Abstract
Background
Analysis of K- and N-RAS mutations is mandatory before planning treatment of metastatic colorectal cancer, because only RAS wild-type (WT) patients can benefit from treatment with anti-EGFR monoclonal antibodies (cetuximab and panitumumab).
Case report
Here we report the case of a 69-year-old male patient affected by metastatic sigmoid cancer. He underwent left hemicolectomy, and histology diagnosed a well-differentiated, pT4, node-positive adenocarcinoma; KRAS analysis performed with direct sequencing identified a mutation in exon 2 of the KRAS gene (GGT->GTT). After first-line chemotherapy with FOLFOX6 plus bevacizumab, the patient underwent surgical resection of residual liver metastases. Histology showed metastatic deposits from colic adenocarcinoma with extensive coagulative necrosis. Mutational analysis of the KRAS gene was also performed on liver metastases by pyrosequencing assay, and no mutation was identified. Due to the discordant results (GGT->GTT exon 2 KRAS mutation in the primary tumor, and KRAS-WT in the liver metastases), mutational analysis on liver metastasis was repeated using next-generation sequencing and enriching the sample in tumor cells by manual microdissection; the same type of mutation of the primary tumor (GGT->GTT exon 2 KRAS gene) was confirmed.
Conclusions
Accurate tissue sampling and adequately sensitive assays are essential to correctly identify colorectal cancer patients who can be treated with an anti-EGFR monoclonal antibody.
Introduction
The introduction of new cytotoxic and targeted drugs during recent decades has significantly improved the overall survival of patients with metastatic colorectal cancer (mCRC). In particular the addition of monoclonal antibodies (mAbs) targeting the epidermal growth factor receptor (EGFR) (cetuximab and panitumumab) to first-line chemotherapy significantly prolongs survival in the subgroup of patients bearing K- and N-RAS wild type tumors (1). More recent clinical trials also suggest that – for RAS wild-type patients – using an anti-EGFR mAb as first-line therapy (2) is more efficacious than administering it in subsequent treatment lines. Therefore, nowadays testing for RAS mutations before starting first-line treatment of mCRC patients is mandatory.
Several assays for detection of RAS mutations are available, with direct sequencing (Sangertechnique) and real-time polymerase chain reaction (PCR) being the most commonly used in Italy (3). The American Society of Medical Oncology (ASCO) and the College of American Pathologists (CAP) (4), the European Society of Pathology (ESP) (5), and the Italian Society of Pathology and Cytopathology–ltalian division of the International Academy of Pathology (SIAPEC-IAP) (6) have published recommendations for tissue sampling and mutational analysis performance; however, no advice is given on which assay is to be preferred.
A second issue is the most appropriate tumor tissue (primary and/or metastasis) to analyze. RAS mutations are considered an early event in colorectal tumorigenesis, and it is assumed that concordance between the primary tumor and synchronous or metachronous metastases is close to 100%; therefore, sampling of multiple tumor sites (primary tumor and metastases) may not be necessary (7). However, as tumor cells continuously modify their genome, it is possible that a mutation in RAS genes occurs later. Thus it can be absent in primary tumor and present in metastases. The loss of RAS mutation in metastases is a very uncommon event, often considered a “false finding” due to methodological issues.
A further aspect to be considered is tumor heterogeneity. The tissue distribution of RAS mutant cells is mostly homogeneous in colorectal carcinomas (8), and a single dominant clone generally occupies most of the tumor volume. However, this picture can be different between the tumor center and the invasion front, and it can change after radiation or chemotherapy. Thus, tumor sampling for RAS analysis is a fundamental procedure to guarantee the reliability of the entire RAS testing process.
Finally, even though the threshold of RAS-mutated cells within a tumor mass that would define it as “resistant to anti-EGRF mAbs” is uncertain, some retrospective analyses have suggested that cetuximab treatment is ineffective also in tumors harboring a small number of mutated cells, and the most recent assays used in clinical trials have very high sensitivity (≥1%).
Here, we report the case of a mCRC patient bearing a G12V KRAS mutation in the primary tumor detected by direct sequencing analysis, and who showed a KRAS wild-type by pyrosequencing on liver metastases, resected after some cycles of chemotherapy plus bevacizumab. A further analysis by next-generation sequencing (NGS) of the liver metastasis, enriched in tumor cells by manual microdissection, confirmed the presence of a G12V mutation in the KRAS gene.
Case report
In March 2012, a 69-year-old man received a diagnosis of stage IV (isolated liver metastasis) sigmoid cancer, and underwent left hemicolectomy due to the symptomatic substenosing colon mass. The pathological diagnosis was well-differentiated (G1) adenocarcinoma, infiltrating perivisceral fat tissue with images of fistulas (pT4); 2 of the 24 isolated lymph nodes were metastatic (pN1); mutational analysis performed with direct sequencing revealed a mutation in exon 2 of the KRAS gene (GGT->GTT).
After surgery of the primary tumor, the radiological re-staging by contrast-enhanced computed tomography (CT) scan showed multiple liver metastases (at least 4 nodules, ranging from 8 to 53 mm, in S2, S5, S6 and S8).
In May 2012, the patient was referred to the Division of Medical Oncology of Federico II University of Naples (Italy), and he underwent first-line chemotherapy with FOLFOX-6 (leucovorin 200 mg/m2, 5-fluorouracil 400 mg/m2 in bolus, 5-fluorouracil 2,400 mg/m2 46 hours continuing infusion, oxaliplatin 85 mg/m2) plus bevacizumab (5 mg/kg day 1), every 14 days; disease reassessment was performed every 4 cycles. After cycle 8, a partial response was registered (only 2 nodules remained: at S6 and S8); locoregional treatment of the liver metastases was considered feasible, but while waiting for surgery, chemotherapy was continued for a further 4 cycles.
Liver surgery (R0 ultrasound-guided liver resection) was performed on December 2012 at a different institution. Pathological diagnosis reported a moderately differentiated adenocarcinoma originating from colon, which was caudal type homeobox transcription factor 2 (CDX-2) positive at immunohistochemistry. The samples showed extensive coagulative necrosis in about 70% of the specimen. The mutational analysis of KRAS gene, performed by pyrosequencing-based assay, revealed no mutation (the percentage of tumor cells in the analyzed tumor sample was about 50%).
Considering the 2 discordant results (GGT->GTT exon 2-KRAS mutation in the primary tumor, and KRAS-WT in the liver metastases), we decided to repeat the mutational analysis on both primary tumor and liver metastasis using NGS and after enrichment of samples in tumor cells by manual microdissection. The same type of mutation was found in both tissues: GGT->GTT in exon 2 of the KRAS gene (Fig. 1).
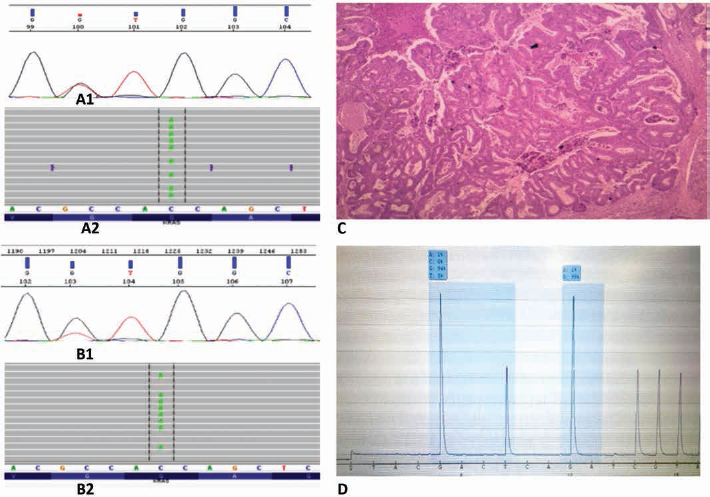
Sanger sequencing (SS) analysis detected a GGT->GTT mutation in exon 2 of KRAS gene (
The patient remained disease-free for about 5 months. In May 2013, he progressed (multiple metastases in several liver segments, ranging from 13 to 32 mm, and 1 nodule in his right lung), and he started second-line chemotherapy with FOLFIRI (leucovorin 200 mg/m2, 5-fluorouracil 400 mg/m2 bolus, 5-fluorouracil 2,400 mg/m2 46 hours continuing infusion, irinotecan 180 mg/m2) plus aflibercept (4 mg/kg day 1) every 14 days, achieving a partial response (only 2 lesions remained in the liver, complete disappearance of the lung nodule) after 8 cycles. Treatment was stopped due to deep vein thrombosis in his left inferior limb and pulmonary embolism, and surgery of the liver metastases was excluded for the same reason. After resolution of the thromboembolic events, the patient refused hepatic surgery, and underwent radiofrequency ablation of the 2 residual liver metastases. On September 2014, the patient experienced a disease progression in his liver (4 lesions, ranging from 14 to 28 mm in S1, S2, S5 and S6), plus 2 nodules in the left lung, and received further 6 months (12 cycles) of chemotherapy with FOLFIRI + bevacizumab, obtaining a stable disease as best response; maintenance with HDFUFA + bevacizumab was administered until December 2015. Treatment was stopped because the patient desired to undergo surgery for bilateral senile cataract. Progression in the liver and lung (dimensional increase of the known lesions) was recorded in June 2016, and regorafenib was administered for 3 cycles, with no benefit. The patient is still alive, receiving supportive care.
Discussion
RAS mutational analysis is fundamental in defining the strategy for treatment of mCRC patients, because only RAS wild-type patients can benefit from therapy with anti-EGFR mAbs (1).
The present case addresses the issue of the discordance of RAS mutational status between primary tumor and metastases, and raises the question of the optimal tumor tissue sampling and the most appropriate assay for RAS testing.
A review of 21 studies (7) testing the concordance of KRAS mutational status between primary tumors and metastases reported an overall concordance rate of 93%. Tissues were mostly formalin-fixed and paraffin-embedded; DNA was isolated by various techniques, the most frequently used being the QIAmp DNAkit (Qiagen, Germantown, MD, USA). In about half of these studies, mutational analysis was performed by PCR followed by sequencing; the other methods were PCR–restriction fragment length polymorphism (PCR-RFLP), allele-specific PCR (AS-PCR), allele-specific oligonucleotide hybridization (ASO), single-strand conformation polymorphism (SSCP) and pyrosequencing. Sub-analyses showed (a) the highest concordance (95%) between primary tumor and liver metastases, and the lowest (86%) between primary and lymph node deposits; (b) the discordance between primary and metastases was more frequent (14%) in KRAS-MUT as compared with KRAS-WT primary tumors (5%).
Discordance of RAS mutational status between primary tumor and metastases could possibly be explained by heterogeneity of the primary tumor, by technical issues or by late acquisition or loss of KRAS mutations during disease progression. In fact, even though discordance of mutational status is a possible event, it is very rare, and thus, current Italian and international guidelines do not recommend to rebiopsy a metastatic lesion for RAS testing if there is sufficient available material from the primary biopsy or resection (4-6).
To date, there is no method considered as the gold standard for RAS mutations testing, and many institutions use laboratory-based assays and not commercially diagnostic kits. The CAP guidelines state that laboratory-developed tests should be used only if the laboratory has carried out required validation testing and received accreditation (4), while the ESP recommends that laboratories develop standardized operating procedures and testing requirements (5). Currently, several methods have been approved for K- and N-RAS mutational analysis, which differ in time to be performed, technical complexity and sensitivity (9); in particular, direct sequencing of the PCR products has a relatively low sensitivity (10%-25%), while pyrosequencing is more sensitive (5%-10%). In this scenario, in our clinical case, the discordance between primary (exon 2-KRAS mutation, by Sanger sequencing) and liver metastases (wild-type by pyrosequencing) cannot be due merely to the sensitivity of the testing procedure.
The relationship between the percentage of tumor cells and the mutational rate is well established; when low-sensitivity methods (such as direct sequencing) are used, paraffin-embedded material containing more than 30% of neoplastic cells should be selected. The liver metastases, resected after 12 cycles of chemotherapy plus bevacizumab, showed extensive necrosis; thus, the absence of mutation might be due to poor tissue sampling, even though the number of neoplastic cells seemed to be sufficient. Tumor cell enrichment by manual microdissection or macrodissection of the formalin-fixed paraffin-embedded blocks before DNA extraction can be a very useful procedure to increase the sensitivity of mutation testing, limiting the presence of DNA coming from peritumoral surrounding cells, and capturing multiple neoplastic clones, contrasting heterogeneity and DNA damages, especially in case of tissue sampling after chemotherapy or radiation therapy.
NGS technologies offer rapid and sensitive (lower than 1%) screening of entire genomes or genome subregions, examining hundreds of potentially druggable alterations in cancer-related genes, thus it can be very useful in clinical practice when multiple genes have to be explored. In our institution, a strong correlation was found between point mutations (including RAS) identified by NGS (Ion Torrent Personal Genome Machine; Life Technologies, Carlsbad, CA, USA) and Sanger sequencing in colorectal cancer tissues (10). Therefore, following the discordant results derived from 2 different assays performed on 2 different tumor sites, NGS technique was applied, and the presence of the same mutation in exon 2 of the KRAS gene was confirmed in both tissues.
In summary, the best assay for RAS mutation testing has not been identified yet, and, probably, it will never be. Selection of the best technique to use can be based on the percentage of tumor cells in the sample, the DNA quantity and quality, the desired level of sensitivity and the laboratory expertise with a given technology.
The lesson learned by the reported case is that when discordant results in mutational status are found, inadequate methodology or inadequate selection of the specimen should be considered as possible explanations, and analysis with more stringent tissue sampling and with a more sensitive assay should be performed.
Footnotes
Financial support: No grants or funding have been received for this study.
Conflict of interest: None of the authors have any conflict of interest with the present manuscript.
