Abstract
Introduction
Neuroblastoma (NB) is one of the children’s most common solid tumors, accounting for approximately 8% of pediatric malignancies and 15% of childhood cancer deaths. Somatic mutations in several genes, such as
Methods
This study analyzed 219 whole-exome sequencing datasets with somatic mutations detected by MuTect from paired normal and tumor samples.
Results
We prioritized mutations in 8 candidate genes (
Conclusion
Our study suggests 2 novel variants of
Keywords
Introduction
With the development of precision medicine, discriminating genomic factors have gained prognostic and therapeutic implications. Many gene mutations, including somatic and germline mutations, are identified using whole-exome, genome, or transcriptome sequencing. Both have improved our understanding of carcinogenesis and influenced the development of cancer treatment plans, including neuroblastoma (NB). NB is one of the most common solid tumors in children, accounting for approximately 8% of all pediatric malignancies and 15% of childhood cancer deaths.
1
A four-center case-control study indicated that LIN28A SNPs (single nucleotide polymorphisms), especially rs34787247 G>A, may increase NB risk.
2
In contrast, a multi-center case-control study using 263 cases and 715 controls to examine the association of
Whole-exome sequencing (WXS, also known as WES) is a genomic technique that is gradually being optimized to identify mutations in increasing proportions of the protein-coding regions of genes.
6
It is now routinely used and has revealed some rare and common gene variants in NB.
7
The high-level amplification of
Methods
Datasets
All of the sequencing data were obtained from the National Cancer Institute (NCI) Office of Cancer Genomics Therapeutically Applicable Research To Generate Effective Treatments (TARGET) NB project (https://ocg.cancer.gov/programs/target, assessed on 6 October 2020). The datasets were downloaded from The Cancer Genome Project (TCGA) Genomic Data Commons (GDC) Data Portal (https://docs.gdc.cancer.gov, assessed on 6 October 2020) using the GDC data transfer tool (https://gdc.cancer.gov/access-data/gdc-data-transfer-tool, assessed on 6 October 2020). A total of 127 RNA-seq and 219 WXS files were downloaded from the site. Of the downloaded sequences, both RNA-seq and WXS were available for 85 patients. In addition, 219 variant call format files (.vcf, specifically WXS.mutect2.raw_somatic_mutations.vcf) created using MuTect2 16 were downloaded for the patients on which WXS was performed to compare to our analysis of the raw sequencing files.
Variant Analysis for WXS Samples
Each VCF file from the WXS samples, ANNOVAR,
17
annotated variants (Figure 1A) was created. For each annotated VCF, we filtered the variants that were not exonic, synonymous SNVs, and those for which the maximum frequency in the population is > .001 in gnomAD.
18
We also filtered out the variants that had flags such as “alt_allele_in_normal,” “panel_of_normals,” or “germline_risk” in MuTect2
16
and required the remaining variants to have 'PASS' flags in more than one of the 219 WXS samples. Ultimately, using these criteria, we obtained 9 variants of the 8 genes. Workflow of the search for variants through filtering and analysis. The whole transcriptome sequencing (WTS): RNA-seq data. Exome sequencing (also known as whole-exome sequencing, WES, or WXS) is a technique for sequencing all the expressed genes in a genome. Population AF is the maximum allele frequency in the population obtained from GnomAD (the genome aggregation database). (A) Pipeline for filtering variants in WXS exome data from 219 patients. (B) Pipeline to call variants from 127 WTS RNA-seq datasets to analyze variants in the 8 genes for which variants meeting the criteria were found in (A). Shapes in dots: Inputs; boxes in dashes: outputs.
Variant Analysis for RNA-Seq Samples
As shown in Figure 1B, we wrote a python pipeline to call variants from 127 RNA-seq samples with BAM files as input [Binary Alignment Map (BAM) is the comprehensive raw data of genome sequencing; BAM is the compressed binary representation of SAM (Sequence Alignment Map), a compact and index-able representation of nucleotide sequence alignments]. We first used Picard 19 to mark duplicates and then used Genome Analysis Toolkit (GATK) 20 to split Reads with N in Cigar, generated and applied a recalibration table for Base Quality Score Recalibration (BQSR), and used HaplotypeCaller to call variants. After that, variants were filtered using GATK 20 with the options “-window 35 -cluster 3 -filterName Filter -filter ‘QD <2.0’ -filterName Filter -filter ‘FS >30.0.’” The filtered variants were annotated with ANNOVAR 17 for further analysis. Similarly, we filtered out non-exonic or synonymous SNV variants with >.001 maximum frequency in the population.
Additional methods included assessing the allele-specific or allele-imbalanced gene expression levels and manually examining the alignment files to locate candidates. We also obtained the GTeX data and the cBioPortal (The cBioPortal for Cancer Genomics website, https://www.cbioportal.org/, assessed on 6 October 2020) for comparative analyses.
Results
Clinical Characteristics
Clinical Characteristics in WXS- and WTS-Identified NB Samples.

Whole-Exome Sequencing Identifies Candidate Genes
A List of 9 Mutations Detected from MuTect2, Its Functional Impacts (To Protein), Population Frequency, Number of Occurrences in the Current Data Set, and the Respective Alt/Ref Reads Count.

RNA-Sequencing Analysis of Candidate Gene Allelic Expression
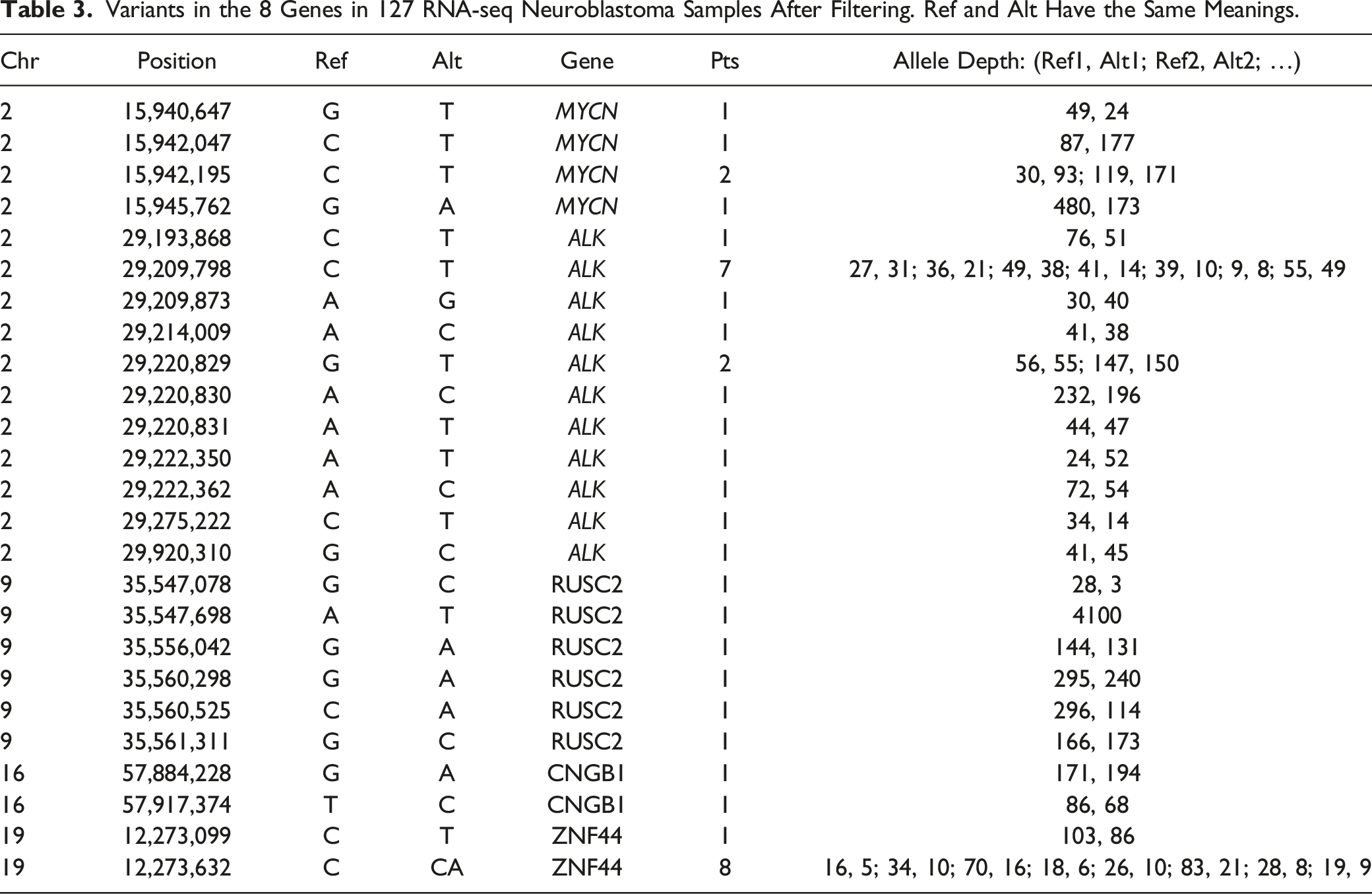
Variants in the 8 Genes in 127 RNA-seq Neuroblastoma Samples After Filtering. Ref and Alt Have the Same Meanings.

Patterns of 2
Moreover, among these mutated sites, we observed 2 novel variants of ZNF44 variant patterns by IGV plot for chr19:12,273,632/C>CA in the 
Discussion
NB is a solid tumor that can develop from immature nerve cells in several areas of the body. It most commonly affects children and rarely occurs in adults.
1
This study analyzed WXS and RNA-seq data from NB patients to identify somatic mutations and their allele-specific expression. As most somatic mutations are identified from DNA-seq techniques such as WXS, the allelic expression of those mutations is often unknown. Proteins, the functional units of a live cell, are made from mRNA, so a somatic mutation may have very different effects on the cellular function that vary with its allelic expression profile. Our study confirmed multiple known NB mutations and identified
Our study explored 2 cohorts of NB patients with either WXS or WTS data available. The overlapping rates of the 2 cohorts were high: 38.8% (85/219) of the WXS group and 66.9% (85/127) of the WTS (Whole transcriptome sequencing) group. It has been reported that using WXS identification, mutation frequencies of somatic genes, including
As expected, there were some differences between WTS and WXS analysis. WXS identified a gene
WTS has been hailed as a promising approach with distinct advantages, especially for determining transcriptome characteristics.
25
However, WTS is not suitable for the discovery of DNA mutations. Thus, combining WXS and WTS can provide complementary perspectives on gene mutations. In our study, an interesting finding from RNA-seq analysis was that in 1 patient sample harboring 2 different Proposed workflow of pediatric cancer for lifetime management. Collective information is based on the publications26-29 related to the single-cell subclonal evolution of pediatric tumors. This workflow might track 2 new variants of ZNF44 and validate them as a novel candidate driver gene for neuroblastoma as details in the discussions.
Highly relevant to our study, an Italian group aimed to determine the differential genetic landscapes between short survival (SS) and long survival (LS) in high-risk (HR) neuroblastoma (NB) (HR-NB) patients at stage M.
5
The significant percentage of patients who demonstrate rapid disease progression despite multimodal treatment presents one of the biggest problems for oncologists treating high-risk (HR-NB) patients. About 60% of these HR-NBs develop fatal conditions within 5 years of diagnosis. They focused on a cohort of stage M NB patients from the Italian NB Registry with complete clinical data, and follow-up over 10 years was considered, including SS (n = 14) and LS (n = 15). They found ZNF44 mutations in only 2 SS patients (#1965, #2578), about 14%, but not in LS patients. The percentage of mutated ZNF44 in the total of patients at the M stage, including SS and LS, is about 6%. They pointed out, “In SS patients, 4 genes (SMO,
Nevertheless, ZNF44 had been barely investigated in previous research, even though the identification of gain-of-function mutations in the
In summary, to date, few studies have explored gene mutations in NB, like simultaneously using both WXS and WTS. Our study revealed that these 2 methods present different perspectives and meaningful results. Specifically, we found that allele-specific expression assessed by RNA-seq can be quite different even for the same gene mutations, which underscores the importance of WTS in cancer research. Furthermore, we identified gene mutations through both methods, validating some well-known NB genes such as
Conclusion
We discovered 2 novel
Supplemental Material
Supplemental Material - RNA-Sequencing Combined With Genome-Wide Allele-Specific Expression Patterning Identifies ZNF44 Variants as a Potential New Driver Gene for Pediatric Neuroblastoma
Supplemental Material for RNA-Sequencing Combined With Genome-Wide Allele-Specific Expression Patterning Identifies ZNF44 Variants as a Potential New Driver Gene for Pediatric Neuroblastoma by Lan Sun, Xiaoqing Li, Lingli Tu, Andres Stucky, Chuan Huang, Xuelian Chen, Jin Cai, and Shengwen C. Li in Cancer Control.
Footnotes
Acknowledgments
We thank Jiang F. Zhong for his guidance and critical reading of the manuscript.
Author Contributions
Writing—initial draft preparation, XL and LS. Writing—review and revision, SCL; Conceptualization, JC, SCL; Methodology and formal analysis, LT and CH; Software and data curation, AS and XC. All authors have read and agreed to the published version of the manuscript.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study is partly supported by the Natural Science Foundation of Chongqing, China (cstc2020jcyj-msxmX1063). This work was also supported in part by the CHOC Children’s–UC Irvine Child Health Research Awards #16004004, CHOC-UCI Child Health Research Grant #16004003, and CHOC CSO Grant #16986004.
Ethical Approval
The data sets came from a public database without ethical issues. Specifically, all of the sequencing data were obtained from the National Cancer Institute (NCI) Office of Cancer Genomics Therapeutically Applicable Research To Generate Effective Treatments (TARGET) neuroblastoma project (https://ocg.cancer.gov/programs/target, assessed on 6 October 2020). The datasets were downloaded from The Cancer Genome Project (TCGA) Genomic Data Commons (GDC) Data Portal (https://docs.gdc.cancer.gov, assessed on 6 October 2020) using the GDC data transfer tool (![]() , assessed on 6 October 2020).
, assessed on 6 October 2020).
Data Availability
The results published here are based on data generated by the Therapeutically Applicable Research to Generate Effective Treatments (https://ocg.cancer.gov/programs/target, assessed on 6 October 2020) initiative phs000467. The data used for this analysis are available at ![]() .
.
Supplemental Material
The results published here are based upon data generated by the Therapeutically Applicable Research to Generate Effective Treatments (https://ocg.cancer.gov/programs/target) initiative, phs000467. The data used for this analysis are available at ![]() . Supplemental material for this article is available online.
. Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
