Abstract
OBJECTIVES:
Pancreatic cancer is one of the most lethal malignancies worldwide. Pancreatic adenosquamous carcinoma (PASC) is a rare histological type of pancreatic carcinoma with a poor prognosis. The median survival time after diagnosis is less than one year. It is believed that the pathogenesis of PASC is different from pancreatic adenocarcinoma. In this study, we tried to reveal the intrinsic gene mutations associated with PASC through whole exome sequencing.
METHODS:
Both cancerous and paracancerous tissues were collected from 12 pathologically diagnosed PASC patients. Their clinical characteristics were collected, and patient survival information was obtained through follow-up. The correlations between the mutations and clinical characteristics were analysed.
RESULTS:
Germline mutations were identified in MAP3K1 (9 cases), PDE4DIP (7), BCR (7), ALK (6), USP6 (5), AR (4), HLA-A (4), SPEN (4), KMT2D (3), NUTM2B (3), ZFHX3 (3), and MN1 (3), while somatic mutations were found in TP53 (5), KRAS (3), HRNR (3), and OBSCN (3). Peripheral tissue invasion was associated with somatic mutations in KRAS (
CONCLUSION:
In conclusion, susceptibility genes including MAP3K1, PDE4DIP, and BCR are frequently found to be mutated in the germlines of PASC patients. Somatic mutations in KRAS, OBSCN, and HRNR and germline mutations in USP6 are related to tumour invasion and metastasis, reinforcing the necessity of translating these potential biomarkers into clinical practice.
Abbreviations
Introduction
Pancreatic cancer is one of the most lethal malignancies worldwide, with 90,100 estimated new cases and 79,400 deaths in China in 2015 [1]. The majority of patients have regional lymph node (29%) or distant metastases (52%) at initial diagnosis, and their prognosis is dismal with a 5-year survival rate of 11.5% or 2.7%, respectively [2]. Pancreatic adenosquamous carcinoma (PASC) constitutes 1–4% of pancreatic carcinomas, and is a rare, clinically aggressive histological form with a poor prognosis [3, 4, 5]. The median survival time after diagnosis is less than one year [3].
PASC contains both ductal adenocarcinoma and squamous carcinoma within the same tumour. It is widely accepted that PASC is derived from pancreatic tissue, rather than a combination of two sources of malignant tumours. Nevertheless, very little is known about the pathogenesis of PASC due to its rarity, especially the genomic features. Recently, Fang et al. performed whole genome sequencing of 17 PASC tumours [6]. Combined with the 11 PASC cases from the International Cancer Genome Consortium, the most frequently mutated genes included KRAS, TP53, and SMAD4. We have performed whole exome sequencing analysis of 12 PASC patients in order to identify additional cancer-related gene mutations, in hopes of further understanding PASC pathogenesis.
Materials and methods
Patients
Cancerous and paracancerous tissues were collected from 12 patients with pathologically diagnosed PASC at the Department of General Surgery of Changhai Hospital, Shanghai between February 2015 and January 2016. Demographic and clinical baseline characteristics were collected from medical records, and survival data were obtained through follow-up. The present study was approved by the Institution Review Board of the Changhai Hospital, Shanghai. All patients or their families signed informed consent forms before the initiation of the study. All methods were performed in accordance with relevant guidelines and regulations.
Sample acquisition and preparation
DNA was purified from the macro-dissected frozen cancer tissues and paracancerous tissues. Sections from the frozen tissues (
Whole exome sequencing
Indexed samples were pooled for loading onto flow cells for sequencing on an Illumina sequencing platform (HiSeq
Demographic and clinical characteristics of the 12 patients with pancreatic adenosquamous carcinoma
Demographic and clinical characteristics of the 12 patients with pancreatic adenosquamous carcinoma
CBD: Common bile duct. Data were collected from clinical medical records. Immunohistochemistry was used to assess the p53 expression using mouse anti-human p53 monoclonal antibody (Maixin-Bio Co., Fuzhou, China). p53 was interpreted as positive if
The Illumina next-generation sequencing (NGS) platform was used for whole exome sequencing, and the results were saved in the FASTQ file format. All the raw sequencing reads in FASTQ format were quality controlled using FastQC v0.11.3 [8]. The trimmomatic V0.32 (trimmomatic-0.36.jar PE-threads N R1 R2 R1.paired.fq.gz R1.unpaired.fq.gz R2.paired.fq.gz R2.unpaired.fq.gz ILLUMINACLIP: adapter. fa:2:30: 10 LEADING:3 TRAILING:3 SLIDINGWINDOW: 4:15 MINLEN:36) was used to trim adapters and also bases with Phred quality scores under 30 [9]. Trimmed reads longer than 36 bp were mapped to human geno-me sequence hg19 (NCBI Build 37,
Both germline and somatic mutations associated with PASC were identified. Normal tissue was used to call the germline Single Nucleotide Variants (SNVs) and short insertions and deletions (INDELs) using GATK HaplotypeCaller. Raw germline variants were filtered using the standard hard filtering parameters according to GATK Best Practice recommendations [12, 13]. Comparing normal and cancer tissues, somatic mutations (SNPs and INDELs) were identified using GATK Mutect2. Both germline and somatic variants which passed the filtering criteria were annotated with ANNOVAR [14]. Variants were considered as potential candidates when three of five prediction tools called them damaging (SIFT, Polyphen2_HDIV, Polyphen2_HVAR, LRT, and MutationTaster) [15, 16, 17, 18].
Statistical analysis
All statistical analyses were carried out using SPSS (SPSS Inc, Woking, Surrey, UK) version 22. Correlations between germline or somatic mutations and clinical characteristics or outcomes were analysed using the Chi-square test or
Results
Subjects characteristics
We conducted whole exome sequencing of 12 PASC patients including three females and nine males. Their demographic and clinical characteristics are shown in Table 1. The primary tumour size ranged from 3 to 12 cm. Eight out of 12 patients had lymphatic metastasis. Common bile duct invasion occurred in three cases, duodenum invasion in two cases, stomach invasion in two, left adrenal gland invasion in one, spleen invasion in one, colon invasion in one, while four patients did not experience peripheral tissue invasion. At the end of our data collection, seven patients had died from local recurrence, liver metastasis and/or pulmonary metastasis, while the remaining five cases had achieved good clinical outcomes.
Identified germline and somatic mutations
A total of 122 germline mutated genes were identified, harbouring mostly missense mutations (Supplemental Table). There are 21 genes which are often mutated in pancreatic cancer, of which four were also mutated in the PASC samples in this study, including TP53 (
Germline mutation spectrum in 12 PASC patients with 36 genes that were identified in at least two samples.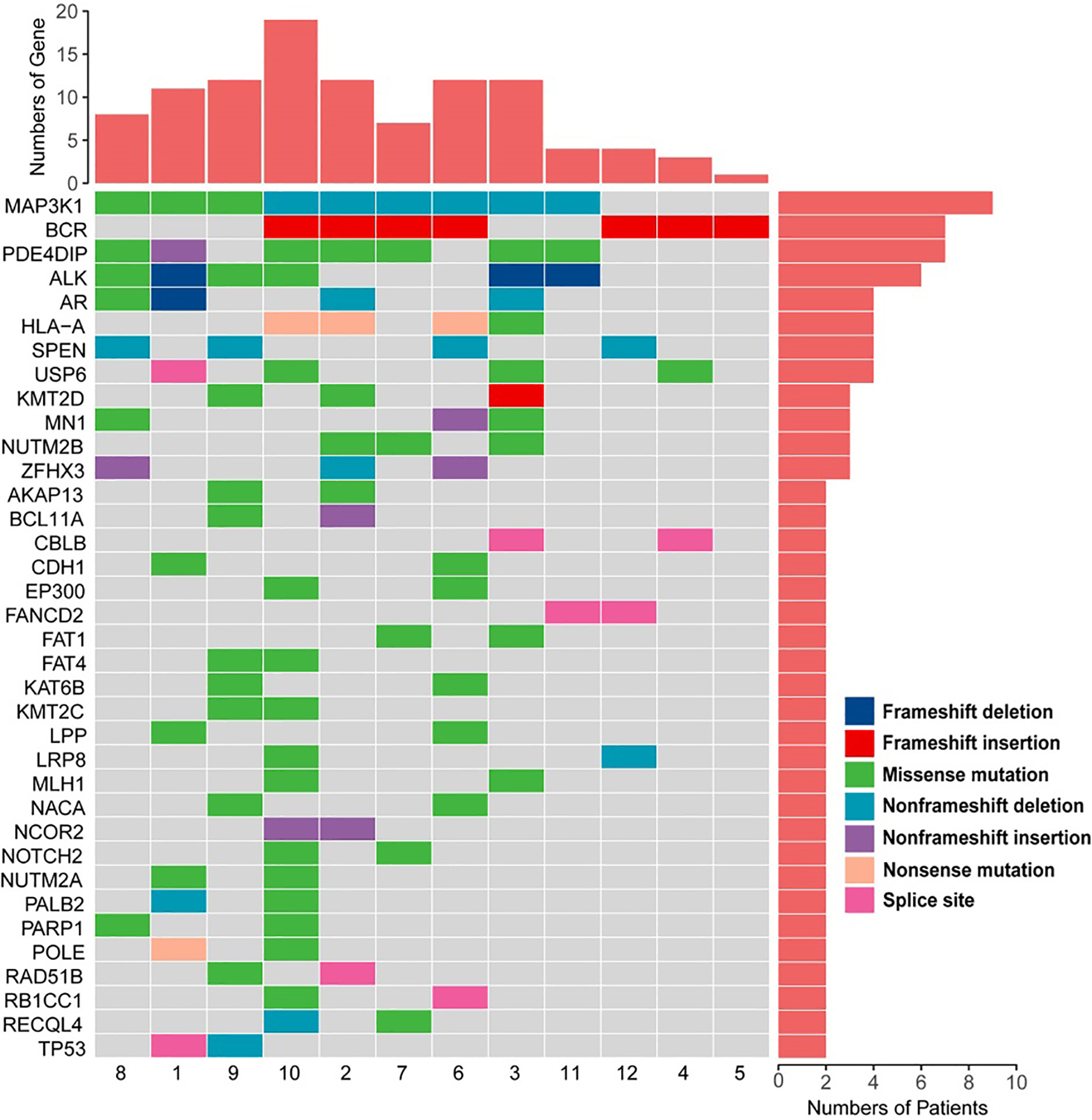
Moreover, multiple somatic mutations were found in PASC cancer tissues that did not occur in paracancerous tissues. A total of 488 somatically mutated genes were identified (Supplemental Table). Figure 2 shows the genes with somatic mutations found in at least two samples. The majority of the mutations (74.3%; 52/70) were missense mutations. TP53, as expected, was the most frequently mutated gene in tumours, followed by KRAS, OBSCN, and HRNR. TP53 had 3 missense mutations, 1 nonsense mutation, and 1 inframe deletion; all mutations in KRAS, OBSCN, and HRNR were missense mutations. Among the genes with somatic mutations in two or more samples, pathogenic variants were found in TP53, KRAS, TGFBR2, KLF17, and CASP9.
Associations between gene mutations and clinical characteristics and outcomes were analysed (Fig. 3 and Table 2). The results show that germline mutations in USP6 are negatively correlated with lymphatic metastasis (4/4 patients without lymphatic metastasis vs. 1/8 patients with lymphatic metastasis;
Association between gene mutations and clinical characteristics and outcome
Association between gene mutations and clinical characteristics and outcome
The values in this table were the
Somatic mutation spectrum in 12 PASC samples with 32 genes that were identified in at least two samples.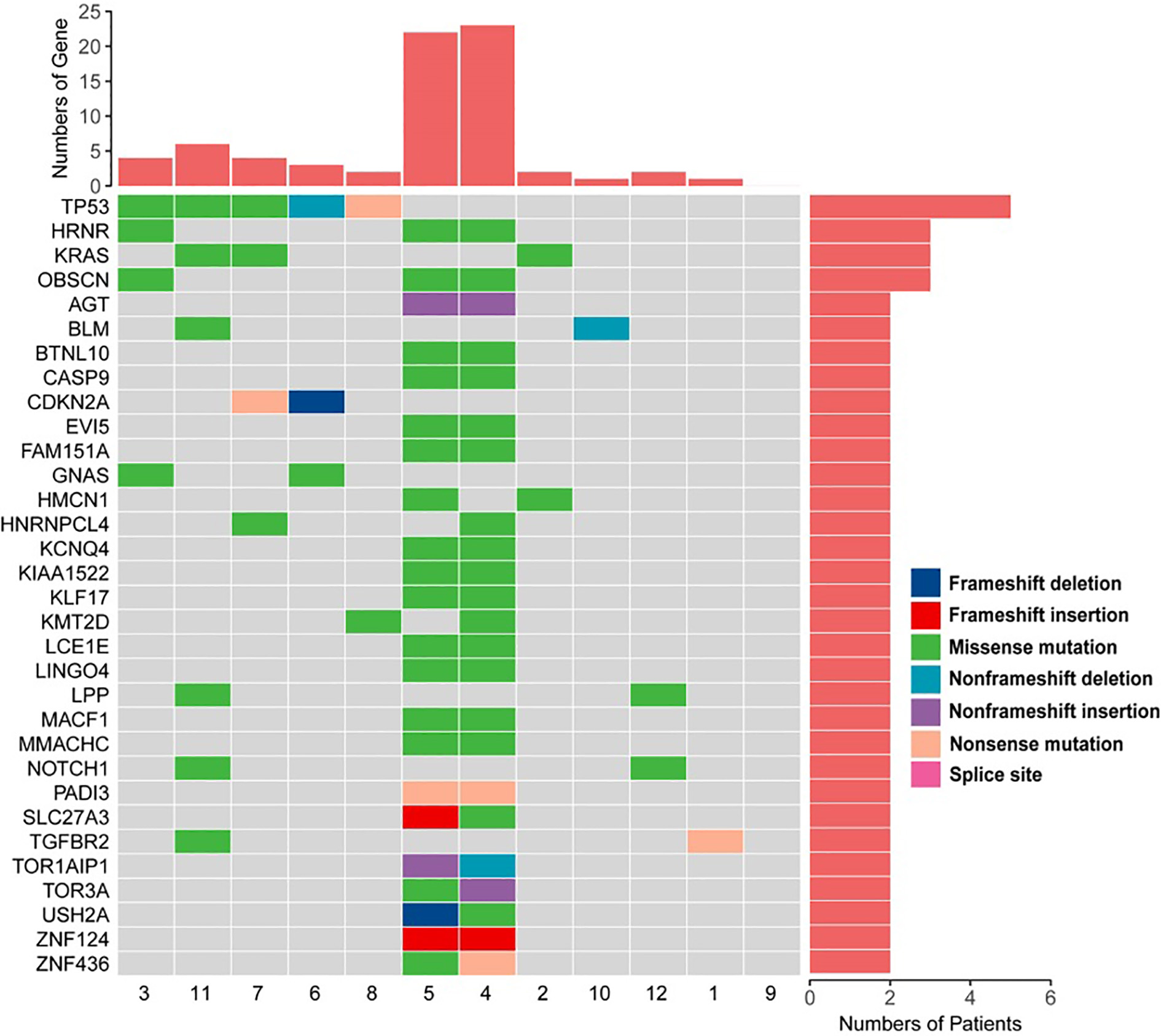
Integrative clustering of key mutated genes in 12 pancreatic adenosquamous carcinoma patients. Clinical features: gender, outcomes, lymphatic metastasis, and peripheral tissue invasion.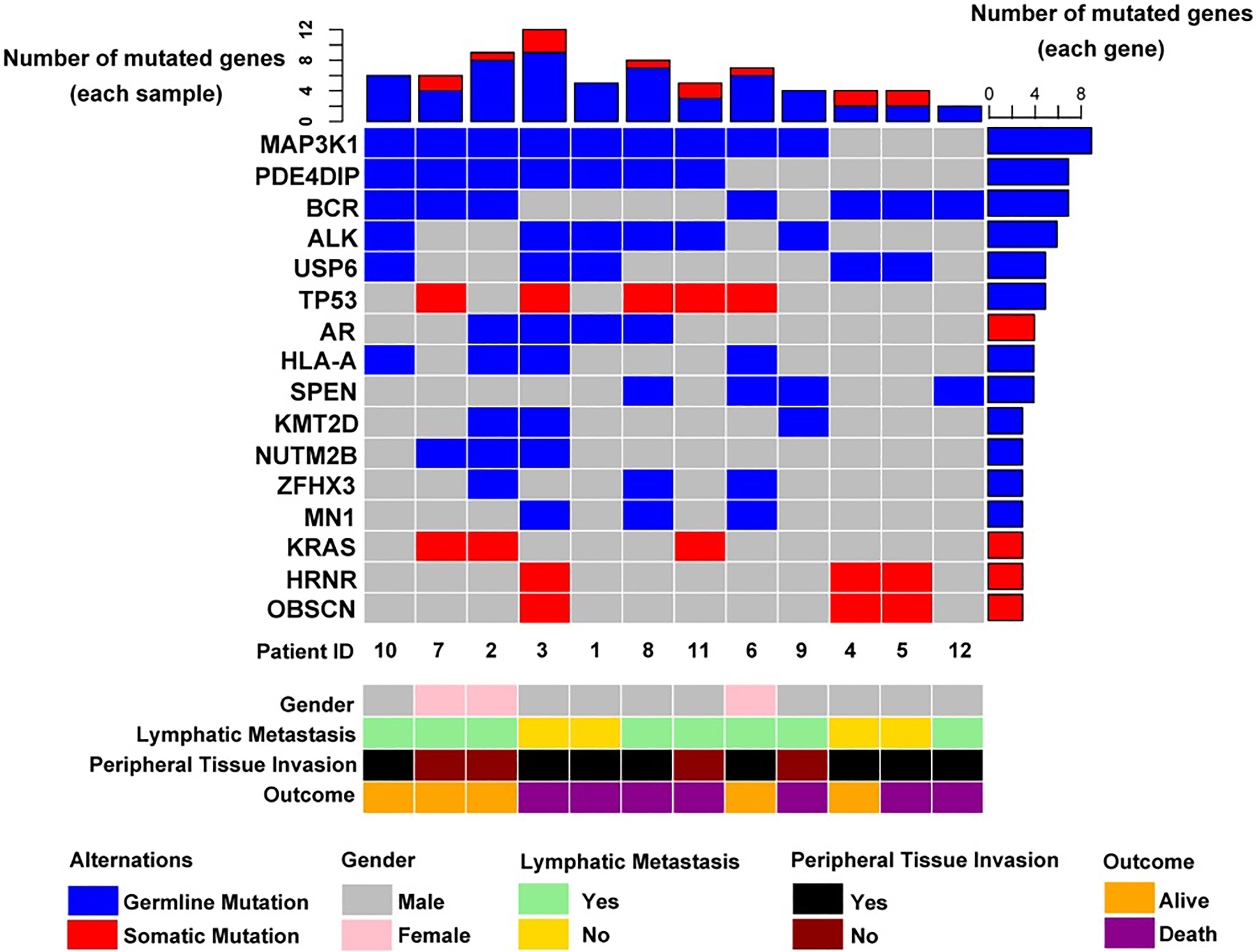
As a rare tumour type, limited studies have been performed regarding the genomic features of PASC [6, 19]. In the present study, we performed a comprehensive analysis of germline and somatic alterations in 12 pathologically diagnosed PASC patients by whole exome sequencing. To our knowledge, this is only the second study of PASC in a relatively large population.
Compared to the research by Fang et al. [6], the present study identified more PASC susceptibility genes. There are approximately 21 well-recognized pancreatic cancer susceptibility genes, including KRAS, TP53, SMAD4, MLL3, TGFBR2, ARID1A, SMARCA4, PBRM1, SF3B1, PCDH18, MAP2K4, EPC1, ATM, ARID2, CREBBP, AHANK, ANK3, ACVR1B, EP300, PCSK6, and ARHGAP3, among which three genes including KRAS, TP53 and SMAD4 were also identified as the most frequently mutated genes by Fang et al. [6]. Germline mutations in TP53, EP300, SMARCA4, and CREBBP were also detected in PASC patients at a low frequency, suggesting that these genes are also PASC susceptibility genes. We found an additional 12 germline mutated genes occurring in three or more PASC patients. The top three frequently mutated genes were MAP3K1, PDE4DIP, and BCR. All three genes had novel germline variants related to PASC.
MAPKs (mitogen-activated protein kinases) are key mediators of signalling pathways and are involved in multiple aspects of cell physiology [20, 21]. Dysregulation of MAPK-mediated signalling is a common event in tumorigenesis. MAP3Ks (mitogen-activated protein kinase kinase kinases) catalyse the first phosphorylation in a cascade of three sequential phosphorylation steps. MAP3K1 participates in the ERK and JNK pathways and the NF-
PDE4DIP encodes phosphodiesterase 4D interacting protein, a regulatory binding protein associated with calcium-mediated cAMP signalling (a pathway which is known to be dysregulated in cancer). Somatic mutations in PDE4DIP have been reported in some types of cancers, such as neuroendocrine Merkel cell carcinoma [28], ovarian cancer [29], malignant peritoneal mesothelioma [30] and pancreatic adenocarcinoma [31]. The malignant disease driven by the BCR-ABL oncogene is chronic myeloid leukaemia [32, 33]. Neither PDE4DIP nor BCR germline mutations have been reported in PASC patients. In fact, to our knowledge, mutations in the PDE4DIP and BCR genes are extremely rare in East Asian populations. Variants in these genes have not been reported in the dbSNP database yet. However, germline mutations in PDE4DIP and BCR were significantly enriched in the patients in this study, indicating that the susceptibility mutations might be unique to the Chinese population. We will focus on these genes in our future studies.
In addition to identifying germline and somatic alterations in patients with PASC, we also investigated the relationship between these mutations and clinical characteristics and outcomes. The results show that somatic mutations in KRAS are associated with diminished peripheral tissue invasion, and germline mutations in USP6 and somatic mutations in OBSCN and HRNR are associated with a decreased incidence of lymphatic metastasis. None of the identified mutations correlate with the outcomes in PASC patients; however, the small sample size, and varied efficacy of different therapies might have affected this analysis.
KRAS is a proto-oncogene encoding monomeric membrane-localised guanine nucleotide (GTP/GDP)-binding protein. Somatic mutations in KRAS have been reported in pancreatic cancer [25]. Other studies have shown that most pancreatic ductal adenocarcinoma patients and PASC patients carry KRAS activating mutations [34, 35, 36]. Consistently, we also identified somatic mutations in KRAS in our cohort. These findings suggest that mutations in KRAS are a common molecular feature of pancreatic cancer. Rachagani et al. reported that knockdown of the activated Kras
The OBSCN gene encodes obscurin, which belongs to a family of giant cytoskeletal proteins. Mutations in OBSCN are common in cancers [39, 40, 41], but not in PASC. We found somatic mutations in OBSCN in PASC patients, and these were negatively correlated with lymphatic metastasis. Altered expression of obscurin was detected in breast cancer tumour tissue rather than in matched normal tissue. Furthermore, loss of obscurin causes breast cancer cells to fail to form adhesion junctions, thus promoting tumour metastasis [41]. Whether the somatic mutations in OBSCN lead to a similar outcome in PASC should be cautiously investigated.
However, it should be noted that there were several limitations in this study. Firstly, the number of samples is a little small. PASC is a rare, clinically aggressive histological tumour, which constitutes only 1–4% of pancreatic carcinomas, and therefore, it is very difficult to collect PASC samples. There were only 12 samples included in this study. Considering the small sample size, our results should be further confirmed in a validation cohort or by experimental methods. Although the limited samples may bias results, we believe that this study has taken an important step towards revealing the underlying pathogenesis of PASC, and may aid in the determination of novel therapeutic targets. Secondly, the relationship between the identified mutations and PASC development/progression is only basing on data analysis with the absent of functional study. Whether these additional mutations are biologically meaningful should be further validated in vitro or in vivo models. We hope that these useful clues will help other researchers to carry out relevant research.
Conclusions
Three PASC susceptibility genes were identified: MAP3K1, PDE4DIP, and BCR. Additionally, we found negative associations between tumour metastasis and germline mutations in USP6 and somatic mutations in OBSCN and HRNR, as well as a negative correlation between peripheral tissue invasion and somatic KRAS mutations.
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-190236.
sj-xlsx-1-cbm-10.3233_CBM-190236.xlsx - Supplemental material
Supplemental material, sj-xlsx-1-cbm-10.3233_CBM-190236.xlsx
Footnotes
Acknowledgments
This work was funded by the National Natural Science Foundation of China (Grant NO. 81672830, 81502054, 81871992 and 61905279), PLA Youth Talent Training Program Foundation (Grant NO. 16QNP095), Scientific Research Project of Shanghai Municipal Commission of Health and Family Planning (Grant NO. 201840054), Scientific Research and Innovation Project of Shanghai Municipal Education Commission (Grant NO. 2017-01-07-00-07-E00012) and China’s National Key Research and Development Project (Grant NO. SQ2019YFC130043).
Conflict of interest
The authors declare no conflict of interest.
