Abstract
Long non-coding RNAs (lncRNAs) are non-protein-coding transcripts that play important roles in tumorigenesis and tumor progression. Our study aimed to explore the role of lncRNA MAGI2-AS3 in breast cancer metastatic progression. In the present study, our results showed that MAGI2-AS3 can inhibit the migration and invasion of breast cancer cells. In addition, an increase in MAGI2-AS3 can inhibit microRNA-374a (miR-374a) expression in breast cancer cells. Bioinformatic analysis predicted the correlation between MAGI2-AS3 and miR-374a. Phosphatase and tensin homolog (PTEN) was found to be an novel mRNA target of miR-374a. MAGI2-AS3 upregulation inhibited breast cancer metastatic progression by decreasing miR-374a and enhancing PTEN expression. Together, our data revealed that lncRNA MAGI2-AS3 is involved in breast cancer cell progression by regulating the miR-374a-PTEN axis. These findings offer new insight into treatment strategies for breast cancer.
Introduction
Sequence of miR-374a mimics and negative miRNA mimic controls
Sequence of miR-374a mimics and negative miRNA mimic controls
Breast cancer is one of the most common cancers in women worldwide [1]. Despite advances in diagnosis and treatment, breast cancer patient survival rates remain low [2]. Approximately 252,710 new cases and 40,610 related deaths are expected to occur among US women in 2017 [3]. As the abnormal proliferation and invasion of breast cancer cells are major contributors to breast cancer patient death [4], it is important to understand these molecular mechanisms to improve clinical treatment.
Long non-coding RNAs (lncRNAs) are a group of RNA transcripts longer than 200 nucleotides that have no protein-coding capacity [5]. Recently, lncRNAs have been shown to be involved in various physiological and pathological processes, such as cell differentiation and apoptosis, X chromosome imprinting, and even tumorigenesis [6, 7]. Accumulating evidence suggests that abnormal lncRNA expression is often observed in various cancers and contributes to their initiation and development [8, 9]. In breast cancer, some lncRNAs have been identified as oncogenes or tumor suppressors and have potential as diagnostic or prognostic markers for breast cancer patients [10, 11].
MAGI2-AS3 is a newly discovered lncRNA, and its role and underlying mechanism are poorly understood in breast cancer. In the present study, we examined the biological functions of MAGI2-AS3 in breast cancer cell migration and invasion using a series of in vitro experiments. Moreover, we investigated the molecular mechanisms by which MAGI2-AS3 exerts its anti-oncogene role in breast cancer.
Cell culture and treatment
The human breast cancer cell lines MDA-MB-231 and MCF-7 were used in this study. All cells were obtained from Jiangsu University (Zhenjiang, Jiangsu, China). MDA-MB-231 and MCF-7 cells were cultured in high-glucose Dulbecco’s modified Eagle’s medium (DMEM, Thermo Fisher Scientific, Waltham, MA, USA) supplemented with 10% fetal bovine serum (FBS, Gibco-BRL, Carlsbad, CA, USA). All cells were maintained in a humidified atmosphere of 5% CO
Lentiviral vector production and infection
MAGI2-AS3 lentivirus particles (LV-MAGI2-AS3) and control lentivirus particles (LV-NC) were purchased from GenePharma Biotechnology (Shanghai, China). Lentivirus infection was performed according to the manufacturer’s instructions. Cells at 70% confluence were treated with 5
Transient miRNA transfection
A breast cancer cell line was selected for miR-374a transfection. miR-374a mimics and mimic negative controls (mimic NC) were synthesized and purified by GenePharma (Shanghai, China). The sequences are listed in Table 1. Briefly, the cells were grown overnight and then transfected with 100 nM miR-374a mimics or mimics NC using Lipofectamine
Wound healing assay
The breast cancer cells were seeded into 6-well plates and then allowed to grow until 100% confluence. Next, the cell layer was scratched through the central axis using a sterile plastic pipette tip, and loose cells were washed away with phosphate-buffered saline. Wound healing was observed and photographed at 0 and 24 h in three randomly selected microscopic fields for each condition and time-point. The degree of motility 24 h after confluent cells had been scratched was expressed as the percentage of wound closure as follows: (distance of cell migration at 24 h/width of scratch at 0 h)
Cell migration and invasion assays
The cell migration assay was performed using Transwell inserts (Corning, Blacksburg, VA, USA), and the cell invasion assay was performed with a CytoSelect 24-well cell invasion assay kit (Cell Biolabs, Inc., San Diego, CA, USA) according to the manufacturers’ instructions. First, the breast cancer cells were starved in DMEM without FBS for 2 h. Then, 5
Sequence of primers used for qRT-PCR
Sequence of primers used for qRT-PCR
Total RNA was isolated from tissues and cells using RNAiso Plus (TAKARA Biotechnology, Dalian, China) according to the manufacturer’s instructions. To measure MAGI2-AS3, miR-374a and phosphatase and tensin homolog (PTEN) mRNA expression, qRT-PCR was performed using a SYBR Green-based PCR kit (TAKARA Biotechnology, Dalian, China). An ABI Step- OnePlus RealTime PCR System (Applied Biosystems, Foster City, CA, USA) was used for the quantitative detection of lncRNA, miRNA and mRNA. The expression of each gene was quantified by measuring the cycle threshold (Ct) values and normalized using the 2
Western blotting
Proteins extracted from cells were separated by sodium dodecyl sulfate-polyacrylamide gel electroph-oresis (SDS-PAGE; Beyotime Institute of Biotechnology, Shanghai, China) and transferred onto polyvinylidene fluoride (PVDF, Beyotime Institute of Biotechnology, Shanghai, China) membranes. The PVDF membranes were incubated at 4
Luciferase reporter assay
The putative miRNA binding sites in the MAGI2-AS3 sequences were predicted by StarBase V2.0 (
Statistical analysis
Statistical analyses were performed using GraphPad software (GraphPad Prism version 5.0, San Diego, USA). Differences between groups were evaluated for statistical significance using
Results
lncRNA MAGI2-AS3 inhibits breast cancer cell migration and invasion
To confirm the migration and invasion effect of MAGI2-AS3 in breast cancer cells, we upregulated MAGI2-AS3 in MDA-MB-231 and MCF-7 cells using a lentivirus. The results showed that MAGI2-AS3 lentivirus (LV-MAGI2-AS3) transfection significantly increased MAGI2-AS3 expression in these cell lines (Fig. 1A). A wound healing assay revealed that MAGI2-AS3 overexpression significantly decreased the migratory capability of both MDA-MB-231 and MCF-7 cells (Fig. 1B). Transwell experiments revealed that MAGI2-AS3 overexpression apparently inhibited migration and invasion in both MDA-MB-231 and MCF-7 cells (Fig. 1C).
MAGI2-AS3 inhibited breast cancer cell migration and invasion. (A) MAGI2-AS3 levels were significantly higher in response to LV-MAGI2-AS3 infection than in response to LV-NC infection in MDA-MB-231 and MCF-7 cells. (B and C) lncRNA MAGI2-AS3 suppresses the migration and invasion of MDA-MB-231 and MCF-7 cells. The bar graphs display the mean 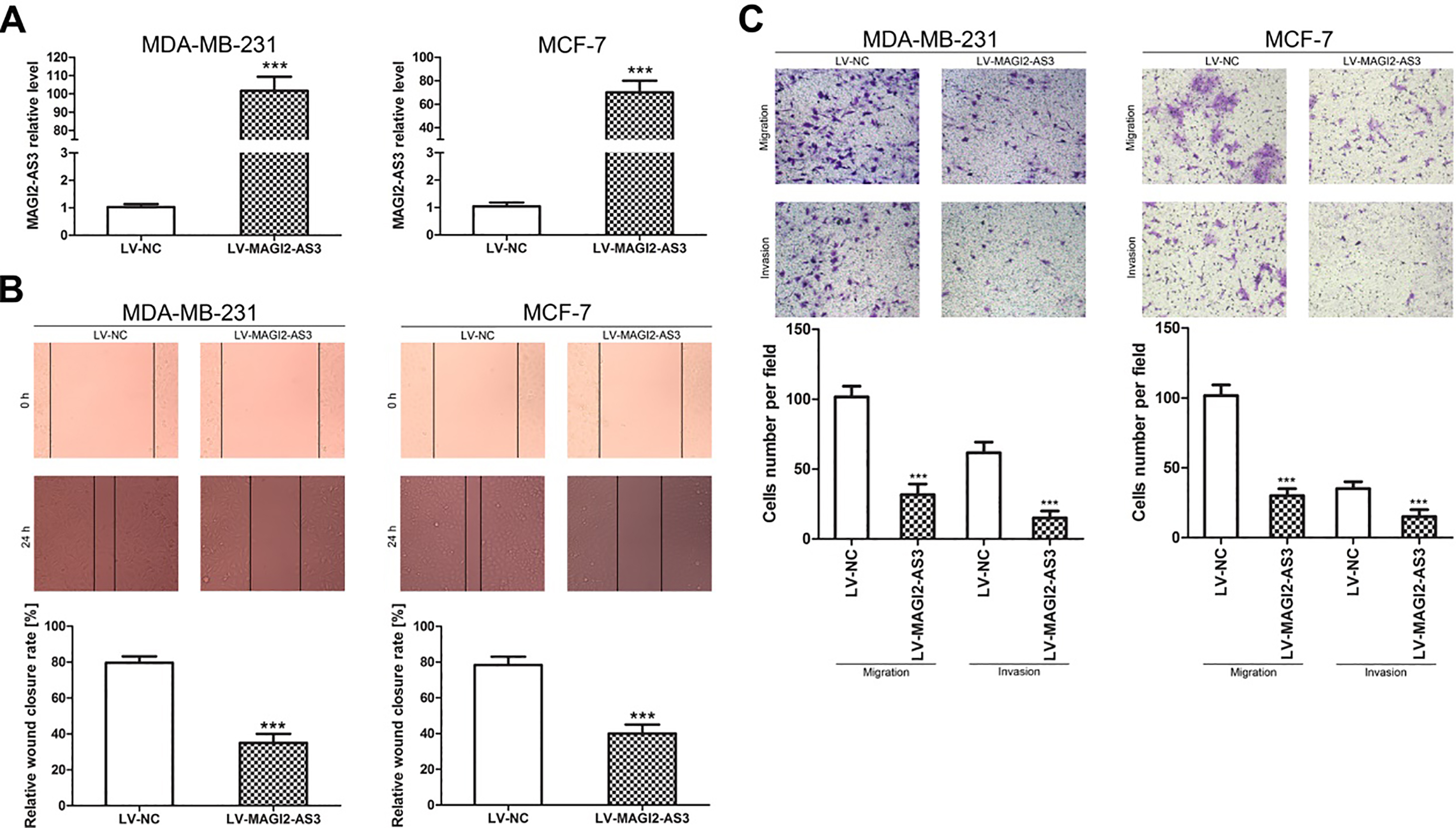
Because lncRNAs function as miRNA decoys, we predicted miRNA recognition sequences in MAGI2-AS3 using StarBase 2.0 software. We found a putative miR-374a-binding site in the MAGI2-AS3 transcript (Fig. 2A). First, we investigated the correlation between MAGI2-AS3 and miR-374a expression. As shown in Figure 2B, MAGI2-AS3 overexpression decreased miR-374a expression in MDA-MB-231 and MCF-7 cells. To further clarify the association between MAGI2-AS3 and miR-374a, luciferase activity was measured in MDA-MB-231 and MCF-7 cells co-transfected with WT-MAGI2-AS3 or MUT-MAGI2-AS3 reporter plasmid and miR-374a mimic or mimic NC. We found that miR-374a overexpression significantly decreased the luciferase activity of WT-MAGI2-AS3 but not that of MUT-MAGI2-AS3 (Fig. 2C). These results implied that miR-374a was a direct target of MAGI2-AS3 in breast cancer cells.
miR-374a is a direct target of MAGI2-AS3 in breast cancer cells. (A) miR-347a binding sites in MAGI2-AS3 were predicted by bioinformatics analysis. (B) Relative qRT-PCR analysis revealed that MAGI2-AS3 overexpression significantly decreased miR-374a levels in MDA-MB-231 and MCF-7 cells. (C) Luciferase reporter assays were performed using MDA-MB-231 and MCF-7 cells co-transfected with miR-374a mimic or mimic NC and WT-MAGI2-AS3 or MUT-MAGI2-AS3 reporter plasmids. The bar graphs display the mean 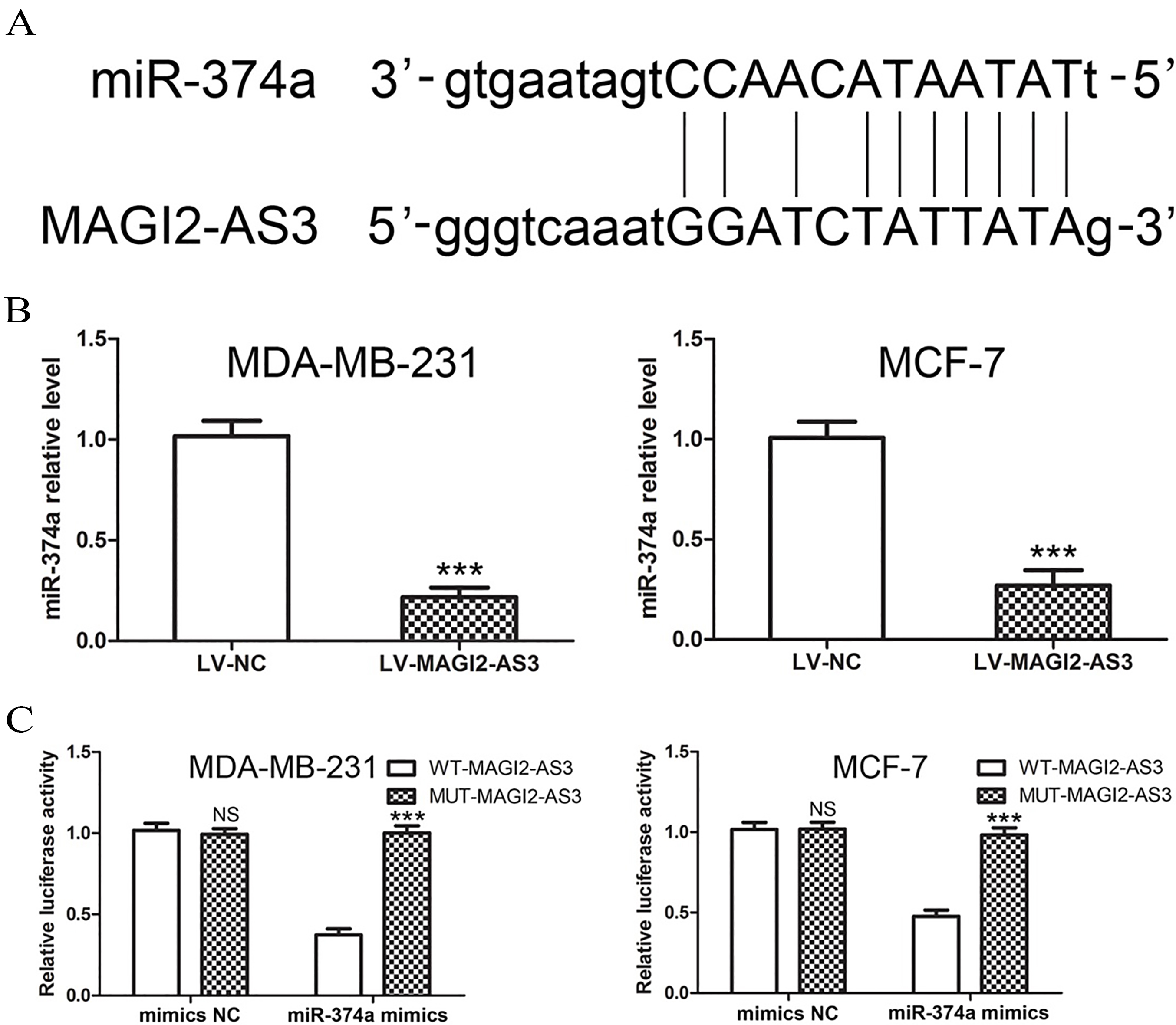
To confirm the function of miR-374a, we transfected miR-374a mimics or mimics NC sequences into breast cancer cells. Transfection efficiency was estimated by fluorescence microscopy 6 h after transfection, and miR-374a expression was determined by qPCR at 24 h. The results showed that the miR-374a mimics significantly increased miR-374a expression in these cells (Fig. 3A). The functional analyses showed significantly greater cell migration and invasion (Fig. 3B and C) in cells transfected with miRNA-374a mimics than in those transfected with mimics NC. To further explore the mechanism by which miR-374a promotes breast cancer cell migration and invasion, we performed bioinformatic analyses to identify the target genes of miR-374a using the miRanda (www.microrna.org) database. We found that PTEN is targeted by miR-374a as PTEN has potential complementary binding sites for miR-374a within its 3’-UTR (Fig. 3D). Several previous studies have suggested that PTEN is a tumor suppressor gene involved in cancer cell proliferation, migration, invasion and apoptosis, as well as angiogenesis, through several signaling pathways [12, 13]. We performed western blotting analysis to assess the effects of miR-374a on PTEN expression. PTEN protein expression was lower in MDA-MB-231 and MCF-7 cells after miR-374a mimic treatment than after mimics NC treatment (Fig. 3E). To confirm PTEN as a direct target of miR-374a, we engineered luciferase reporter constructs containing the WT or MUT 3’UTR of the PTEN gene. A luciferase reporter assay showed that miR-374a significantly decreased the luciferase activity of the PTEN 3’UTR but not that of the MUT in MDA-MB-231 and MCF-7 cells (Fig. 3F). Taken together, these data show that miR-374a promotes breast cancer cell migration and invasion by targeting PTEN.
miR-374a promotes breast cancer cell migration and invasion by regulating PTEN. (A) miR-374a levels were significantly higher in response to miR-374a mimic infection than in response to mimics NC infection in MDA-MB-231 and MCF-7 cells. (B and C) miR-374a suppresses migration and invasion in MDA-MB-231 and MCF-7 cells. (D) The seed sequence of miR-374a matches the 3’-UTR of PTEN. (E) PTEN protein levels were significantly decreased in MDA-MB-231 and MCF-7 cells transfected with miR-374a mimics according to western blotting assay. (F) Effect of miR-374a on PTEN expression as determined by a luciferase reporter assay. The bar graphs display the mean 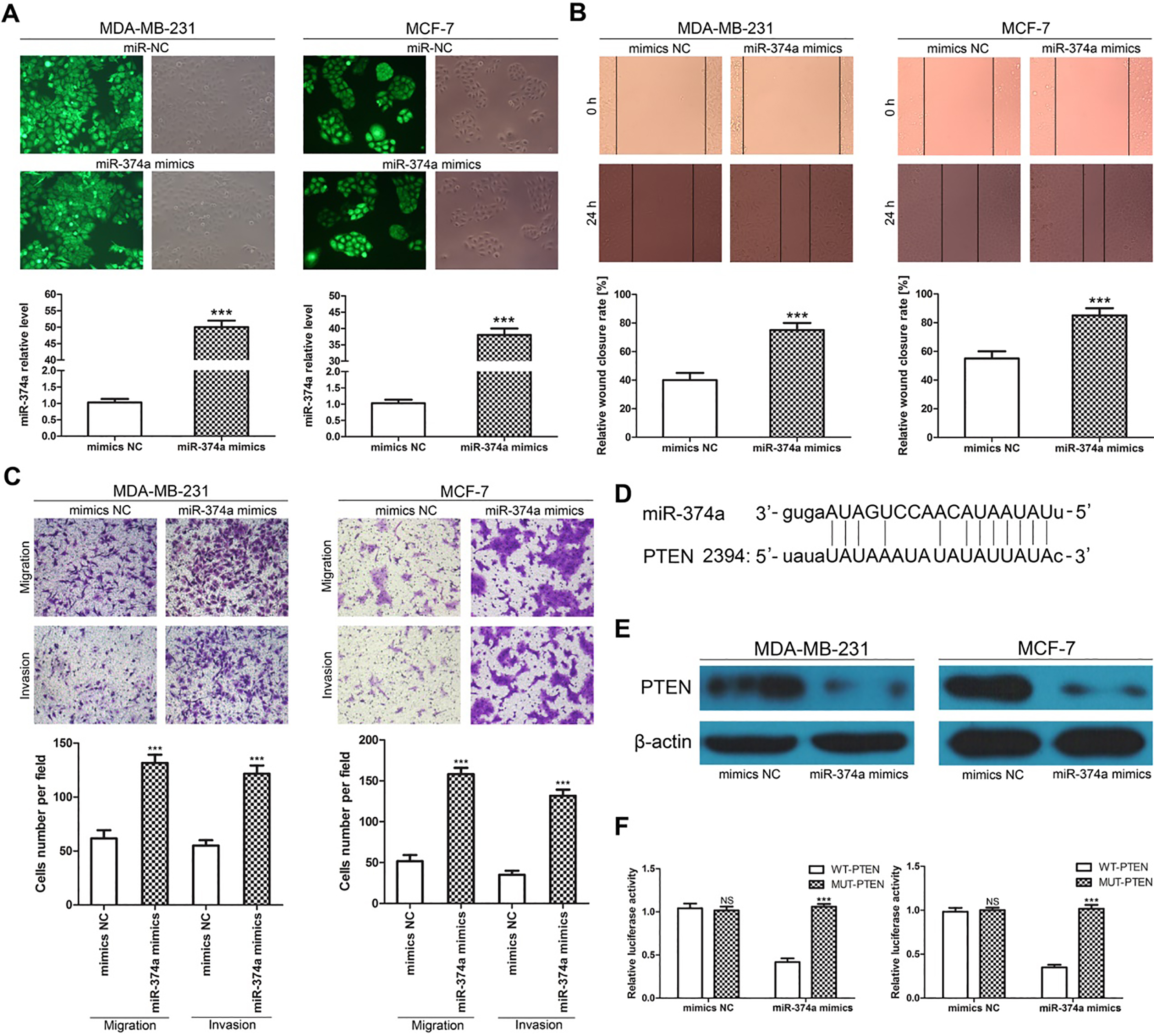
To test whether MAGI2-AS3 affected PTEN expression via sponging miR-374a in breast cancer cells, MDA-MB-231 or MCF-7 cells were transfected with LV-MAGI2-AS3, miR-374a mimics or cotransfected with LV-MAGI2-AS3 and miR-374a mimics, and PTEN expression was detected. As shown in Fig. 4A and B, transfection with LV-MAGI2-AS3 significantly increased PTEN expression, transfection with miR-374a mimics significantly decreased PTEN expression compared with transfection with LV-MAGI2-AS3, and cotransfection with LV-MAGI2-AS3 and miR-374a mimics increased PTEN expression compared with cotransfection with miR-374a mimics in both MDA-MB-231 and MCF-7 cells. In terms of cell migration and invasion, transwell assays showed that cotransfection with LV-MAGI2-AS3 and miR-374a mimics significantly decreased cell migration and invasion compared with transfection with miR-374a mimics. These results indicated that MAGI2-AS3 controls PTEN expression by regulating miR-374a in breast cancer cells.
MAGI2-AS3 suppresses breast cancer cell migration and invasion by upregulating PTEN. (A and B) MAGI2-AS3 controls PTEN expression by regulating miR-374a in MDA-MB-231 and MCF-7 cells as analyzed by western blotting. (C) lncRNA MAGI2-AS3 suppresses the migration and invasion of MDA-MB-231 and MCF-7 cells by regulating miR-374a. The bar graphs display the mean 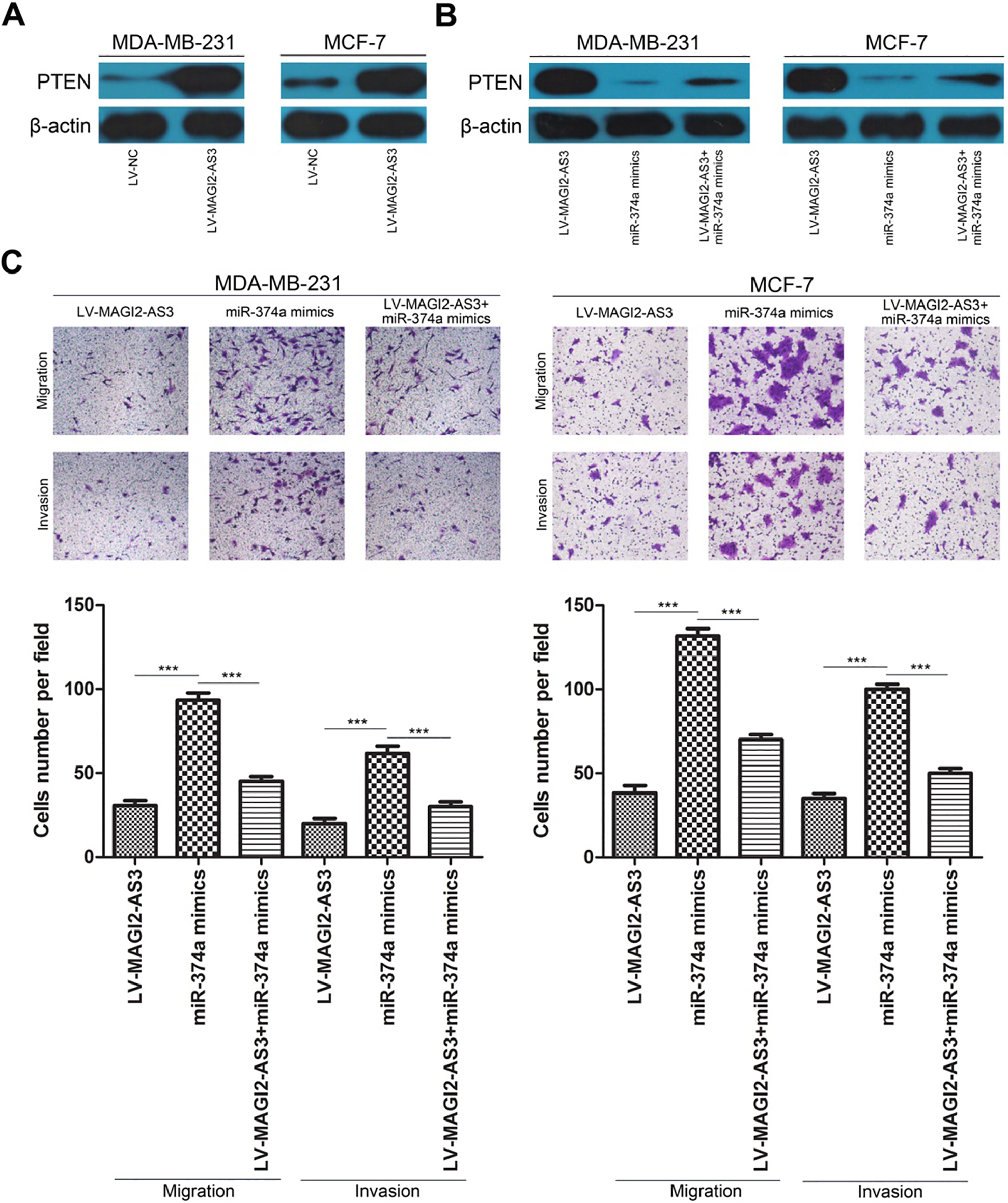
lncRNAs are a type of non-coding RNA that are 200 nt long and do not have protein-coding capacity [14]. They have been presented as new regulators of various biological processes in tumors because they serve as proto-oncogenes or tumor suppressor genes [15, 16]. Increasing evidence has demonstrated that lncRNA dysregulation is frequently present in breast cancer and has a great impact on the formation and progression of various tumors, including breast cancer [17]. For example, lncRNA MALAT1 was upregulated in breast cancer, and MALAT1 suppression inhibited cell proliferation, migration, and invasion and promoted apoptosis and cell cycle arrest in breast cancer [18, 19]. lncRNA Z38 is highly expressed, and Z38 silencing significantly inhibits cell proliferation and tumorigenesis in breast cancer [20]. In a previous study, we investigated the effect of lncRNA MAGI2-AS3 on breast cancer cell proliferation and explored the underlying mechanisms [21]. In the present study, we aimed to explore the role of lncRNA MAGI2-AS3 in breast cancer metastatic progression. To determine the role of MAGI2-AS3 in breast cancer, we transfected LV-MAGI2-AS3 into breast cancer cells to induce its overexpression. Our data showed that exogenous MAGI2-AS3 overexpression significantly inhibited breast cancer cell migration and invasion by wound healing and Transwell assays.
Previous evidence has shown that lncRNAs can serve as a competitive endogenous RNA (ceRNA) to sponge miRNAs [22]. To further explore the underlying molecular mechanism by which MAGI2-AS3 regulates breast cancer, we performed a bioinformatics prediction analysis and found that miR-374a was a potential candidate. In this study, our data showed that MAGI2-AS3 upregulation decreased miR-374a expression in both MDA-MB-231 and MCF-7 cells. Moreover, dual luciferase reporter assays further confirmed direct binding between miR-374a and MAGI2-AS3. Previous studies have found that miR-374a is upregulated in several types of tumors, suggesting that miR-374a may play pivotal roles in cancer tumorigenesis [23, 24, 25, 26]. Our data also suggest that miR-374a overexpression could promote breast cancer cell migration and invasion. These results implied that MAGI2-AS3 exerted tumor-suppressive effects via downregulating miR-374a in breast cancer cells.
The aforementioned results showed that MAGI2-AS3 elicited its anti-cancer effects by acting as a sponge for miR-374a in breast cancer cells. This effect might affect the ability of miR-374a to bind to its targets. A previous study demonstrated that miR-374a inhibited cell metastasis by targeting PTEN [27]. PTEN is well recognized as a tumor suppressor gene in human cancer; it is involved in regulating cancer cell biological behavior and plays an important role in the progression and development of various types of human cancer [28]. Here, we found that MAGI2-AS3 overexpression significantly decreased miR-374a expression and increased PTEN expression at the mRNA and protein levels, respectively, in breast cancer cells. Meanwhile, cotransfection with LV-MAGI2-AS3 and miR-374a mimics increased PTEN expression compared with transfection with miR-374a mimics in both MDA-MB-231 and MCF-7 cells. Furthermore, transwell assay showed that cotransfection with LV-MAGI2-AS3 and miR-374a mimics significantly decreased cell migration and invasion compared with transfection with miR-374a mimics. These results suggest that MAGI2-AS3 exerts its anti-oncogene function, at least in part, by regulating the miR-374a/PTEN axis.
In conclusion, our studies revealed that MAGI2-AS3 may serve as a therapeutic target for breast cancer metastatic progression and may function as a molecular sponge of miR-374a to inhibit breast cancer cell migration and invasion by regulating PTEN expression. These findings provide a novel mechanism for the occurrence and development of breast cancer.
Footnotes
Acknowledgments
This work was supported by grants from the National Natural Science Foundation of China (no. 8160 2330), the Project of Shandong Province Higher Educational Science and Technology Program (no. J18KA 299), Zibo key research and development plan (no. 2018kj010140) and the Natural Scientific Foundation of Shandong Province (no. ZR2015PH056). We also thank Professor Sun of Jiangsu University for providing all of the cell lines.
Conflict of interest
The authors declare that there are no conflicts of interest.
