Abstract
BACKGROUND:
Breast cancer is a worldwide leading cause of cancer mortality and it is associated with numerous tumor suppressor genes and oncogenes. Growing evidence exists that different KLFs play pivotal roles of in human malignancies. However, the function of KLFs in breast cancer development has remained uncovered.
OBJECTIVE:
To explore the potential prognostic biomarkers among KLFs in breast cancer.
METHODS:
In the present study, by using multiple large open databases, such as Oncomine database, Kaplan-Meier Plotter, and bc-GenExMiner online software, we deeply analyzed the expressions and clinical values about KLFs in patients with breast cancer.
RESULTS:
KLF4/5/8/9/10/15 were significantly down-regulated in breast cancer samples. KLF11 exerts significantly negative effect on the prognosis of patients, whereas expressions of KLF4/15 were associated with better prognosis. Moreover, the vital genes KLF4/11/15 showed significant association with clinical parameters including age, estrogen receptor, progesterone receptor, epidermal growth factor receptor-2, Scarff-Bloom-Richardson grade, and Nottingham prognostic index.
CONCLUSIONS:
Bioinformatics analysis suggested that KLF4/11/15, compared to other KLFs, might be potential prognostic indicators and treatment targets for breast cancer patients.
Expressions of KLFs in 20 types of cancer vs. corresponding normal tissues using the Oncomine database. Red and blue represent the numbers of datasets with statistically significant mRNA high-expression and low-expression of KLF family genes, respectively. Statistically significant values and fold change were demarcated as 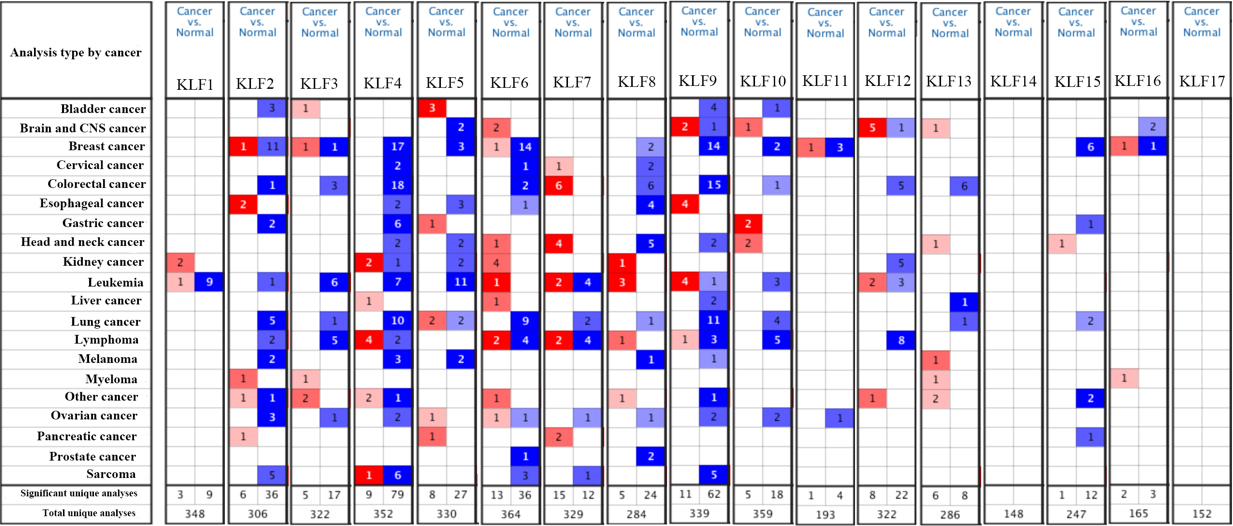
Breast cancer, the most common malignant tumor, is the second cause of cancer-related deaths in women [1]. Breast cancer is associated with inactivation of thousands of tumor suppressor genes and activation of oncogenes [2]. Although advances have been achieved in early diagnosis and systematic therapies, a portion of cases still relapse and present with metastases. It is a fascinating strategy to explore more valuable biomarkers for the diagnosis, treatment and prognosis of patients with breast cancer. The breast cancer molecular subtypes relies on expression of estrogen receptor (ER), progesterone receptor (PR), epidermal growth factor receptor-2 (HER-2) and Ki-67, providing important prognostic and therapeutic information [3]. A highly conserved family of zinc finger transcription factors is involved in Krüppel-like factors (KLFs). As of currently, It is established that KLFs play an important role in the progress of diseases, especially in human malignant tumors, such as breast cancer, colon cancer and so on [4]. Although evidences about the core role of KLFs in cancers are accumulating, it is still not completely clear about the expression and status of KLFs in breast cancer. The complicated mechanisms of KLFs function in cancers might suggest novel targets for prognosis and therapy [5]. Therefore, identifying suitable targets from KLFs for prognostic analysis and individual therapy of breast cancer urgently needs attention all over the world. However, to our best of knowledge, the clinical significance of KLFs in breast cancer has not been explored yet using the databases available about cancer.
In the present study, by using multiple large open databases, such as Oncomine database, Kaplan-Meier Plotter, and bc-GenExMiner online software, we deep-ly analyzed the expressions and clinical values about KLFs in patients with breast cancer. Furthermore, we then selected more meaningful indicators from 17 KLFs for prognosis and therapy in breast cancer.
Materials and methods
ONCOMINE data-mining analysis
ONCOMINE is a public online cancer database for RNA and DNA sequences [6]. In the present study, firstly, it was used to facilitate data-mining the transcriptional expressions of KLFs in different kinds of cancer. We compared RNA expressions of KLFs in breast cancer samples with those in normal samples using Student’s t-test. Statistically significant values and fold change were defined as
Kaplan-Meier curve showed the better RFS based on mRNA level of KLFs. Breast cancer patients with increased expressions of KLF1/2/3/ 4/6/7/9/12/15/16/17 presented with better RFS.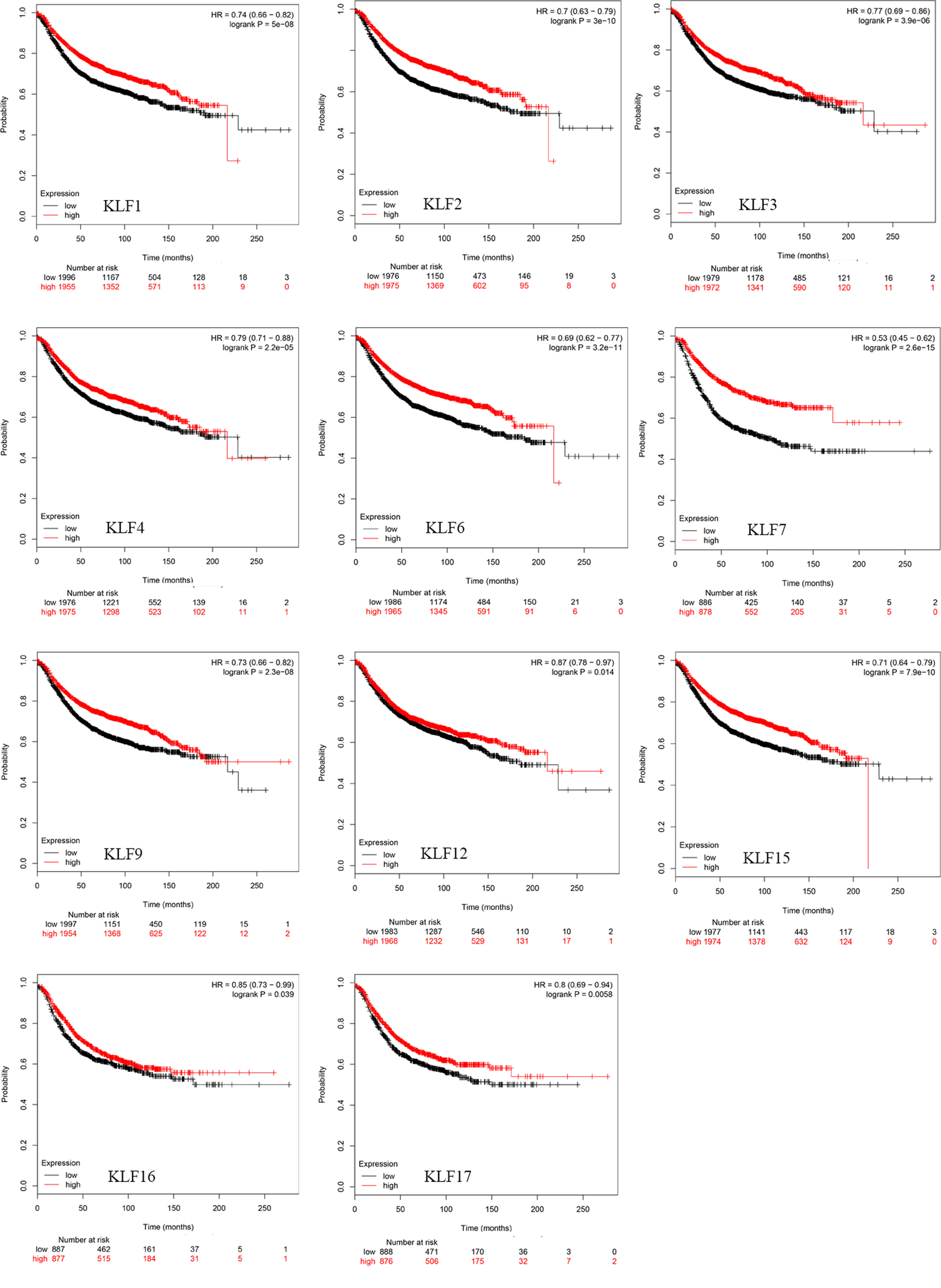
Breast Cancer Gene-Expression Miner v4.1 (bc-GenExMiner v4.1), a statistical mining tool of 264 independent datasets, is of three classical functions: gene expression, cancer prognosis, and correlation [7, 8]. All data were updated in October 2018. KLFs were analyzed with clinical parameters including age, nodal status, ER, PR, HER-2, Scarff-Bloom-Richardson (SBR) grade, and Nottingham prognostic index (NPI). We also analyzed the prognostic value of metastatic recurrence event in the prognosis module about KLFs in breast cancer and selected the potential novel prognostic markers.
Kaplan-meier plotter
The Kaplan Meier Plotter (
Prognostic association of KLFs expressions in breast cancer based on bc-GenExMiner v4.1. (event: metastatic recurrence)
Prognostic association of KLFs expressions in breast cancer based on bc-GenExMiner v4.1. (event: metastatic recurrence)
Dysregulated expressions of KLFs in breast cancer patients
We used ONCOMINE database to identify all the 17 KLF family genes. As shown in Fig. 1, the expressions of KLFs in tumor and normal tissues were compared in 20 kinds of cancer including bladder cancer, breast cancer and so on. The mRNA expressions of KLF4/5/8/9/10/15 were significantly down-regulated in breast cancer samples in multiple datasets (blue). In addtion, KLF4/15 were also lower expressed in gastric cancer and lung cancer samples. The mRNA expressions of KLF1/7/12/13/14/17 were neither up-regulated nor down-regulated in breast cancer tissues compared with normal samples and the expressions of KLF2/3/6/11 in breast cancer were different among multiple datasets in Oncomine.
Kaplan-Meier curve showed the worse RFS based on mRNA level of KLFs. Breast cancer patients with increased expressions of KLF5/8/11/13 presented with worse RFS.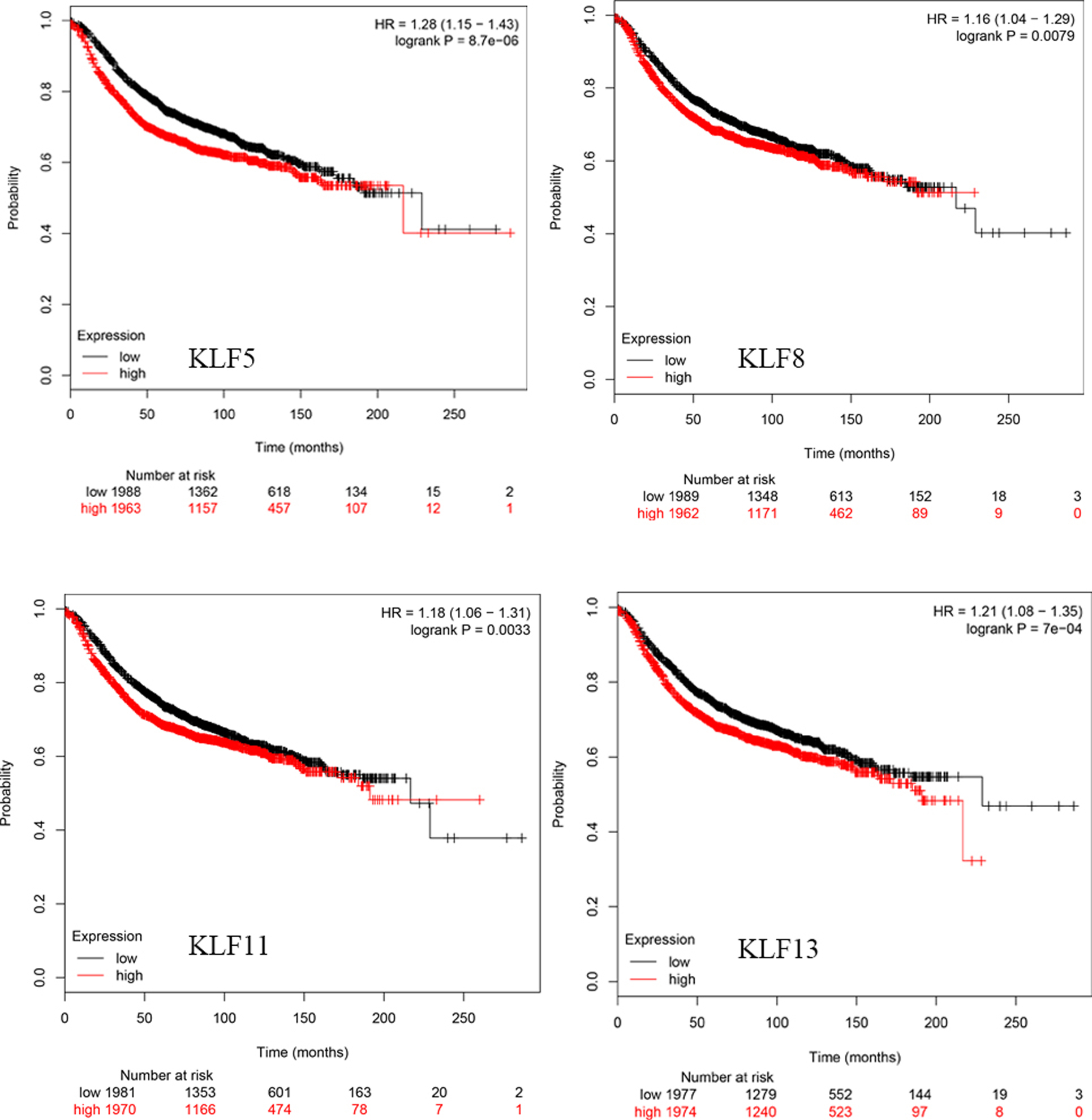
The Kaplan-Meier curves about survival information indicated that increased
KLF1/2/3/4/6/7/9/12/15/16/17 (Fig. 2) mRNA levels and reduced KLF5/8/11/13 (Fig. 3) mRNA levels are significantly associated with better RFS (
Prognostic association of KLFs expressions in breast cancer based on kaplan-meier plotter (event: relapse-free survival)
Prognostic association of KLFs expressions in breast cancer based on kaplan-meier plotter (event: relapse-free survival)
Kaplan-Meier curve showed the OS based on mRNA level of KLFs. (A) Breast cancer patients with increased expression of KLF7/12/15 presented with preferable OS. (B) Breast cancer patients with induced expression of KLF5/8/11 presented with worse OS.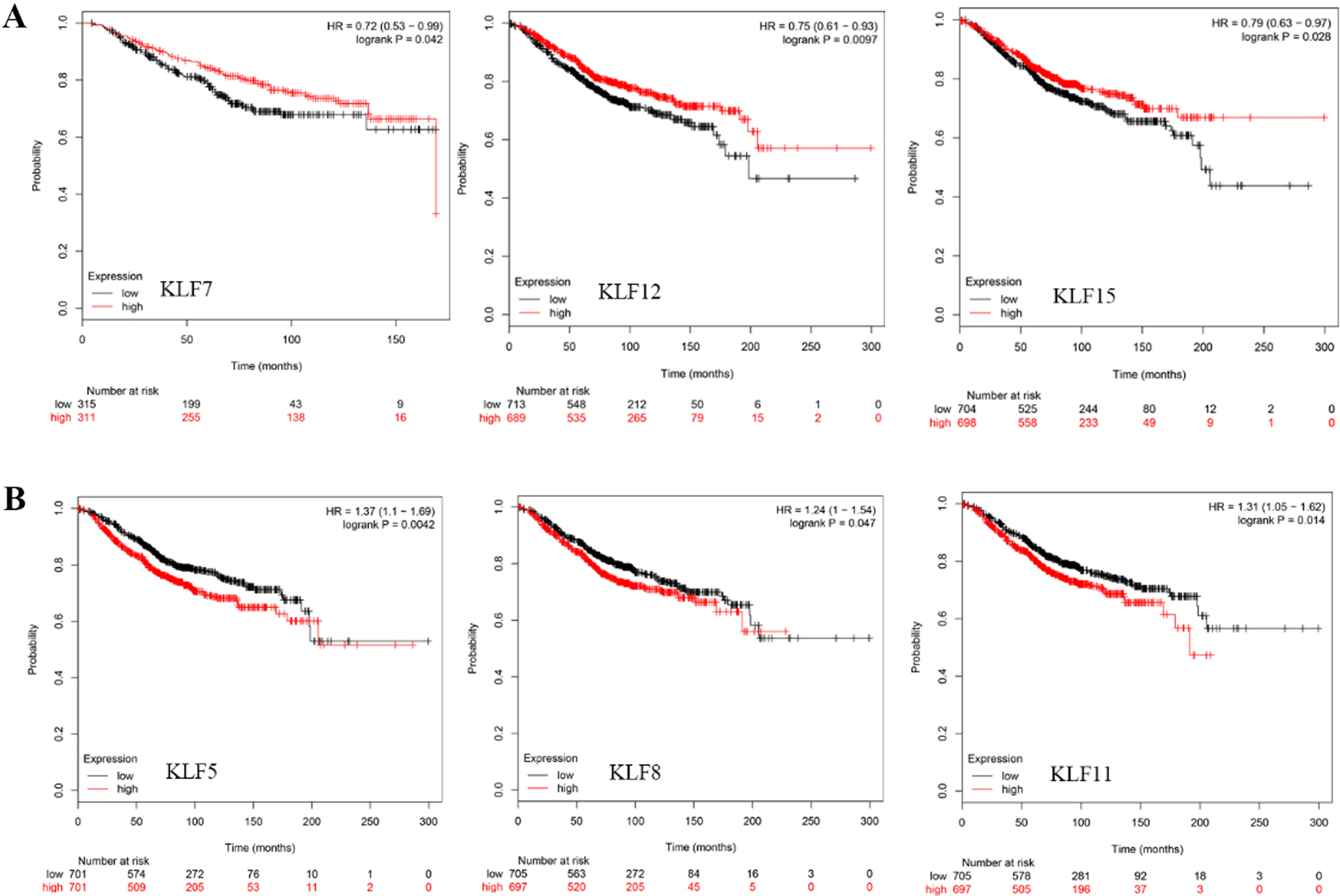
The relationship between mRNA expressions of KLFs with the SBR grade and the NPI value. (A) Patients with higher grade tumors (SBR2 and SBR3) tended to express higher levels of KLF11 and low levels of KLF4/15. (B) Patients with higher NPI value tended to express higher levels of KLF11and low level of KLF4.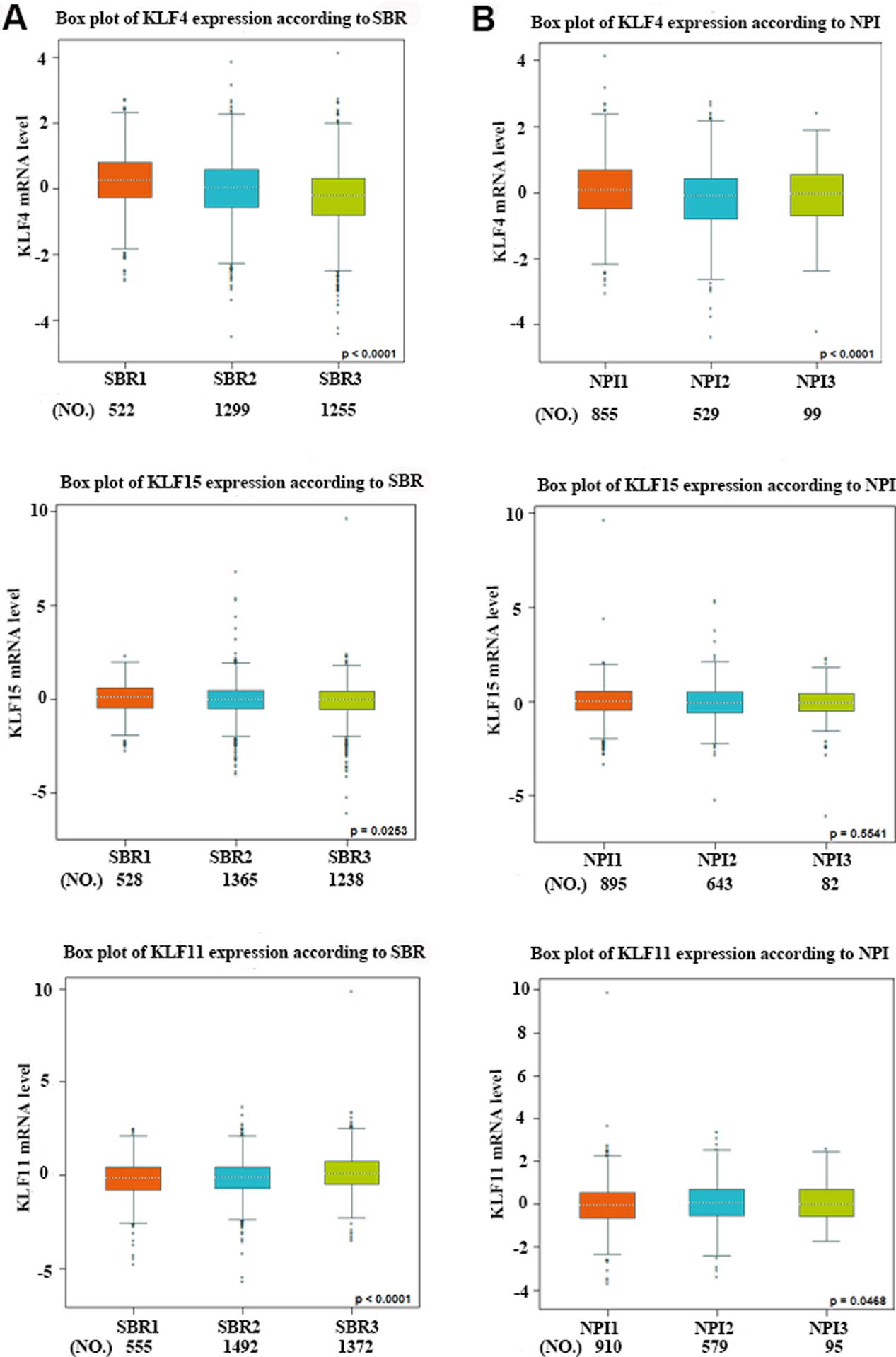
Prognostic association of KLFs expressions in breast cancer based on Kaplan-Meier Plotter (Event: Overall Survival)
Using the databases mentioned above, we next focused on whether KLF4/15 and KLF11 play core roles in breast cancer progression. Using bc-GenExMiner v4.1 software, we implemented Welch’s test to compare the abnormal expressions of KLF4/15/11 among different groups of patients according to clinical pathological features. The Scarff, Bloom and Richardson (SBR) grade is a widely accepted prognostic evaluation system, suggesting that patients with poorly differentiated tumors (e.g. SBR2 and SBR3) have a worse outcome compared with those who have well-differentiated tumors (SBR1) [10]. According to SBR criterion, patients with higher grade tumors expressed increased KLF11, and decreased KLF4/15, which were consistent with our previous finding (Fig. 5A). As we know, Nottingham Prognostic Index (NPI) is also a well-known prognostic model for breast cancer patients [11]. Higher expression of KLF11 and lower expression of KLF4 were correlated with higher NPI value, which were also consistent with our previous data (Fig. 5B).
Considering age criterion (Fig. 6A), no significant difference between
The relationship between mRNA expressions of KLF4/15/11 with age (A), nodal status (B) and basal-like status (C) of breast cancer using the bc-GenExMiner online software.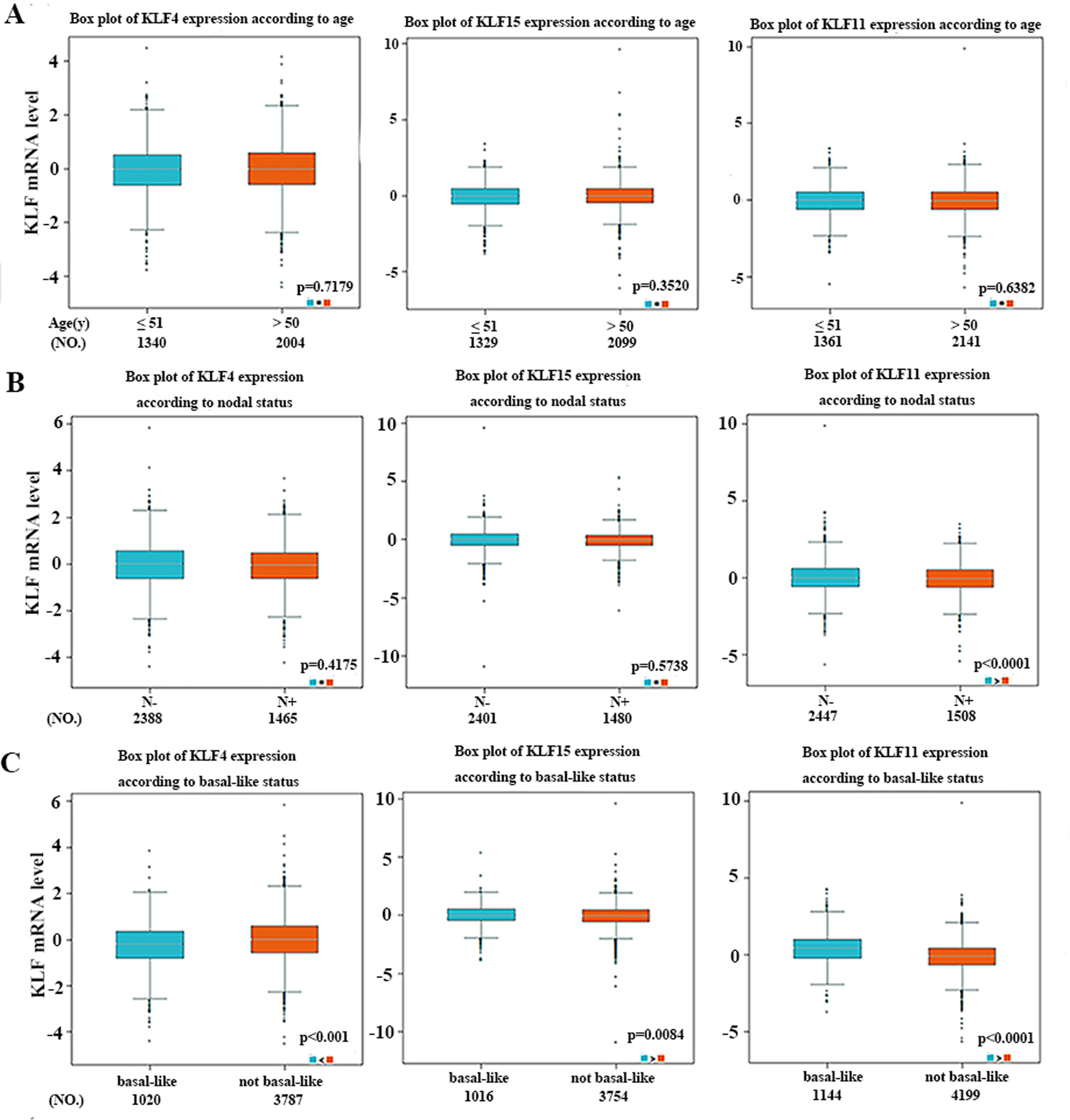
The relationship between mRNA expressions of KLF4/15/11 with protein expressions of ER (A), PR (B), and HER-2 (C) of breast cancer using the bc-GenExMiner online software.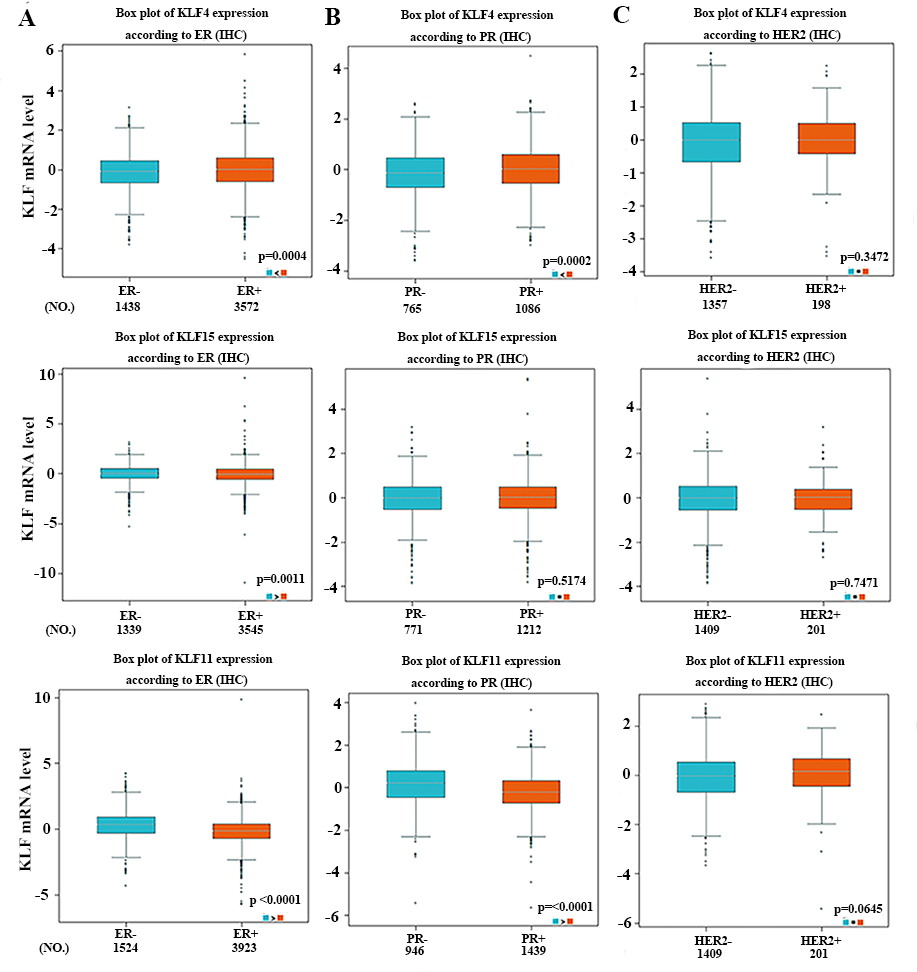
Although a general statement for KLFs in cancer progression have been uncovered recently, most relationships between KLFs and cancers have not been fully characterized. Therefore, we firstly need a thorough scan to determine the expression of KLFs in different cancer types. In the present work, the expressions of total 17 KLFs were compared in breast cancer samples and normal tissues in Oncomine database, including down-regulated KLF4/5/8/9/10/15 in breast cancer samples, and KLF1/7/12/13/14/17 with no significant reported dysregulation. Using the Kaplan-Meier survival analysis, we found that increased KLF1/ 2/3/4/6/7/9/12/15/16/17 were associated with preferable RFS and (or) OS whereas increased KLF5/8/11/13 were correlated with worse RFS and (or) OS. Besides, bc-GenExMiner online software provided metastatic recurrence data showing that KLF4/15 were related to better prognosis and higher expressions of KLF6/10/11 /16 exerted negative effects on patient prognosis. Therefore, combination of the above results shows that KLF4/11/15 could be identified as potential good/bad prognostic targets of breast cancer. Interestingly, we also found that KLF4/15 were down-regulated in gastric and lung cancer samples, suggesting that KLF4/15 might be prognostic targets for other cancers. However, multiple bioinformatics methods are required to investigate the role of KLF4/15 in gastric cancer and lung cancer.
KLF4 is an epithelium enriched regulator, and it has been reported to act as an oncogene in oral and skin squamous cell cancers [12, 13]. However, the role of KLF4 remains controversial in breast cancer. Several studies have demonstrated that KLF4 acted as an oncogene in breast cancer [14] while tumor suppressive role of KLF4 in breast cancer has also been found [15]. Moreover, KLF4 expression levels were inversely correlated with tumor grade [16]. In our study, analyses of the Oncomine database we found that mRNA level of KLF4 in breast tumor tissues was lower than that in normal ones. What’s more, we found that KLF4 was related to better prognosis by using both Kaplan-Meier survival analysis and bc-GenExMiner online software. So, KLF4 could be identified as potential good prognostic targets of breast cancer reasonablely.
Recent studies reported that KLF15 were expressed in various tissues, including kidney, liver, heart, and so on. It is also named as kidney-enriched KLF (KKLF) [17, 18]. However, its role in human malignant tumors has been reported scarcely. It has been reported that KLF15 might exert anti-growth effects on multiple cancer cells, including pancreas cancer and breast cancer [19, 20]. Moreover, KLF15 has been reported to decrease cell proliferation in the human breast cancer cell line T47D in an estrogen-dependent pathway [20]. Hence, KLF5 might play a role in regulating breast cancer proliferation. Our study also confirmed that high levels of KLF15 conferred a RFS and OS advantage in breast cancer patients and it may be a promising prognostic indicator.
KLF11 was first found in osteoblast cells. Since then, in is reported that KLF11 was also widely expressed in multiple tissues such as breast, lung, kidney, colon, and pancreatic tissues [21]. It has been well studied that KLF11 is involved in the development of many kinds of human tumors, such as ovarian and pancreatic cancer [22]. However, further studies are necessary to uncover how KLF11 is involved in breast cancer biology and progression because there is little information on KLF11 in breast cancer. In our study, higher expression of KLF11 exerted negative effects on patient prognosis, which confirmed by bc-GenExMiner online software and Kaplan-Meier Plotter. Higher levels of KLF11 indicated worse RFS and OS in all breast patients, suggesting that KLF11 might be a potential therapeutic target.
In addition, we found that basal-like breast cancer individuals expressed significantly increased KLF11 and reduced KLF4. Comparing to ER-negative or PR-negative tumors, hormone receptor positive tumors tended to express lower level of KLF11 and high level of KLF4. As we know, ER-positive and PR-positive breast cancer patients generally have a better prognosis. Therefore, these results indicated that lower expression of KLF11 and higher expression of KLF4 may predict a preferable prognosis. Nevertheless, KLF15 mRNA expression was significantly higher in basal-like breast cancer patients and ER-positive tumors tended to express lower levels of KLF15 compared with ER-negative ones. Those results above contradicted our above findings of KLF15 as tumor suppressors. Such difference might be explained by the fact that KLF15 expression in the database were originated from RNA sequences and that clinical samples and experimental conditions were quite different. Further investigations are required to more precisely elucidate the biological behaviors of KLF15. Of note, patients with higher grade tumors express higher levels of KLF11, and lower levels of KLF4/15, consistent with our previous founding respectively. Moreover, we found that higher expression of KLF11 and lower expression of KLF4 were associated with higher NPI value, also consistent with our previous data.
In the current study, we deeply analyzed expressions and clinical values for all 17 KLFs in breast cancer. Bioinformatics analysis suggested that KLF4/11/15, compared to other KLFs, might be potential prognostic indicators and treatment targets for breast cancer patients. KLFs expression and activity are altered in human cancers, and individual KLFs can be tumor suppressors or oncogenes, often with context-dependent functions depending on tissue, tumor type or cancer stage. The study might supply a more comprehensive understanding of the complicated dysregulation in the molecular biology of breast cancer. However, other multiple in-vivo experiments and more in-depth researches are required to confirm the prognostic and therapeutic potentials of these KLFs.
Footnotes
Acknowledgments
This work was financially supported by Natural Science Foundation of China (81502294 and 81602336) and the Project of Jiangsu Province Traditional Chinese Medicine Bureau (YB201952).
Conflict of interest
The authors declare no conflict of interest.
