Abstract
Objective
Major depressive disorder (MDD) is a recurrent disorder with an increasing incidence. Alterations in key signaling pathways of the nervous system, such as the Wnt and MAPK pathways, mediated through microRNAs (miRNAs) provide crucial information regarding the etiopathology of MDD. We aimed to analyze whether the heterogeneity of literature findings regarding differential expression of miRNAs in the blood could arise from their different distributions among blood compartments.
Methods
We performed a pilot study analyzing the differential expression of miR-26a, miR-494, miR-30c, miR-93, and miR-101 and investigated their levels in white blood cells, total plasma (TP), exosomes from plasma, and exosome depleted plasma (EDP) in patients with MDD before and after antidepressant treatment with escitalopram and in healthy controls.
Results
MiR-494 was more abundant in EDP, and miR-26a and miR-30c were predominantly more abundant in TP relative to other blood compartments. Moreover, miR-30c, miR-101, and miR-26a, were significantly downregulated in TP of patients with MDD compared with controls. After antidepressant treatment, only miR-494 was significantly differently expressed in EDP.
Conclusions
This proof-of-principle study suggests that identifying the miRNA abundance in different blood compartments is crucial for biomarker development and could enrich the current knowledge regarding MDD pathophysiology.
Introduction
Major depressive disorder (MDD) is a debilitating neuropsychiatric disorder and exhibits the highest prevalence of all neuropsychiatric disorders in the general population (2%–20% worldwide). 1 It is predicted to be the second leading cause of global disease burden by 2030. 2 Because it is associated with declines in many psychosocial aspects, poor quality of life, functional impairment, morbidity, and a high degree of mortality by suicide, MDD is considered to be a major health concern. 3
Regarding the pathophysiology of MDD, neural remodeling and structural plasticity have been repeatedly linked to MDD, including diminished synaptic connectivity between the frontal lobe and other brain regions, neural atrophy, and numerous morphological modifications (decreased hippocampal volume, modified shape, and alteration of the number of dendrites).4,5
In addition, it is also believed that up to 50% of MDD cases are caused by genetic factors. 6 Of note, the serotonin transporter (5HTT/SLC6A4) and serotonin receptor 2A, brain-derived neurotrophic factor (BDNF), and tryptophan hydroxylase (rate-limiting enzyme for the synthesis of cerebral serotonin) are among the most well studied genes that are frequently associated with MDD. More recently, genome-wide association studies revealed numerous single nucleotide polymorphisms occurring in MDD, including polymorphisms of aczonin, a protein responsible for monoaminergic neurotransmission in the brain. 7
In this context, the analysis of disease-specific genes expressed in certain brain regions has contributed to acquiring novel insights into the molecular mechanisms involved in the pathology of MDD. Clearly, alterations at different levels of key cellular networks with implications in the disposition and cognition of an individual will lead to impaired synaptic function and defective structural plasticity and thus to the development of MDD. 2
Recently, studies of microRNAs (miRNAs) have received increased attention because of the potential role of miRNAs in the etiology and pathophysiology of mental disorders. MiRNAs are small, non-coding RNAs consisting of 18 to 24 nucleotides that play an essential role in gene regulation at the post-transcriptional level by causing messenger RNA (mRNA) degradation or inhibition of mRNA translation. 8 MiRNAs are involved in many physiological processes and pathological conditions, including MDD and suicidal behavior. 9 In MDD, miRNAs play key roles in neurogenesis, synaptic plasticity, and regulation of specific genes crucial for the signaling pathways involved in this condition. 10
Furthermore, miRNAs are found in most body fluids, such as whole blood, plasma, serum, exosomes, cerebrospinal fluid, saliva, urine, and tears, and are generally very stable and easy to collect. Although they have a broad distribution among body fluids, given their heterogeneous origin, it is imperative to discover miRNAs of cerebral origin. In neuropsychiatric disorders, miRNAs are released actively or passively from neurons into the circulating blood 11 and are enclosed in exosomes or other microvesicles. 12 Therefore, their presence in peripheral tissues is of critical interest for their potential use as biomarkers of MDD. Studies show that in psychiatric illnesses, miRNAs can be easily detected in whole blood, plasma, serum, peripheral nucleated blood cells (PNBCs), and exosomes.13,14 There is a high degree of heterogeneity in the literature regarding the analytical assessment of circulating miRNAs as biomarkers of MDD, but these miRNAs were investigated in different blood compartments.
Herein, we have investigated the distribution among intracellular and extracellular blood compartments of several miRNAs that were reported in our previously published screening study, as well as that from other studies, as being differentially expressed in patients with MDD compared with normal controls or according to MDD treatment.15,16 According to previous studies, we selected miR-26a, miR-494, miR-30c, miR-93, and miR-101 and assessed their relative abundance in white blood cells, plasma, exosomes from plasma, and exosome depleted plasma (EDP) in patients with MDD before and after antidepressant treatment and in healthy controls.
Materials and methods
Study population and sample collection
The exclusion criteria for healthy controls were as follows: the presence of any psychiatric diagnosis on axis I of the Diagnostic and Statistical Manual of Mental Disorders, 4th Edition, Text Revision (DSM-IV-TR), either at the time of assessment or in the 12 months prior to the initiation of the research; the presence or intention of pregnancy during the study duration (12 weeks starting from the baseline); and the presence of any medical condition requiring chronic drug treatment. Diagnostic and sampling methods were performed as previously described. 16 Briefly, all patients included in the study fulfilled the DSM-IV-TR diagnostic criteria for MDD.
All subjects provided written informed consent, and the study was approved by the Ethics Committee for Scientific Research of the Timisoara County Emergency Hospital (Approval No. 61/29 October 2013e).
Blood samples were collected in ethylenediaminetetraacetic acid-containing tubes from patients with MDD before and 12 weeks after antidepressant treatment with escitalopram. 16 The patients received one daily dose of 10 mg of escitalopram. Treatment response was measured after 12 weeks of antidepressant treatment using the same tools used for diagnosis (Structured Clinical Interview for Psychiatric Disorders of axis I of the DSM-IV-TR and the Hamilton Depression Rating Scale, 17-item version). 16 Blood samples were collected only once from healthy controls. Plasma and peripheral nucleated blood cells were separated, aliquoted, and immediately stored at −80°C until further use.
miRNA analysis
Exosomes were separated from plasma using the Exosome Isolation Kit from plasma (Life Technologies, Waltham, MA, USA) according to the manufacturer’s instructions. Total RNA (containing miRNAs) was extracted from total plasma (TP), exosomes from plasma, EDP and PNBCs, using the Qiagen miRNeasy kit (Qiagen, Hilden, Germany) following the manufacturer’s protocol. In all samples, except PNBCs, 25 fmol/mL of Caenorhabditis elegans miR-39 (cel-miR-39) was added as a normalization control, as per the manufacturer’s instructions. RNU-24 was used as an endogenous control for PNBCs.
The relative abundances of individual miRNAs were measured in triplicate with TaqMan MicroRNA kits (Life Technologies), according to the manufacturer’s instructions, using a 7900HT real time PCR instrument (Thermo Fisher Scientific, Rockford, IL, USA). The following TaqMan gene expression assays were used: miR-26a assay ID: 000405, miR-494 assay ID: 002365, miR-30c assay ID: 000419, miR-93 assay ID: 001090, and miR-101 assay ID: 002253. The relative expression of miRNAs was calculated using the delta-delta Ct method.17,18
Statistical analysis
GraphPad Prism (GraphPad, San Diego, CA, USA) was used for the statistical analysis and graphical presentation of data. A D’Agostino-Pearson omnibus normality test was performed to assess the Gaussian distribution of the data. A Mann–Whitney U-test was used to compare miRNA expression between the case and control groups, and the Wilcoxon signed-rank test was used for comparisons of different blood compartments before and after treatment in patients.
Results
We investigated 11 patients with MDD and 11 healthy controls (Table 1). The controls were partners of eligible and enrolled study subjects, with an age difference less than 5 years, and aged between 18 and 65 years. The relative expression of the investigated miRNAs in patients compared with controls in the various blood compartments is presented in Table 2.
Clinical and demographic characteristics of included subjects.

MDD, major depressive disorder; HDRS-17, 17-item Hamilton Depression Rating Scale.
Relative expression of the investigated miRNAs among blood compartments in patients compared with controls.

FC, fold change; TP, total plasma; EXO, exosomes from plasma; EDP, exosome depleted plasma; PNBCs, peripheral nucleated blood cells.
In this pilot project, we found that miR-30c, miR-101, and miR-26a were significantly downregulated in TP of patients with MDD compared with controls (p < 0.05, Figure 1). Among extracellular compartments, the same changes in miRNA levels were also found to be statistically significant in EDP (Figure 2) and PNBCs (Figure 3), but not in exosomes. There were no statistically significant differences in the miR-494 levels between the case and control groups in any of the blood compartments.
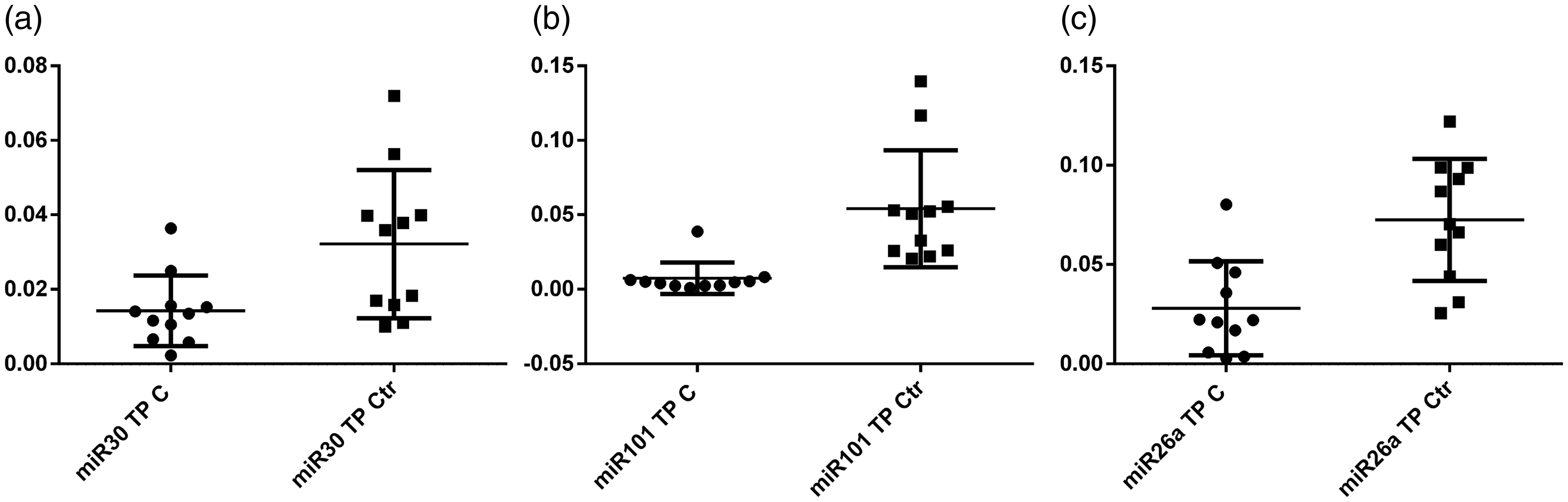
Relative quantities of miR-30c (a), miR-101 (b), and miR-26a (c) in total plasma (TP) in the case (C) and control (Ctr) groups (p < 0.05).

Relative quantities of miR-30c (a), miR-101 (b), and miR-26a (c) in exosome depleted plasma in the case (C) and control (Ctr) groups (p < 0.05).

Relative quantities of miR-30c (a), miR-101 (b), and miR-26a (c) in peripheral nucleated blood cells in the case (C) and control (Ctr) groups (p < 0.05).
The relative expression of the investigated miRNAs among blood compartments in patients before and after treatment is presented in Table 3.
Relative expression of the investigated miRNAs among blood compartments in patients before and after treatment.

FC, fold change; TP, total plasma; EXO, exosomes from plasma; EDP, exosome depleted plasma.
In this pilot project, we found that only miR-494 was significantly differentially expressed in EDP after treatment (p = 0.008). Before treatment, EDP exhibited significantly higher miR-494 levels than exosomes (p < 0.001), and no difference was found compared with TP. However, TP had significantly more miR-494 than exosomes (p < 0.001). This distribution pattern of miR-494 among blood compartments was similar after treatment. A similar distribution pattern among blood compartments was found for miR-101, regardless of treatment status.
MiR-26a was significantly more abundant in TP than in both EDP and exosomes (p < 0.001), and no difference was observed between EDP and exosomes, suggesting that extracellular miR-26a is found both inside and outside of exosomes. These differences remained similar after treatment. A similar pattern of distribution among blood compartments was found for miR-30c, regardless of treatment status.
None of the expression levels of the miRNAs that were investigated in this study changed according to treatment in PNBCs. We did not compare PNBCs with extracellular compartments because different endogenous controls were used.
Discussion
The aim of this small-scale, pilot study was to provide a comparative analysis regarding the expression levels of some mRNAs to produce a global snapshot of the heterogeneity and differences among various blood compartments in the same individual at different time points (before and after treatment) and between patients suffering from MDD and healthy controls. Nevertheless, our findings provided insights into the predominance of certain miRNAs in the blood compartments investigated.
Thus, we confirmed the differential expression of four of the five miRNAs investigated in extracellular blood compartments of patients with MDD compared with controls. This difference was significant only for the TP and EDP, suggesting that these miRNAs are preponderantly transported outside of exosomes. MiR-494 was differentially expressed in the blood of patients with MDD after treatment with escitalopram; however, this difference was significant only for EDP. Comparison among extracellular blood components suggested that miR-494 is carried in blood preponderantly outside exosomes. MiR-26a was present at similar levels in both exosomes and EDP.
In a similar manner, Bocchio-Chiavetto et al. 15 conducted a genome-wide quantitative analysis of miRNA in the blood of 10 patients with MDD before and after 12 weeks of treatment with escitalopram. They found that 30 miRNAs were differentially expressed after escitalopram treatment: 28 miRNAs were upregulated, and two miRNAs were downregulated. Among 13 miRNAs that may regulate gene expression in the brain, miR-26a and miR-494 were shown to play a role in neural plasticity, stress responses, and the pathogenetic mechanisms of several neuropsychiatric diseases. Both miRNAs have been reported to play a role in the pathogenesis of psychiatric disorders and in the mechanism of action of antipsychotic drugs and mood stabilizers. Additionally, miR-26a widely contributes to the action of the neurotrophin BDNF in the brain, and miR-494 is downregulated in the prefrontal cortex of patients with MDD.
Belzeaux et al. 14 examined the expression profiles of different miRNAs in peripheral blood mononuclear cells (PBMCs) from 16 patients with MDD and 13 controls at baseline and at 2 and 8 weeks after treatment. Fourteen miRNAs showed dysregulation (nine miRNAs were upregulated and five were downregulated). Among these 14 dysregulated miRNAs, miR-494 was downregulated at baseline in patients with MDD compared with controls.
Using the same sample specimen and blood compartment as in the previously mentioned study (PBMCs), Fan et al. 19 analyzed miRNA expression levels in 81 patients with MDD and 46 healthy volunteers and found five miRNAs that were significantly upregulated (miRNA-26b, miRNA-1972, miRNA-4485, miRNA-4498, and miRNA-4743). Moreover, gene prediction analysis revealed that differentially-expressed miRNAs may have implications in the development of MDD because a significant association was found between the aforementioned miRNAs and crucial pathways for diverse brain functions. Moreover, one study found that 29 interconnected miRNAs exhibited blunted expression in the prefrontal cortex of 18 antidepressant-free depressed suicidal subjects and had a strong impact on neuronal plasticity and axonal growth. 20 Furthermore, miR-1202 was downregulated in the ventrolateral prefrontal cortex, which correlates with an increase in metabotropic glutamate receptor 4 (GRM4), which is responsible for neurotransmission. Ex vivo experiments showed marked upregulation of miR-1202 expression upon chronic exposure to either imipramine or citalopram. In terms of signaling pathways, Wnt and MAPK signaling are among the most studied for their role in MDD through mediation of cell proliferation and hippocampal plasticity.21,22
Interestingly, we observed significant alterations in miRNA expression in different blood compartments. However, because they could not be directly linked to treatment with the antidepressant escitalopram, we assumed that these differences in expression levels may occur because of the pathophysiological mechanism of MDD alone or may only reflect an epiphenomenon of the actual MDD mechanisms, with changes in miRNAs being secondary to the actual pathophysiological mechanisms.
In addition, as mounting evidence increasingly suggests that a multitude of miRNA species have major implications and roles in the etiopathogenesis of MDD, the clinical use of miRNAs as diagnostic biomarkers and therapeutic targets holds great promise for future use. 23 It is therefore crucial to identify the source of miRNAs in blood compartments, especially because various factors could influence changes in their expression. One such factor is stress, which is a major risk factor for depression. Furthermore, various behavioral and nutritional changes that occur in subjects with MDD may also cause alterations in peripheral sources but not play a role in the etiology and pathological mechanisms of MDD. Another aspect to consider is that miRNAs act mostly as a fine tuning mechanism of gene expression, and the effect is not usually to silence gene expression but is rather more nuanced by decreasing the levels of proteins; therefore, even small changes in miRNA expression could have a biological effect, as previously described.24,25
In our pilot study, we attempted to profile the expression levels of different miRNAs in four blood compartments. However, our study had some limitations that must be acknowledged. These limitations arise primarily from the limited sample size, and therefore studies with larger sample sizes are needed. In this respect, future large-scale studies could eventually draw definitive conclusions regarding the distribution of miR-494 and miR-26a among blood compartments in patients with MDD in relation to antidepressant treatment.
Conclusion
MDD is a complex and heterogenous disease with numerous molecular mechanisms that remain to be elucidated. MiRNAs have been shown to be involved in gene regulation of important targets of the nervous system and brain function, with potential correlations in the development of MDD. The vast origin of miRNAs in the blood circulation makes it imperative to identify their source. Thus, we found that some miRNAs are more abundant in certain blood compartments than others (miR-494 was more abundant in EDP and miR-26a and miR-30c were more abundant in TP), and the investigated miRNAs had a general tendency of being significantly (p < 0.05) downregulated in the TP of patients relative to healthy subjects. Regarding the differential expression of the selected miRNAs before and after antidepressant treatment, only miR-494 was significantly differentially expressed after treatment in EDP (p = 0.008). MiR-26a and miR-30c were expressed at similar levels before and after antidepressant treatment in different blood compartments. None of the expression levels of the miRNAs that were investigated in this study changed in PNBCs according to treatment. Future large-scale studies are needed to confirm these findings to assess the biomarker potential of miRNAs in MDD.
