78.4% (29/37). The sensitivity was 72.7% in 22 patients with stage I and II CRC, and 86.7% (13/15) in patients with stage III and IV CRC. Preoperative CEA levels of 35 patients were collected among all eligible patients. However, only in 11 patients, the CEA were detected positive (
Dynamic ctDNA monitoring of patients H03 and H21. A. Dynamic ctDNA monitoring of patient H03. B. Dynamic ctDNA monitoring for patient H21. The orange shaded box shows the period of adjuvant chemotherapy. The blue lines shows the time point of surgery and died or reccured. VAF, variant allele fractions; ctDNA, circulating tumor DNA.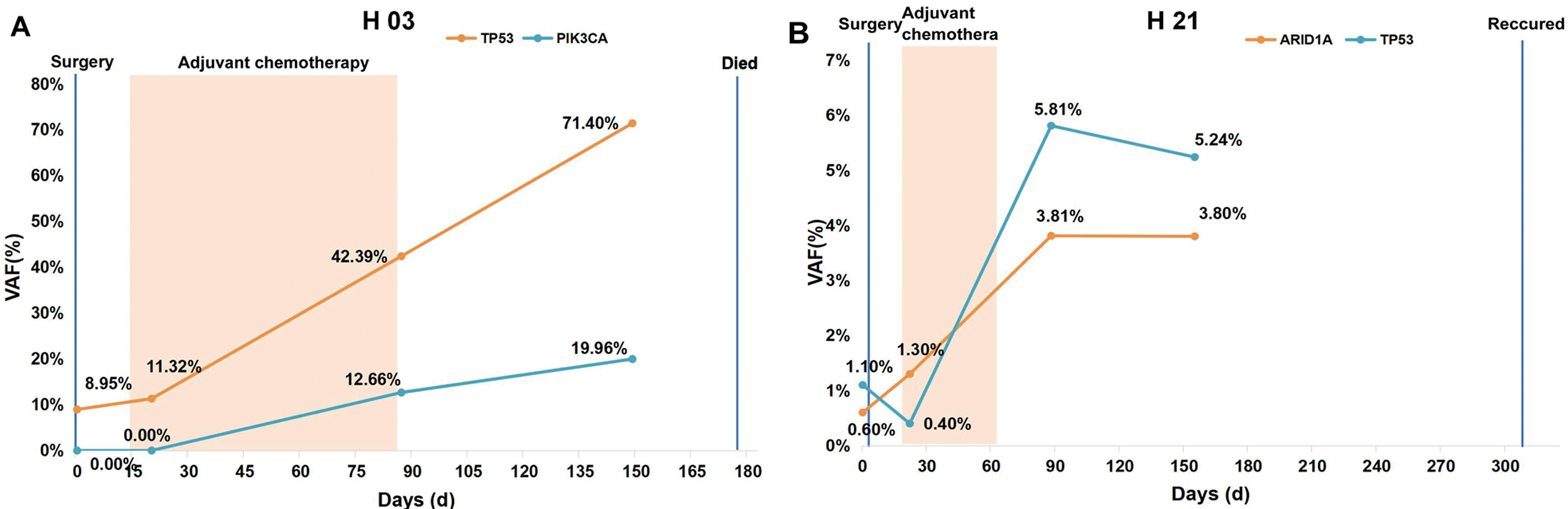
The yield of preoperative plasma cfDNA in CC was little higher than in RC (media yield of preoperative ctDNA per mL plasma was 8.25 ng vs 7.51 ng). Six of the eight preoperative ctDNA-negative patients had RC and only two had CC, which differed significantly (
Postoperative blood ctDNA for CRC recurrence prediction
A total of 98 postoperative plasma samples were collected from 39 enrolled patients. The number of mutations detected in the 1
In this study, ctDNA was also used to predict recurrence in CRC. The 1
Discussion
In this study, the clinical implications for CRC diagnosis was investigated with preoperative plasma ctDNA. The obtained data is consistent with previous studies [29]. The sensitivity of preoperative ctDNA (78.4%) is much higher than that of CEA (31.4%). In particular, in this study, ctDNA was also found to be effective for diagnosing early stage CRC. The sensitivity can reach up to 72.7%, while other biomarkers such as CEA and ECT2 are less sensitive [30, 31].
The recent technological improvement for detecting low-frequency mutations has improved ctDNA-based cancer diagnosis. The quantity of ctDNA likely depends on tumor load. For a tumor load of 30 g, only 0.15% of the cell-free circulating DNA would be released from the tumor [32]. Therefore, the sequencing depth was increased to about 10,000
As ctDNA is mostly released from the apoptosis and necrosis of tumor cells, it can provide more comprehensive information for the comparison of tumor tissues, and can be used to overcome problems introduced by tumor heterogeneity [33, 34]. For example, a missense mutation in the exon2 of NRAS (G37A) was detected in preoperative plasma ctDNA of patient H19, but it was not detected in paired tumor tissue.
The clinical value of preoperative ctDNA in CC and RC was found to differ. The concentration of preoperative plasma cfDNA in CC exceeded that of RC. Furthermore, the mutation-positive rate of preoperative ctDNA in CC was also higher than in RC (92.3% vs 45.5%,
Patients with increased VAFs of mutations detected in the 1
In this study, 85 genes mutated most frequently in CRC were analyzed via targeted capture sequencing. The sensitivity and specificity were 75% and 81.8%, respectively, which are superior to conventional clinical risk factors. Furthermore, the sensitivity and specificity can be further improved by using larger targeted capture panels.
Conclusion
This study suggests three potential values of ctDNA sequencing for diagnosing and monitoring of CRC. Firstly, mutations can be detected from preoperative ctDNA samples for even stage I/II CRC patients, suggesting that ctDNA can be a potentially promising biomarker for the diagnosis of CRC. Secondly, CC tumors may be more likely to release tumor DNA into blood than RC tumors, suggesting that different thresholds for CC and RC patients may be considered when evaluating sequencing results. Finally, the mutation statuses of postoperative ctDNA are significantly correlated with recurrence for CRC patients, suggesting that it may be beneficial to monitor CRC patients by sequencing their post-surgery ctDNA samples.
Footnotes
Acknowledgments
We thank the clinicians took part in the study design, sample collection and clinical information collection, meanwhile, the researchers and engineers from HaploX Biotechnology took part in the experiments and sequencing data analysis. The present study was funded by grants from Health and Family Planning Commission of Shenzhen Municipality (grant No. 201601016) for patient enrolment and sample collection, by Shenzhen Science and Technology Innovation Committee Technical Research Project (grant No. JSGG20180703164202084) for DNA sequencing, and by Shenzhen Strategic Emerging Industry Development Special Fund (grant No. 20170922151538732) for data analysis.
Conflict of interest
None.
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-190257.
