Abstract
BACKGROUND:
Shisha smoking has been associated with multiple diseases including oral cancer. However, a mechanistic study to investigate alteration of secreted proteins in oral cells due to shisha smoking is lacking.
OBJECTIVES:
Elucidation of differentially secreted proteins by immortalized human normal oral keratinocytes (OKF6/TERT1) upon chronic exposure to shisha.
METHODS:
OKF6/TERT1 was chronically treated with 0.5% shisha extract for 8 months. Conditioned media from shisha treated (OKF6/TERT1-Shisha) and untreated (OKF6/TERT1-Parental) cells were subjected to TMT-based quantitative proteomic analysis. Bioinformatics analysis of differentially secreted proteins was carried out using SignalP, SecretomeP and TMHMM. Immunoblot validation of selected proteins was carried out to confirm the proteomics results.
RESULTS:
Proteomic analysis of OKF6/TERT1-Parental and OKF6/TERT1-Shisha secretome resulted in the identification of 1,598 proteins, of which 218 proteins were found to be differentially secreted (
CONCLUSION:
This study serves as a useful resource to understand the effect of chronic shisha smoking on the milieu of secreted proteins of oral cells. In vivo studies are warranted to supplement our in vitro data to elucidate the role of these proteins as early diagnostic biomarkers for oral carcinogenesis among shisha smokers.
Introduction
Tobacco smoking has been established as a major risk factor for oral cancer. In addition to cigarette smoking, tobacco usage in the form of shisha (waterpipe) smoking has gained popularity globally [1]. The use of shisha is prevalent in the Middle Eastern countries, with a gradual shift towards younger demographic due to increasing popularity and lack of stringent regulations [2, 3]. A multi-country survey of Arab nations recorded higher preference towards shisha smoking compared to cigarette smoking or smokeless tobacco among youth [4]. Studies have demonstrated that levels of polycyclic aromatic hydrocarbons and volatile aldehydes present in mainstream shisha smoke are similar or higher compared to cigarette smoke [5, 6]. Epidemiological studies have attributed shisha smoking to lung, esophageal and gastric cancers [7, 8, 9]. However, molecular alterations associated with shisha smoking has not been well characterized [10, 11].
Secretory proteins play an important role in cell-to-cell adhesion as well as cell-matrix interactions. Analysis of secreted proteins provides insights into the nature of autocrine and paracrine signaling within the microenvironment of tissues [12]. Changes in the abundance of secretory proteins in cancers have been previously reported [13] and studying them aids in the understanding of disease progression [13, 14]. Secreted proteins are known to be involved in metastasis, and this makes secretome from tumor cells a rich reservoir for identifying potential biomarkers [15, 16, 17]. Although proteins secreted by tumors ultimately reach body fluids such as saliva and blood, their presence can be masked by more abundant proteins, making it difficult to identify and study these proteins [18, 19]. Hence, cell line-based models, which are relatively low interference systems, are preferred to understand the cancer secretome in vitro. Secretome characterization using mass spectrometry has been used for potential biomarker discovery in various cancers such as breast, esophageal and ovarian cancers [20, 21, 22]. Candidates identified through such cell line-based discovery studies can be validated in body fluids, particularly in serum/plasma derived from cancer patients using non-invasive or minimally invasive techniques [23].
In this study, we investigated the effect of chronic shisha exposure on the secretome of normal human oral keratinocytes. We have employed mass spec- trometry-based quantitative proteomics approach to identify differentially secreted proteins in normal oral keratinocytes chronically treated with shisha extract (OKF6/TERT1-Shisha) compared to untreated oral keratinocytes (OKF6/TERT1-Parental). To the best of our knowledge, this is the first study of its kind investigating the molecular alterations of secreted proteins in oral keratinocytes due to chronic shisha exposure. This study can serve as a useful resource for identification of clinically relevant early diagnostic markers for oral cancer among shisha smokers.
Materials and methods
Preparation of shisha extract and cell culture
Shisha extract was prepared using a similar method described previously by Rohatgi et al. [24]. Briefly, 5 g of commercially available shisha was homogenized in 50 ml of phosphate buffered saline (PBS). The mixture was stirred overnight in a shaker incubator maintained at 37
Immortalized non-transformed normal human oral keratinocyte (OKF6/TERT1) cell line was cultured in keratinocyte serum-free media (KSFM) (Life Technologies, Grand Island, NY) supplemented with bovine pituitary extract (25 mg/ml), calcium chloride (0.4 mM), epidermal growth factor (0.2 ng/ml) and 1% penicillin/streptomycin, and maintained in a humidified CO
To study the effects of shisha on oral keratinocytes, 0.5% shisha extract was added to the growth media. Fresh media along with the shisha extract was replenished every 48 hours. Cells were maintained in the culture flask until they were confluent and then passaged into fresh culture flask, and shisha treatment was repeated. The process was continued for a period of 8 months. At the end of the treatment period, cells were allowed to attain 80% confluence under shisha exposure, following which the media was removed and further steps for secretome analysis was carried out. Cells which were treated with shisha extract were termed as ‘OKF6/TERT1-Shisha’. Simultaneously, control cells which were not treated with shisha were maintained alongside for the same duration (termed as OKF6/TERT1-Parental).
Collection and processing of secretome
Secretome collection and processing was carried out as described previously by our group [25]. OKF6/ TERT1-Parental and OKF6/TERT1-Shisha cells were grown to 80% confluence and washed thrice with PBS after removing the supplement-rich media. Cells were then grown in supplement-free media for 8 h. Subsequently, conditioned media was collected and centrifuged at 800 g for 10 min to remove any cell debris. The supernatant was filtered using a 0.22
Protein extraction, TMT labeling and bRPLC fractionation
Protein concentrations from OKF6/TERT1-Parental and OKF6/TERT1-Shisha secretome were determined using bicinchoninic acid (BCA) assay (Thermo Scientific, Bremen, Germany) [26] and equal amount of protein was collected from both conditions for further processing. Samples were reduced using dithiothreitol (DTT) at 60
The pellets from both the treated and untreated conditions were then reconstituted in 4 M urea. Protein samples were then digested using Lysyl Endopep- tidase
Peptide samples were labelled using Tandem Mass Tag (TMT) labels (ThermoScientific, Bremen, Germany) as per manufacturer’s instructions. Peptides derived from OKF6/TERT1-Parental and OKF6/TERT1-Shisha secretome were labelled with TMT tags 130C and 131, respectively. TMT labelled peptides were pooled and subjected to basic pH reverse phase chromatography (bRPLC), as previously described [27]. The 96 fractions obtained were concatenated into 6 fractions. The fractions were vacuum-dried, desalted using C
LC-MS
analysis
Peptide fractions were analysed on Orbitrap Fusion Tribrid mass spectrometer (Thermo Scientific, Bremen, Germany) interfaced with Easy nLC-1000 system (Thermo Scientific, Odense, Denmark). The peptide fractions were reconstituted in 0.1% formic acid (Solvent A) and loaded onto a trap column (75
Data acquisition on the Orbitrap Fusion was carried out using a data-dependent method with synchronous precursor selection MS3 scanning for TMT tags. The scan sequence was started with the acquisition of a full MS or MS1 one spectrum acquired in the Orbitrap (m/z range, 500–1600; resolution, 1.2
Data analysis
Identification and quantification of proteins was carried out using Proteome Discoverer (version 2.1) software suite (Thermo Scientific, Bremen, Germany). Raw files were searched against NCBI human RefSeq protein database (version 75) containing common contaminants, using Sequest HT search algorithm. In the search parameters, carbamidomethylation of cysteine, and TMT label at peptide N-termini and lysine were set as static modifications; while oxidation of methionine was set as a dynamic modification. Trypsin was chosen as the protease with a maximum allowed missed cleavage of 1. Precursor mass tolerance was set to 10 ppm and fragment mass tolerance was set to 0.1 Da. Decoy database search was carried out and peptide spectrum matches with a false discovery rate (FDR) cut-off of 1% was considered for peptide identification. Quantitative node in Proteome Discoverer was used to calculate protein ratios between OKF6/TERT1-Shisha and OKF6/TERT1-Parental using reporter ion intensities. Protein fold-change ratio was calculated as 131/130C (OKF6/TERT1-Shisha/OKF6/TERT1-Parental). Scatter plots were generated using log
Bioinformatics analysis
Proteins identified in conditioned media were assessed for their secretory potential using SignalP [28]. Proteins following non-classical secretion pathways were predicted using SecretomeP, and TMHMM was used to predict transmembrane domain [29, 30]. Proteome of conditioned media was also compared with experimentally annotated databases – ExoCarta [31], Vesiclepedia [32] and Human Protein Atlas (HPA) [33] to identify vesicular proteins based on experimental evidence. In addition, the data was compared with the updated salivary proteome [34] to identify differentially secreted proteins potentially detectable in saliva.
Partial list of differentially secreted proteins identified in OKF6/TERT1 cells chronically treated with shisha extract
Partial list of differentially secreted proteins identified in OKF6/TERT1 cells chronically treated with shisha extract
Quantitative proteomic analysis of OKF6/TERT1-Shisha secretome. (A) Workflow for TMT-based quantitative secretome analysis of OKF6/TERT1 cells chronically treated with 0.5% shisha extract. (B) Scatter plots representing the correlation of protein fold change (OKF6/TERT1-Shisha/OKF6/TERT1-Parental) across three replicates. Pearson correlation coefficient (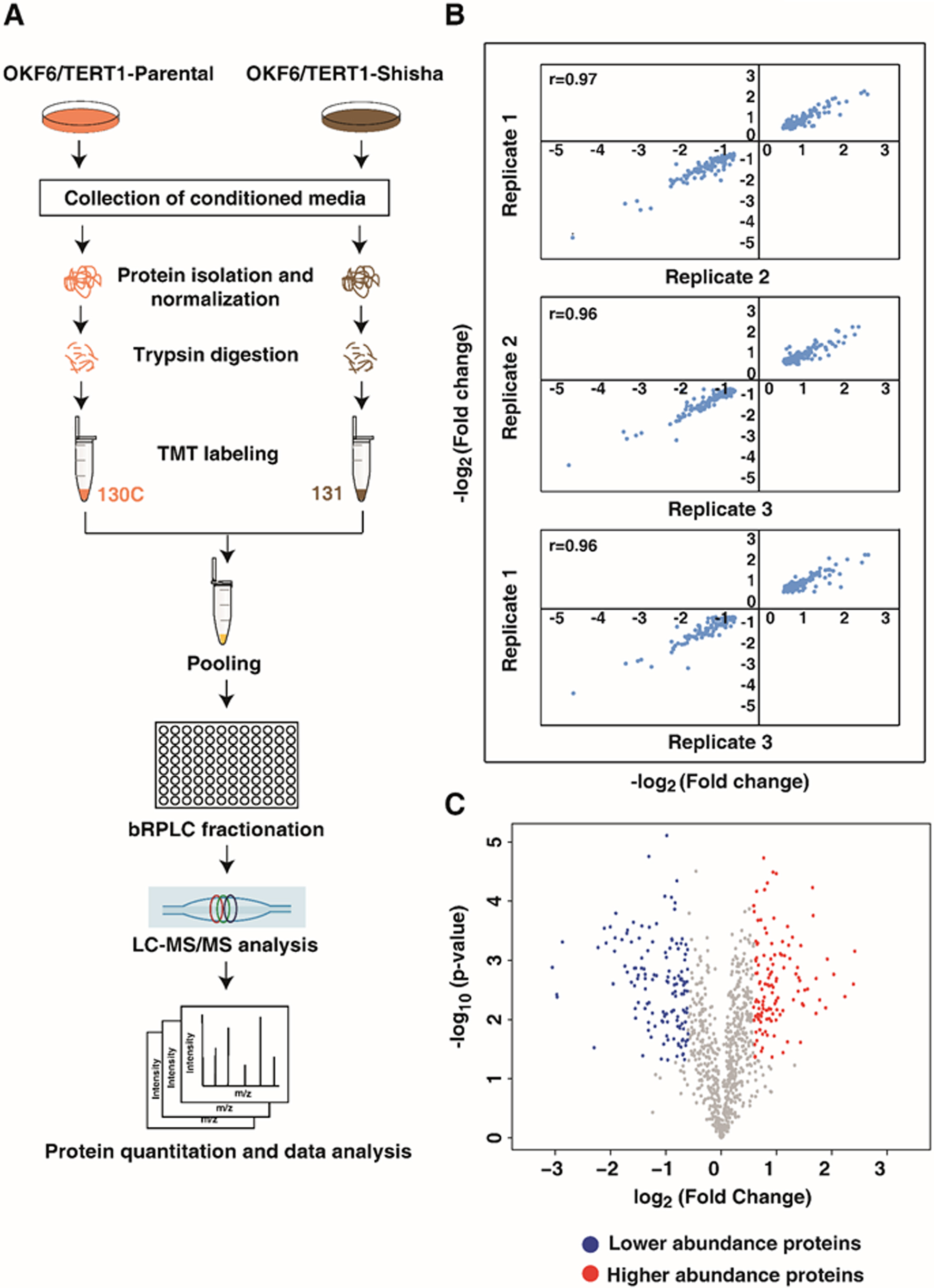
Representative MS/MS spectra of proteins found to be differentially abundant in OKF6/TERT1-Shisha secretome – (A) AKR1C2 (B) LGALS1 (C) HSPH1 (D) MMP9. Inset peaks represent tandem mass tag (TMT) label intensities representing fold change of the protein.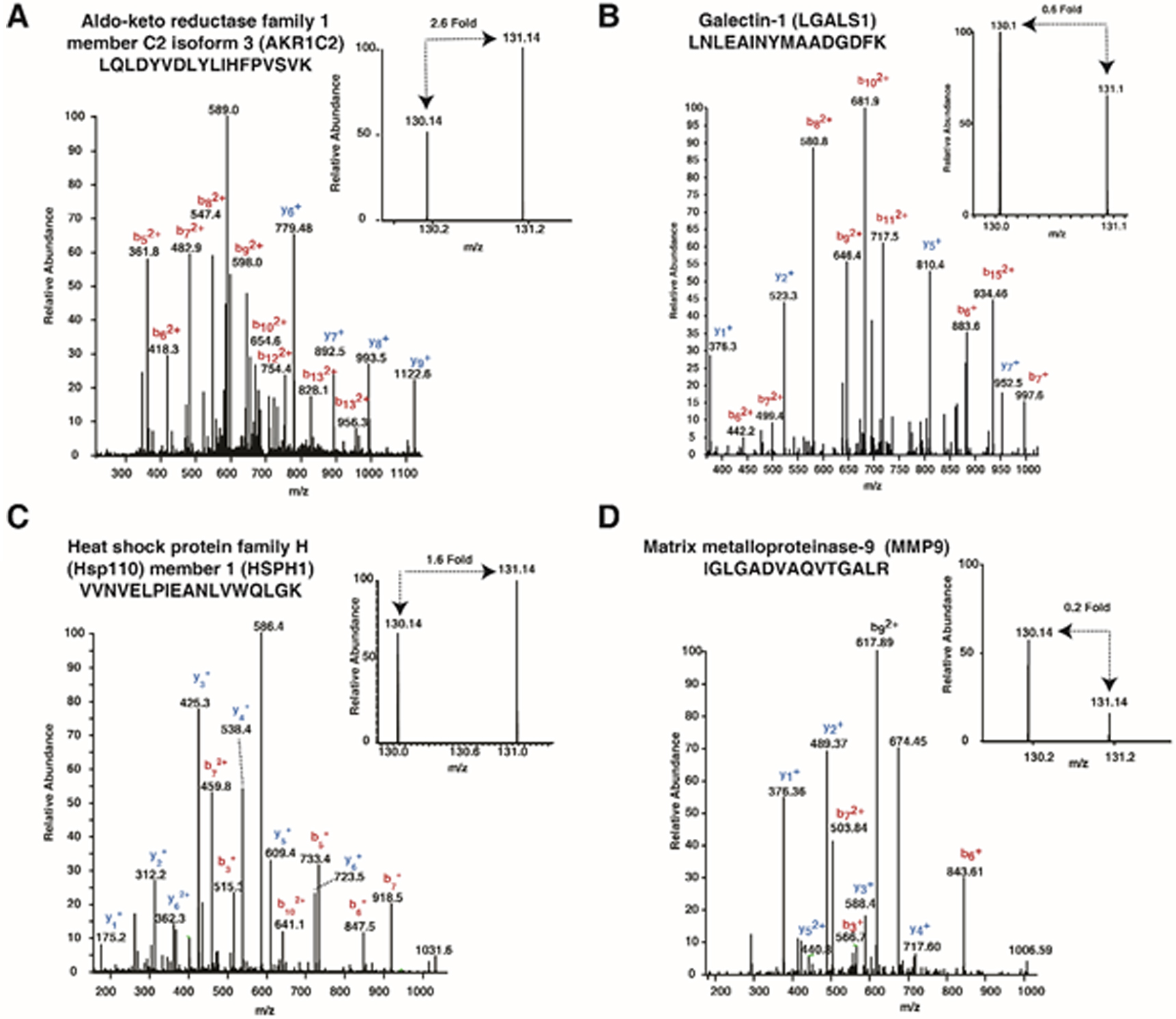
Secretory evidence for differentially abundant proteins in OKF6/TERT1-Shisha secretome. (A) Pie-Chart representation of the number of differentially secreted proteins identified in this study which have experimental and prediction-based evidence for secretory potential. (B) Venn diagram depicting the distribution of differentially secreted proteins identified in this study compared with experimentally validated databases. (C) Venn diagram depicting the differentially secreted proteins identified this study with secretory potential analyzed using prediction-based tools – SignalP and SecretomeP (D) Venn diagram representing the number of proteins differentially abundant in this study reported to be present in normal human salivary proteome (Sivadasan et al., 2015).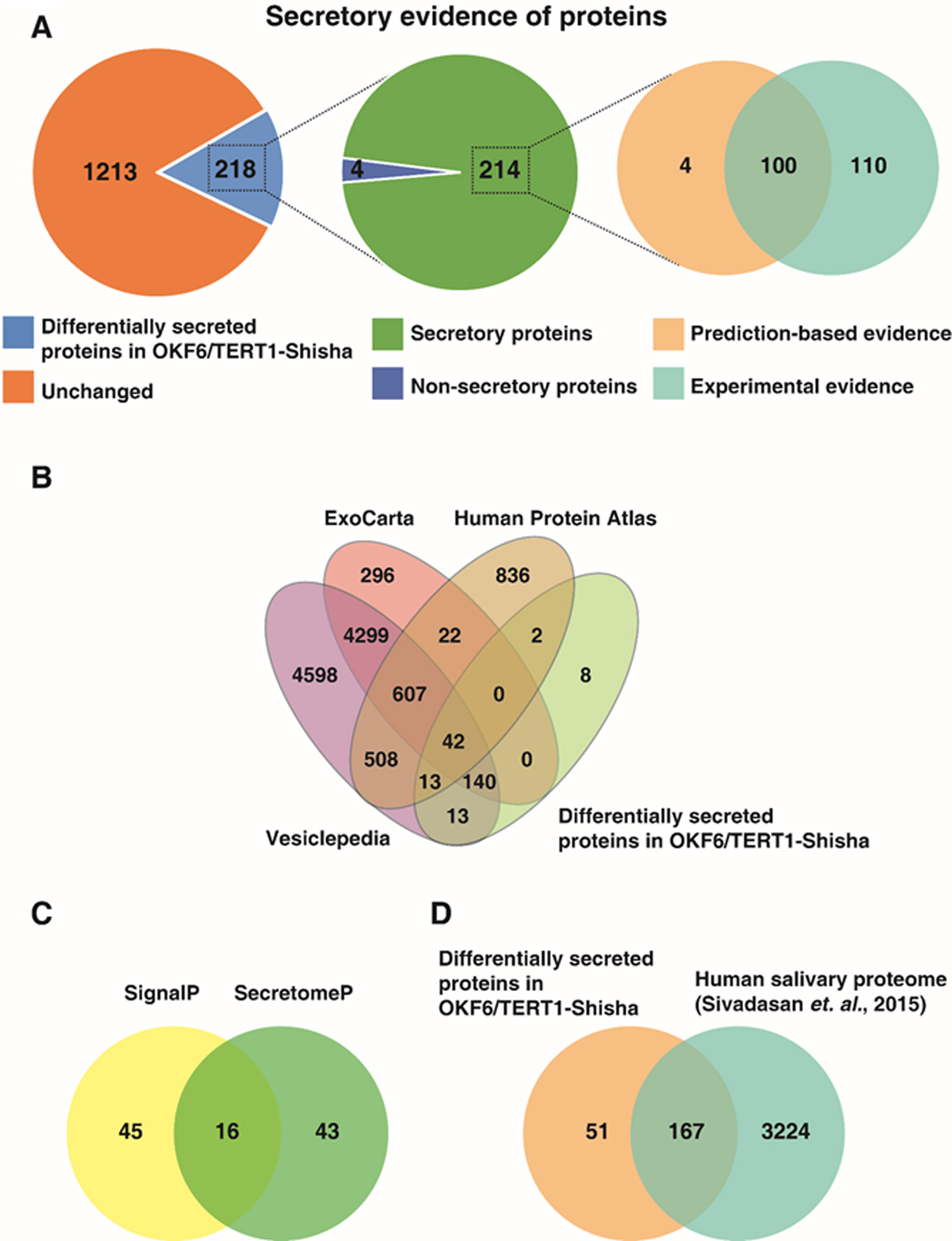
(A) Venn diagram representing differentially secreted proteins identified in this study as compared to proteins reported in published HNSCC/OSCC secretome studies. (B) Western blot validation for selected proteins differentially secreted in OKF6/TERT1-Shisha secretome.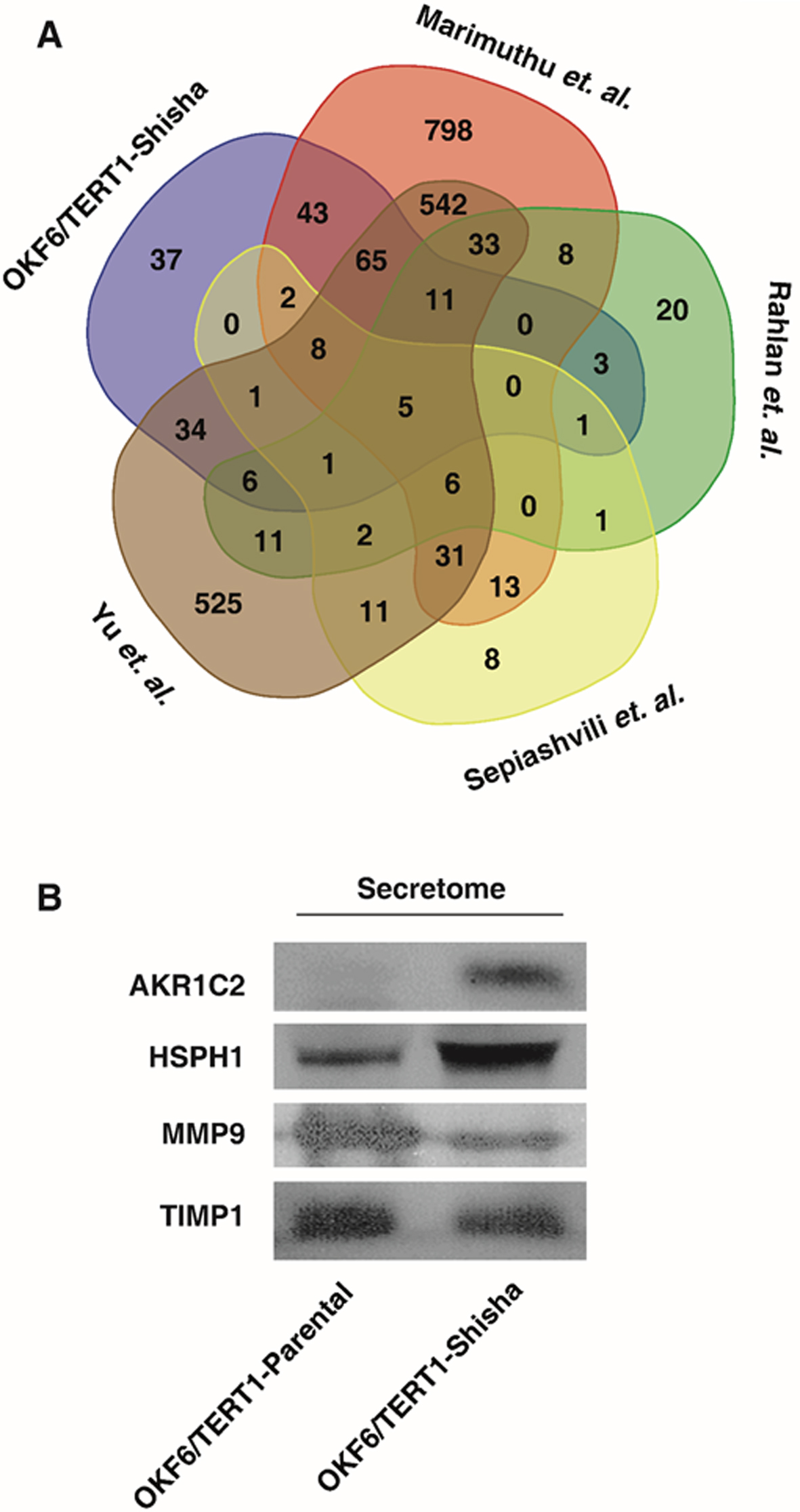
30
Statistical analysis
Statistical significance was calculated in Perseus software suit [35] using student’s t-test.
Data availability
Proteomics data has been submitted to ProteomeXchange Consortium (
Results
Quantitative proteomic analysis of OKF6/TERT1-Shisha secretome
Mass spectrometry-based quantitative proteomic analysis of secretome from OKF6/TERT1-Parental and OKF6/TERT1-Shisha cells resulted in the identification of 1,598 proteins. The workflow employed in this study is depicted in Fig. 1A. High correlation (r
Bioinformatics analysis of dysregulated proteins in OKF6/TERT1-Shisha secretome
Differentially secreted proteins identified in our data were analyzed for secretory potential using prediction-based bioinformatics tools (SignalP and SecretomeP) and by confirmation with databases of experimentally validated secreted proteins (Vesiclepedia, ExoCarta and HPA). Overall, we found secretory evidence for 214 out of 218 differentially secreted proteins in OKF6/TERT1-Shisha (Fig. 3A). Comparison of differentially secreted proteins with the secreted protein-database in Human Protein Atlas showed an overlap of 57 proteins. In addition, we also compared the differentially secreted proteins in our data with ExoCarta and Vesiclepedia databases to identify exosomal proteins which are released into extracellular space in the form of membranous vesicles. In total, we found 208 proteins with experimental evidence for vesicle-mediated exosomal secretion (Fig. 3B, Supplementary Table 2).
We further analyzed the differentially secreted proteins for the presence of N-terminal signal peptides using SignalP software. Proteins which contain N-terminal hydrophobic peptides are released from cells through the classical secretory pathway facilitated by endoplasmic reticulum [42]. Our analysis predicted 61 differentially secreted proteins contain N-terminal signal peptide. In addition, based on SecretomeP analysis, we identified 43 proteins differentially secreted in OKF6/TERT1-Shisha secretome which do not contain a signal peptide but are predicted to be secreted by non-classical pathways (Fig. 3C, Supplementary Table 2). Further, we analyzed proteins for the presence of a transmembrane domain using TMHMM analysis. We identified 25 differentially secreted proteins in OKF6/TERT1-Shisha secretome predicted to contain a transmembrane helical domain(Supplementary Table 2).
We evaluated differentially secreted proteins in shisha exposed cells (compared to parental cells) that have been previously detected in normal human saliva. We compared differentially secreted proteins with the salivary proteome study which cataloged the normal human salivary proteins based on mass spectrometry evidence [34]. We found that 167 proteins differentially secreted in our data overlapped with proteins identified in human saliva, of which 89 proteins were secreted in higher abundance in OKF6/TERT1-Shisha secretome (Fig. 3D, Supplementary Table 2). Taken together, our analyses provides strong evidence of secretory potential for the differentially secreted proteins in our data. In addition, comparison of higher abundant proteins identified in our data with the salivary proteome serves the objective of this study to identify early diagnostic biomarkers for oral cancer in shisha smokers.
Identification of potential cancer biomarkers using the OKF6/TERT1-Shisha secretome data
Based on proteomic evidence, we established that normal oral keratinocytes OKF6/TERT1 chronically treated with shisha extract results in altered secretion of proteins. We compared the differentially secreted proteins identified in our data with four HNSCC studies published previously. Marimuthu et al. carried out iTRAQ-based quantitative proteomic analysis of the secretome of 6 HNSCC cell lines – JHU-O11, JHU-O22, JHU-O28, JHU-O29, FaDu and CAL 27, and compared with the secretome of OKF6/TERT1 [25]. Sepiashvili et al. studied the secretome of HNSCC cell lines FaDu, UTSCC8 and UTSCC42a using mass spectrometry-based proteomic approach and gene expression microarrays [43]. Yu et al. carried out secretome analysis of OSCC cell lines OECM1 and SCC4, using mass spectrometry-based approach and further validated their findings with tissue transcriptome analysis [44]. Rahlan et al. employed proteomics approach to identify proteins in the secretome of four HNSCC cell lines-SCC4, HSC2, SCC38, and AMOSIII [45]. Upon comparison, we found that 134, 131, 21 and 18 proteins reported to be differentially secreted (
Differentially secreted proteins in our data and those reported previously in cancer cell lines indicate that chronic exposure of normal oral keratinocytes to shisha may lead to cellular transformation. Identification of such proteins will facilitate identification of early oral cancer diagnostic biomarkers among shisha users.
Immunoblot validation of differentially secreted proteins identified in OKF6/TERT1-Shisha secretome
To confirm the results of proteomic analysis, we carried out western blot validation of differentially secreted proteins identified in OKF6/TERT1-Shisha. These proteins were also reported to be differentially secreted in studies pertaining to HNSCC secretome as discussed above. In agreement with our mass spectrometry data, we observed significantly increased abundance of AKR1C2 and HSPH1 and decreased abundance of MMP9 compared to OKF6/TERT1-Parental secretome (Fig. 4B).
Discussion
Shisha smoking has been a recreational practice in the Middle Eastern and South East Asian countries since many centuries. However, emergence of shisha smoking among Western nations has been observed recently [46, 47]. Despite being categorized as a health hazard, the pathobiology of shisha smoking-related diseases remains poorly understood. Analysis of secreted proteins in disease conditions has the potential to reveal biomarkers that could be monitored in body fluids. Particularly in the case of oral cancer and head and neck cancers, where tobacco smoking remains a predominant risk factor, identification of potential prognostic biomarkers in saliva and serum have proven to be clinically relevant [48, 49]. In this study, we investigated the effects of chronic shisha exposure on secreted proteins in oral keratinocytes. We chronically treated normal immortalized non-transformed human oral keratinocytes (OKF6/TERT1) with shisha extract for a period of 8 months (OKF6/TERT1-Shisha), followed by proteomic analysis of the secretome which was compared with that of untreated cells (OKF6/TERT1-Parental).
Mass spectrometry-based quantitative proteomic analysis of OKF6/TERT1-Shisha secretome resulted in the identification of 218 differentially secreted proteins (
We compared out data with previous studies pertaining to HNSCC secretome and observed an overlap of several proteins differentially secreted by OKF6/TERT 1-Shisha cells. We observed an increased abundance of pyruvate kinase M1/2 (PKM) in our data, which is known to facilitate adhesion and migration of colon cancer cells by regulating STAT-3 associated signaling [60]. Similarly, our data also shows increased abundance of two eukaryotic elongation factors, EEF1G and EEF2. EEF1G is known to interact with E-cadherin to promote cell adhesion and has been demonstrated to be highly abundant both at mRNA and protein levels in colorectal carcinoma [61, 62]. Secretome of OKF6/TERT1-Shisha cells was found to have higher abundance of proteins reported in cell proliferation including transforming protein RhoA (RHOA) and prostaglandin E synthase 3 (PTGES3) amongst others. Overexpression of RHOA has been reported to promote proliferation and migration of cervical cancer cells [63]. Similarly, studies have shown increased expression of PTGES3 in the oral mucosa of smokers and hypoxia-induced esophageal squamous cell carcinoma (ESCC) [64, 65]. Comparison with published secretome studies revealed that PKM1/2, EEF1G, EEF2, RHOA and PTGES3 are differentially secreted in HNSCC cells and have also been identified in human saliva [34, 35].
Similarly, we detected higher abundance of chaperone proteins such as endoplasmin (HSP90B1), heat shock protein family H (Hsp110) member 1 (HSPH1) which have been reported in several cancers previously [66]. In addition, we also detected decreased abundance of ECM proteins such as matrix metallopeptidase 9 (MMP9), secreted protein acidic and rich in cysteine (SPARC) and cysteine rich angiogenic inducer 61 (CYR61) among others in OKF6/TERT1-Shisha secretome. Dysregulation of certain ECM proteins has been known to promote detachment of cancer cells from their primary tumor and to facilitate metastasis [67, 68]. Expression of SPARC and CYR61 inversely correlates with cell invasion and metastasis in ovarian, prostate and gastric cancer, respectively [69, 70, 71, 72]. These findings indicate that chronic shisha exposure alters the expression of secreted proteins involved in cellular processes linked to tumorigenesis.
Our data also suggests alteration in metabolic pathways in OKF6/TERT1 cells chronically treated with shisha extract. Aberrant metabolism has been widely established as one of the hallmarks of cancer and is often associated with increased energy requirements and requirement of metabolites for anabolic pathways [73]. Although conventionally believed to be restricted to the cytoplasm, studies have demonstrated that glycolytic enzymes have non-canonical extracellular functions as well in cancers, such as cytokine signaling and anti-apoptotic response [74]. Moreover, elevated levels of metabolic enzymes in the serum have been correlated with tumorigenesis and drug resistance [75, 76]. Increased secretion of glycolytic proteins GAPDH, PFKP and GPI has been previously reported in HNSCC secretome [12, 25, 29, 31].
In summary, this study highlights the detrimental effects of shisha smoking on oral keratinocytes in vitro and provides list of candidate proteins that potentially play a role in oral carcinogenesis. We hypothesize that differentially secreted proteins identified in this study may serve as early diagnostic markers among shisha smokers. Further studies using salivary samples from OSCC patients with shisha smoking habits are warranted to establish these proteins as clinically relevant biomarkers.
Conclusions
The current study was carried out to investigate the adverse effects of chronic exposure of shisha in oral keratinocytes by studying their secretory protein profile. Using proteomics-based approach, we identified several differentially secreted proteins in the secretome of shisha-exposed cells. Our data indicates that chronic exposure to shisha results in aberrant oxidative stress response and altered glycolytic pathway in oral keratinocytes. Comparison of our data with previously published OSCC and HNSCC secretome studies showed an overlap of several differentially secreted proteins, many of which were also detectable in human saliva. This study demonstrates molecular aberrations in oral keratinocytes upon chronic shisha exposure and highlights the potential of several identified secretory proteins as biomarkers in oncogenic risk assessment amongst shisha smokers.
Footnotes
Acknowledgments
We thank the Department of Biotechnology (DBT), Government of India for research support to the Institute of Bioinformatics (IOB), Bangalore. NB, KP and JA are recipients of the Senior Research Fellowship from the Council of Scientific and Industrial Research (CSIR), New Delhi, India.
Conflict of interest
The authors declare that there are no conflict of interests.
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-182099.
