Abstract
BACKGROUND:
Tobacco exposure (through smoking or chewing) is one of the predominant risk factors associated with the development of oral squamous cell carcinoma (OSCC). Despite the growing number of patients diagnosed with OSCC, there are few circulating biomarkers for identifying individuals at a higher risk of developing the disease. Successful identification of candidate molecular markers for risk assessment could aid in the early detection of oral lesions and potentially be used for community screening of high-risk populations.
OBJECTIVE:
Identification of differentially expressed proteins in the serum of oral cancer patients which can serve as biomarkers for the diagnosis of the onset of oral cancer among tobacco users.
METHODS:
We employed a tandem mass tag (TMT)-based quantitative proteomics approach to study alterations in the serum proteomes of OSCC patients based on their tobacco exposure habits (chewing and smoking) compared to healthy individuals with no history of using any form of tobacco or any symptoms of the disease.
RESULTS:
Mass spectrometry-based analysis resulted in the identification of distinct signatures in the serum of OSCC patients who either chewed or smoked tobacco. Pathway analysis revealed opposing effects of dysregulated proteins enriched in the complement-coagulation signaling cascades with a high expression of the Serpin family of proteins observed in OSCC patients who chewed tobacco compared to healthy individuals whereas these proteins showed decreased levels in OSCC patients who smoked. ELISA-based validation further confirmed our findings revealing higher expression of SERPINA6 and SERPINF1 across serum of OSCC patients who chewed tobacco compared to healthy individuals.
CONCLUSIONS
: This study serves as a benchmark for the identification of serum-based protein markers that may aid in the identification of high-risk patients who either chew tobacco or smoke tobacco.
Abbreviations
Introduction
Tobacco is predominantly consumed in its smoking form (cigarettes, smoking of ‘bidis,’ or cigars) worldwide and is one of the leading causes of oral cancer [1, 2]. Studies have reported the molecular alterations due to cigarette smoke and its constituents in oral and esophageal cancers [3, 4, 5]. However, in certain South-East Asian countries, another form of tobacco consumption that is widely popular is the chewing tobacco [6]. Numerous studies have reported the use of multiple forms of smokeless tobacco, including chewing tobacco, as a significant risk factor and/or cause for oral squamous cell carcinoma (OSCC) cases [7, 8, 9]. Studies also indicate the increased risk of oral cancer due to chewing tobacco compared to those with no habits of consuming tobacco [10, 11]. We and others have shown that both chewing and smoking tobacco’s adverse effects lead to alteration of multiple metabolic functions/pathways [12, 13]. Our group has previously reported signature molecular alterations in normal oral keratinocytes chronically subjected to cigarette smoke and chewing tobacco [14]. Thus, despite the mode of tobacco intake, its use in any form, smoking or smokeless, is implicated in the development of multiple cancers, including OSCC [15].
Though numerous epidemiological studies have demonstrated the correlation of smoking and chewing tobacco with oral cancers [7, 16], very few have focused on the difference in the effects of cigarette smoke and chewing tobacco at the molecular level. As the constituents between the two forms of tobacco vary significantly [17, 18, 19], it is evident that the mode of action among these two tobacco products also varies considerably. Thus, it is important to understand and differentiate the molecular changes induced by smoking and chewing tobacco, even when it comes to the early detection or screening of the population for risk assessment. Multiple groups have used body fluids such as serum and saliva to identify clinical biomarkers for OSCC [20, 21, 22]. Several serum biomarkers such as adiponectin, cyclin D1, C-Reactive Protein (CRP), growth-differentiation factor (GDF 15), cytokines such as interleukin 1 beta (IL-1
In this study, we employed a quantitative proteomics approach to identify differentially expressed proteins in the serum from OSCC patients based on their tobacco using habits (chewing and smoking) compared to healthy volunteers with no habits and no symptoms of disease or any predisposed medical condition.
Materials and methods
Study subject, sample collection, and storage
Patient and control blood samples were collected after approval from the Institutional and Ethical Review Board at Dr. R. Ahmed Dental College and Hospital, Kolkata, West Bengal, and Yenepoya Medical College, Mangalore, India (YEC-1/2017/166). Informed consent was obtained from all patients and volunteers before the blood samples were collected. Only those patients diagnosed with OSCC with tobacco chewing and smoking habits were included in the study. Patients with mixed habits, any other systemic illness, or who had undergone prior surgery were excluded from the study. Blood samples were collected from healthy volunteers with no tobacco using habits, no symptoms of the disease, and no pre-existing/existing health conditions/systemic illness.
For serum isolation, blood samples were collected in BD Vacutainer Plus plastic serum tubes, kept at room temperature to coagulate for 20–30 min, followed by centrifugation at 3,000 g for 20 min at 4
Sample preparation for MS analysis
LC-MS/MS approach was employed using isotopomer labels, “tandem mass tags” (TMTs), to determine the relative quantification of proteins in the serum [25]. The serum samples (
In-solution digestion and TMT labeling
Post-depletion, an equal amount of protein from the three cohorts were processed for TMT labeling. Briefly, 150
Vacuum-dried peptides were labeled with TMT (Thermo Fisher Scientific) as per manufacturer’s protocol. Two sets of 10-plex TMT experiments were carried out. The control serum sample set used for both the experiments were identical (processed and labeled together). Following TMT labeling, samples were subjected to a StageTip-based strong cation exchange (SCX)-based fractionation adapted from Kulak et al. [27]. Briefly, StageTips are prepared by stacking 5 layers of SCX sorbent in a 200
Mass spectrometry analysis
The Orbitrap Fusion Tribrid mass spectrometer (Thermo Fisher Scientific, Bremen, Germany) interfaced with EASY-nLC 1200 system (Thermo Fisher Scientific) was used for the analysis of TMT-labelled samples. The MS parameters used for analysis were followed as described previously [28]. Briefly, peptides from each fraction were resuspended in 0.1% formic acid (Solvent A) and loaded onto C18 trap column (75 um
Bioinformatics and statistical analysis
Proteome Discoverer 2.2 (Thermo Scientific, Bremen, Germany) was used to carry out protein identification and quantitation. All raw files were searched against Human RefSeq 92 protein database supplemented with common contaminants using SequestHT and Mascot (version 2.5.1, Matrix Science, London, UK) search algorithms. The search parameters included trypsin as the proteolytic enzyme with a maximum allowed missed cleavages of two; oxidation of methionine and acetylation of protein N-terminus was set as dynamic modifications, while static modifications included cysteine carbamidomethylation and TMT modification at N-terminus of the peptide and lysine. Precursor mass tolerance was set to 10 ppm, and fragment mass tolerance was set to 0.05 Da; all the PSMs were identified with 1% FDR. The differential expression ratios between OSCC patients with chewing and smoking habits and control samples were calculated. Proteins with differential expression ratios
A network analysis exploring relationships between SERPINs F1 and A6 and the Complement and Coagulation cascade pathway was carried out using STRINGdb in Cytoscape 3.8.0. The gene set for the pathway was obtained from the Molecular Signatures Database (MSigDB,
Enzyme-linked immunosorbent assay (ELISA)
ELISA-based validation was carried out using serum samples from a total of 48 individuals (24 healthy individuals
Data availability
The raw data files and the MSF files were submitted to the PRIDE partner repository [30] with dataset identifier PXD019014.
Proteomics data overlap between the two study groups. A schematic representing the results of the proteomics analysis of serum samples from OSCC patients who chewed or smoked tobacco.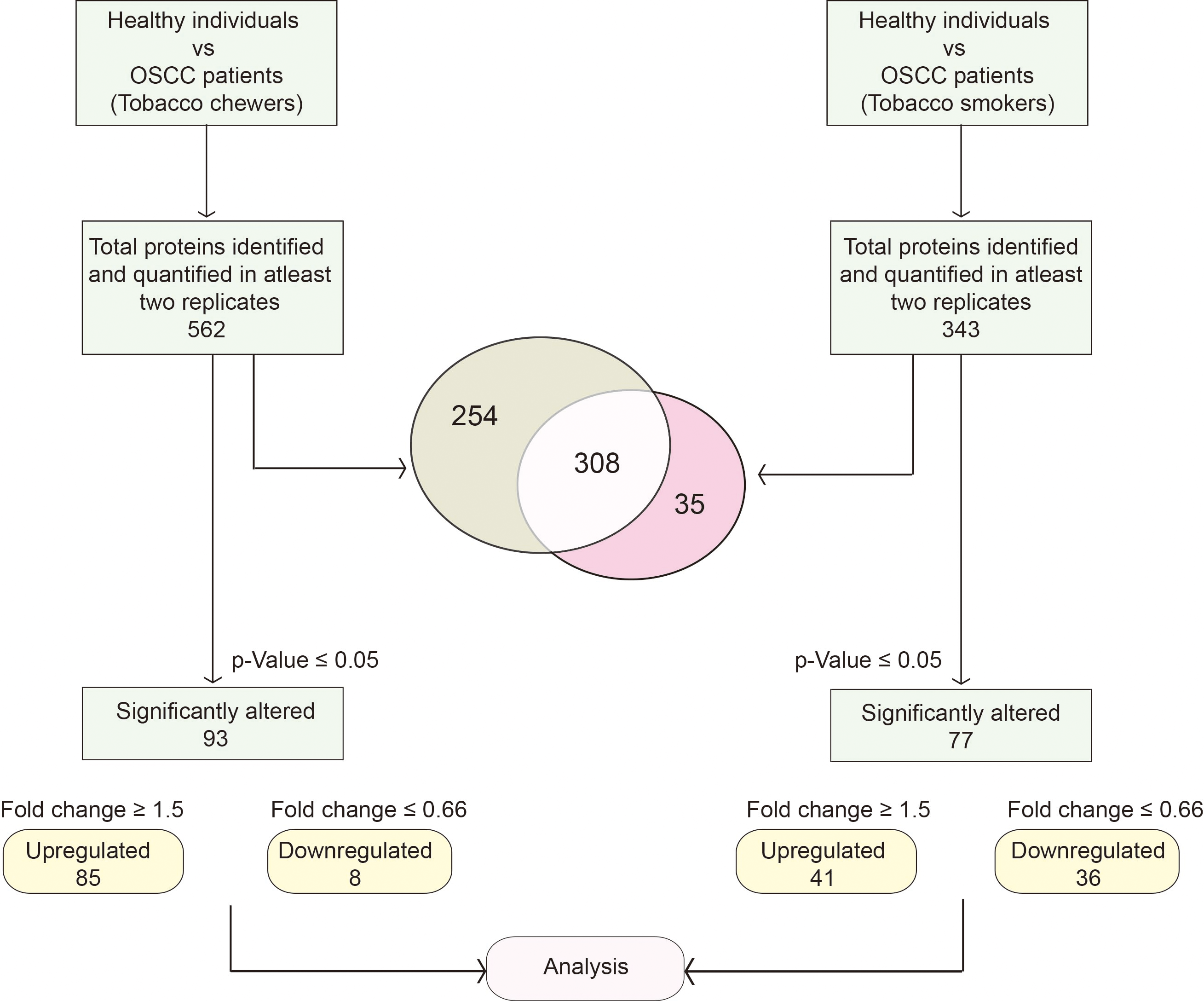
Comparative proteomic analysis of serum from OSCC patients
A total of 15 serum samples, five each from healthy individuals and 10 OSCC patients with tobacco exposure (chewing
A list of top six differentially expressed proteins based on tobacco using habits of OSCC patients
A list of top six differentially expressed proteins based on tobacco using habits of OSCC patients
Differential expression of serum proteins in OSCC patients. Volcano plots of the significantly differentially expressed proteins in the serum of (A) OSCC patients who chewed tobacco and healthy individuals; (B) OSCC patients who smoked and healthy individuals. The red and green dots represent significantly dysregulated proteins (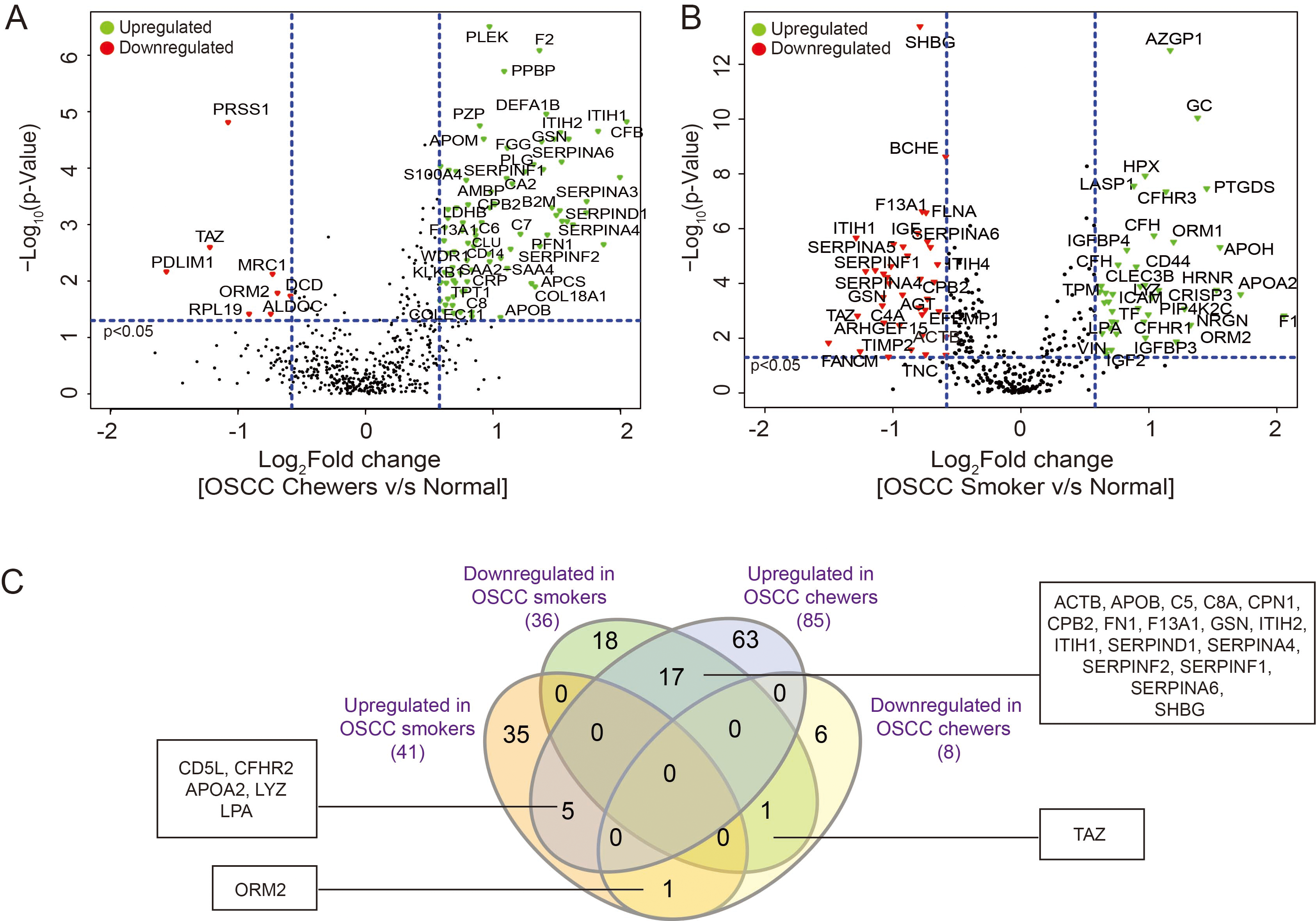
TMT-based quantitative proteomic analysis of the OSCC serum samples categorised based on tobacco usage resulted in the identification of 760 proteins, of which 562 were quantified across at least two replicates in the tobacco-chewing set. Similarly, in the smoking set, 442 proteins were identified, of which 343 proteins were quantified across at least two replicates. A complete list of the proteins and peptides identified in the study are provided in Supplementary Tables 1–4.
The results from these two sets were compared and illustrated using a Venn diagram (Venny 2.1) to identify the number of common and differentially expressed proteins. Following the schema, as shown in Fig. 1, a comparison of identified proteins across the two groups of tobacco users, revealed 254 proteins identified exclusively in OSCC patients who chewed tobacco, whereas 35 were exclusive to OSCC patients who smoked. A total of 308 proteins were shared between the sets (Fig. 1, Supplementary Table 5). Of the 562 proteins quantified in OSCC patients who chewed tobacco, 93 were significantly dysregulated (85 upregulated and 8 downregulated; 1.5-fold cut-off,
Role of the Complement and coagulation cascades pathway in OSCC. The complement and coagulation signaling cascade pathways were found to be enriched based on proteins dysregulated in OSCC patients who smoked and chewed as compared to healthy individuals. The proteins highlighted in grey were not identified in our study.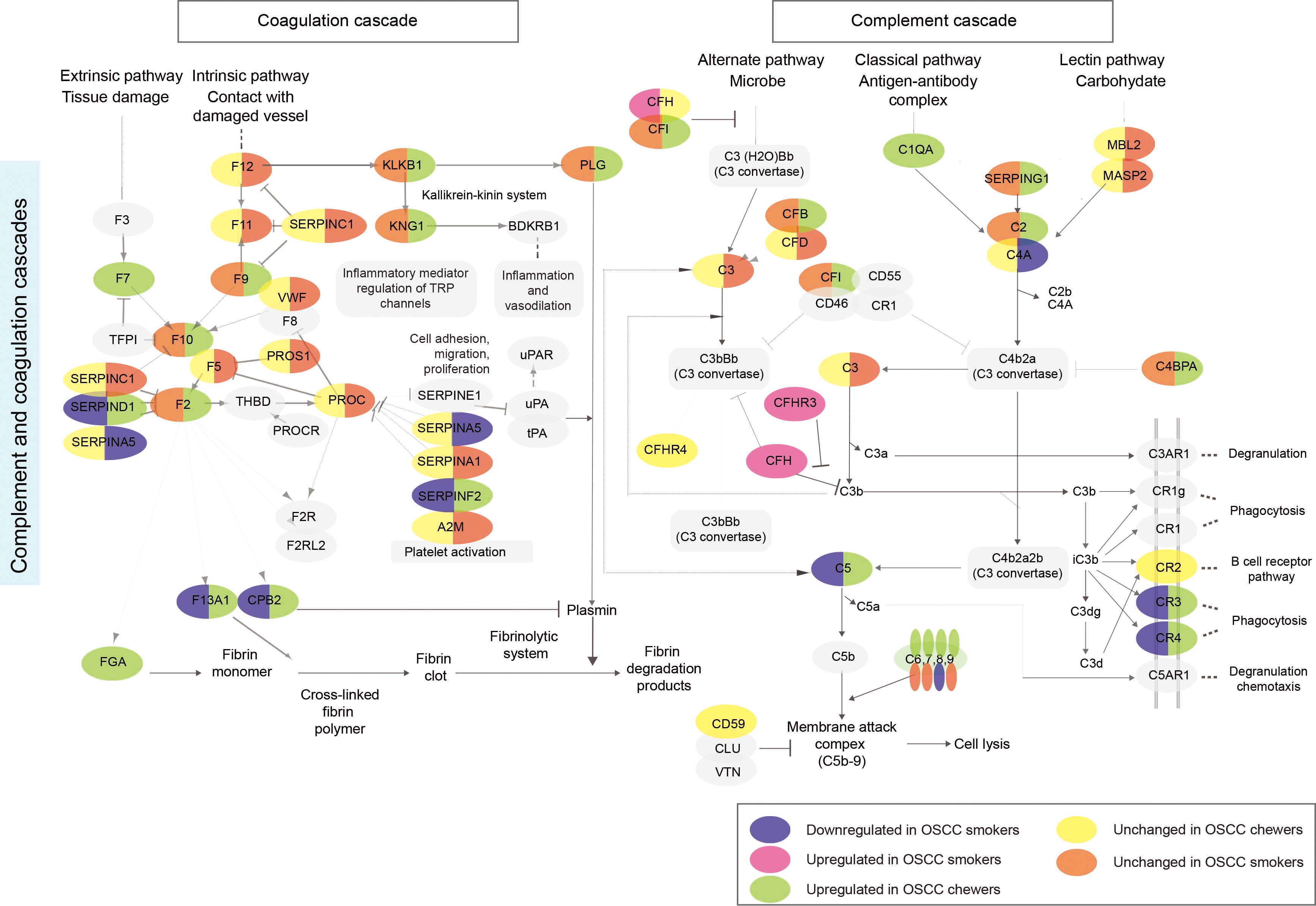
To obtain more insights and to interpret the molecular differences in the serum of OSCC patients as a result of chewing and smoking habits, we studied the respective signaling cascades using DAVID 6.8 [31]. Analysis of the dysregulated proteins from the serum of OSCC patients who chewed or smoked reflected enrichment of 5 pathways, listed in Supplementary Table 7. Interestingly, our analyses demonstrated that different modes of tobacco use exert the opposite effect on the complement and coagulation signaling cascades in OSCC patients (Fig. 3).
Our data showed contrasting expression trends of SERPIN family members in OSCC patients based on the form of tobacco used, indicating a relationship between dysregulation of SERPIN family proteins with tobacco-induced oncogenicity (Fig. 3). While increased expression levels of SERPINA4, SERPINA6, SERPIND1, SERPINF1, and SERPINF2 were observed in the serum of OSCC patients who chewed tobacco, these proteins showed decreased levels in OSCC patients who smoked. Other members, including SERPINA3 and SERPING1, were exclusively increased in OSCC patients who chewed tobacco while SERPINA5 was exclusively decreased in OSCC patients who smoked.
Differential expression of SERPIN family members and their opposing trends of expression between tobacco chewing and smoking OSCC patients. Representative MS/MS spectra and its corresponding scatter plot representation for identified peptides of (A) SERPINA6, and (B) SERPINF1 in OSCC patients who chewed tobacco as compared to healthy individuals; and (C) SERPINA6, and (D) SERPINF1 in OSCC patients who smoked as compared to healthy individuals, respectively. The data presented as vertical scatter plots consisted of patient (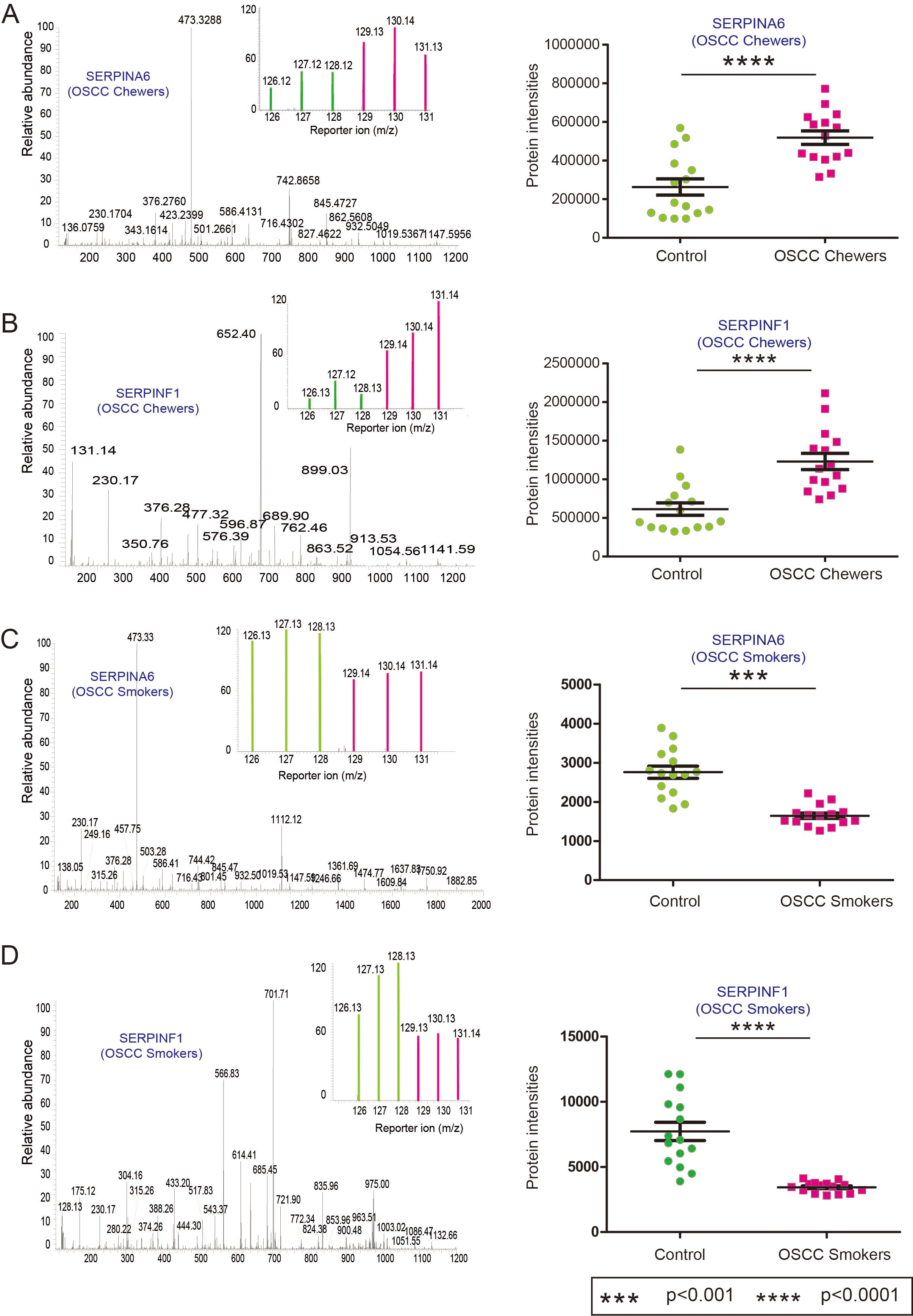
Validation of SERPINA6 and SERPINF1 by ELISA. ELISA-based validation: Expression of (A) SERPINA6 and (B) SERPINF1 in OSCC patients who chewed tobacco compared to healthy controls; Receiver operating characteristic (ROC) curves for (C) SERPINA6 and (D) SERPINF1, respectively. SERPINA6: Serpin family A member 6 or Corticosteroid-Binding Globulin; SERPINF1: Serpin family F member 1. (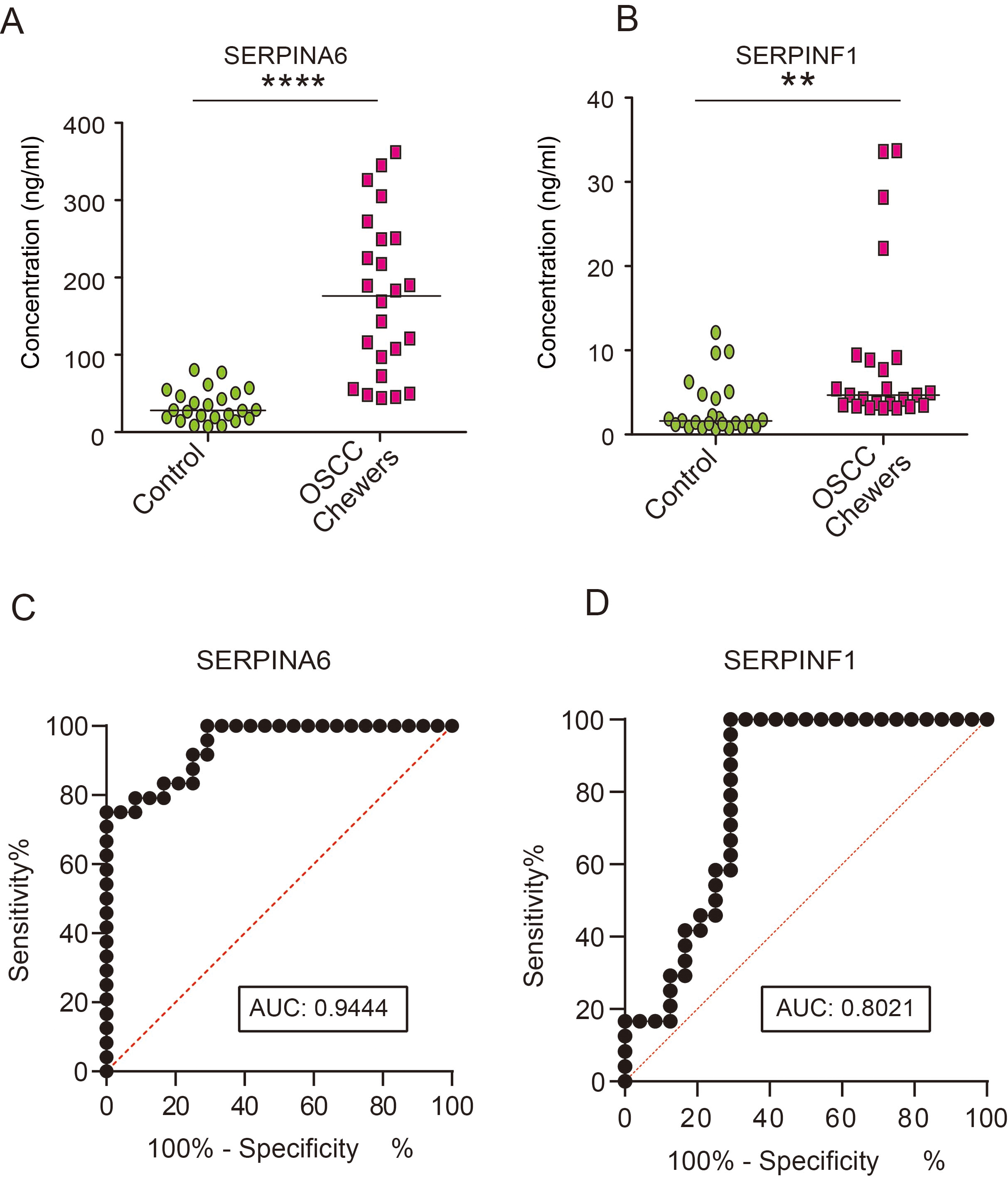
The previous section showed the serpin family to be enriched and showed the opposite expression trend in mass spectrometry data (Fig. 4, Supplementary Fig. 3). We found
We validated the expression of SERPINA6 and SERPINF1 by complementary ELISA-based approach using serum from 24 healthy controls and 24 OSCC patients who chewed tobacco (Supplementary Table 8). The levels of both the serpins were significantly higher in serum of OSCC patients who chewed tobacco compared to healthy controls (Fig. 5A and B) in agreement with our mass spectrometry findings.
A ROC curve analysis was carried out where we found the area under the ROC curve for SERPINA6 to be 0.9444 (95% CI: 0.8873–1.00223,
Discussion
Our mass spectrometry data has generated information for candidate markers and pathways dysregulated in OSCC patients with tobacco chewing and smoking habits. The pathobiology of OSCC resulting from tobacco use, either in chewing or smoking form, differs significantly owing to the difference in composition [17, 18, 19]. We observed dysregulation of proteins associated with plasminogen signaling cascade in the serum of patients who chewed tobacco. Aberrant expression of proteins participating in such pathways includes plasminogen (PLG), fibrinogens (FGA, FGB, FGG), actin (ACTB) contributing to malignant cell cycle progression in various cancers [32, 33]. Whereas in patients who smoked, majority of the proteins dysregulated are known to be involved in the process of platelet degranulation. These proteins have previously been reported to be dysregulated in lung cancer and reported to be involved in smoking-induced epithelial disorders [34, 35].
Comparison of the significantly upregulated proteins between the two groups led to the identification of 5 common proteins which were significantly upregulated in OSCC patients, irrespective of the form of tobacco used. These included CD5 molecule like (CD5L); complement factor H related 2 (CFHR2); lipoprotein (LPA); lysozyme (LYZ); and apolipoprotein A2 (APOA2). Studies have demonstrated that altered expression of LYZ results in tumor growth and metastasis and is associated with poor prognosis in cancer patients [36, 37]. APOA2 is an essential component of high-density lipoprotein and its differential expression has been reported in various types of cancers [38]. Its increased expression has been observed in the serum of patients with prostate cancer [39] and hepatocellular carcinoma [40]; however, its decreased level was reported in pancreatic cancer [41]. A recent study by Chu et al. reported increased levels of APOA2 in the saliva of OSCC patients compared to healthy individuals [42]. Interestingly, we identified an opposite trend in serum levels of ORM2 in OSCC patients, which is a secretory plasma protein and known to be associated with inflammation and multiple cancers [43, 44, 45]. Of the 17 proteins which showed the opposite trend between OSCC patients who chewed tobacco or smoked (i.e upregulated in patients who chewed tobacco and downregulated who smoked), significant enrichment of the members of the serpin family of proteins amongst others were observed. These proteins are mainly associated with metabolic processes and have catalytic activity, and have the potential to serve as distinct signatures to distinguish between the two cohorts based on habits.
Our data revealed opposite effects on the complement and coagulation signaling cascades between patients who chewed tobacco and patients who smoked tobacco. This signaling cascade is one of the significant proteolytic pathways, consisting of serine proteinases with trypsin-like activity and their activators and inhibitors, that protect against infections [46]. The complement pathway constitutes an integral component of humoral immunity, and a coagulation system is critical in hemostasis, and their roles have been implicated in multiple diseases, including cancer [47]. Our data indicate the presence of several hub proteins regulating this pathway, such as components of fibrinogen (FGA, FGB, FGG), plasminogen (PLG), complement factors (C2, C5, C7, C9, C8A, C8G, CPB), coagulation factors (F2, F7, F13A1) along with members of serpins family (SERPIND1, SERPING1) in the serum of OSCC patients who chewed tobacco. Our analysis further indicates a high fibrinogen level in OSCC patients who chewed tobacco, promoting platelet activation (FGA, FGB, FGG), which is in accordance with our pathway analysis. Higher plasma fibrinogen is known to be associated with multiple malignancies such as colon [48], pancreatic [49], nasopharyngeal [50], and oral cancer [51]. Complement factors and serpin family members were found to be upregulated in the OSCC chewer cohort. Both serpin family G member 1 (SERPING1) and serpin family D member 1 (SERPIND1) are known to play a crucial role in the regulation of the complement cascade and studies have revealed elevated levels in the saliva of OSCC patients [52].
In the case of OSCC patients who smoked, we observed decreased expression of complement component 5 (C5), which is a part of the complement cascade known to trigger acute inflammatory responses. Recent studies have reported downstream targets of C5a to induce chronic inflammation during tissue damage, thus promoting tumor growth by recruiting myeloid-derived suppressor cells [53, 54]. Our data also showed decreased expression of several proteins downstream of this pathway, such as C8A, CPB2, and F13A1, in the serum of OSCC patients who smoked. F13A, a key regulator of the coagulation cascade and plays an important role in promoting tumorigenesis and is reported to be downregulated in the serum of colorectal cancer patients [55]. In contrast, these were found to be increased in the serum of OSCC patients who chewed tobacco.
OSCC progression is a multi-step process marked by uncontrolled cell growth, invasion, and migration [56]. Such cellular processes are controlled by proteolytic enzymes such as serine and cysteine proteases, and serine proteases are controlled by members of the SERPIN (serine protease inhibitor) family. Our findings suggest that serine protease inhibitors identified in the study can activate the complement pathway in OSCC patients. Although the role of serpins in cancer is not well established, and the underlying mechanism is not well understood, our data indicates serpins play an essential role in the pathobiology of oral cancer. Our data indicated upregulation of SERPINA6 and SERPINF1 in patients who chewed tobacco whereas these proteins were downregulated in patients who smoked. ELISA-based validation further confirmed overexpression of SERPINA6 and SERPINF1 in the serum of OSCC patients who chewed tobacco. SERPINA6 is a 52 kDa secreted
These results suggest the possible role of SERPINA6 and SERPINF1 as potential biomarkers for OSCC patients with tobacco chewing habits. Further studies are warranted to define the functional role of these proteins in OSCC progression.
Conclusions
In our study, we investigated the underlying molecular alterations in OSCC based on the mode of tobacco intake. The use of tobacco, in both smoke and smokeless form, has been associated with an increased incidence of inflammation and activate complement and coagulation cascade [60, 61, 62]. We observed the dysregulation of coagulation and complement-related proteins in OSCC patients, suggesting its role in the pathogenic process of oncogenicity. However, this signaling cascade showed opposing effects owing to the different modes of tobacco intake. Our findings suggest that an imbalance of this signaling cascade contributes to the molecular basis for tumorigenic phenotype owing to tobacco use in either form. We demonstrate the promising role of SERPINA6 and SERPINF1 as potential early detection biomarkers for OSCC patients who chewed tobacco that should be tested in larger cohorts. However, we were unable to validate the same in the serum of OSCC patients who smoked due to the limited number of samples in the smoking cohort. Also, due to lack of samples, we were unable to study the expression of these proteins in subjects with a chewing or smoking tobacco habit but no oral lesion or symptoms of the disease or subjects with pre-cancerous lesions. We also did not have enough samples to explore if there was an association between the detected proteins in serum and the amount of tobacco exposure of individual patients. The study did not include patients with mixed habits (those who chewed and smoked tobacco), nevertheless, it might be interesting to extend this study by including such a cohort of patients. This work highlights the potential of serum proteins, which can serve as biomarkers to identify high-risk patients who either chew or smoke tobacco products. Further studies with larger cohorts and associated control groups are required in a clinical setting before these molecules can be taken forward as a potential risk assessment panel in OSCC patients who are tobacco users.
Author contributions
Conception: Aditi Chatterjee, T.S. Keshava Prasad, Harsha Gowda, David Sidransky
Interpretation or analysis of data: Varshasnata Mohanty, Yashwanth Subbannayya, Riaz Abdulla, Jay Gopal Ray, Shankargouda Patil, Mandakulutur S. Ganesh, Arnab Pal
Preparation of the manuscript: Varshasnata Mohanty, Yashwanth Subbannayya, Aditi Chatterjee
Revision for important intellectual content: Varshasnata Mohanty, Yashwanth Subbannayya, Aditi Chatterjee, Harsha Gowda, David Sidransky
Supervision: Aditi Chatterjee, T.S. Keshava Prasad, Harsha Gowda, David Sidransky
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-203077.
sj-pdf-1-cbm-10.3233_CBM-203077.pdf - Supplemental material
Supplemental material, sj-pdf-1-cbm-10.3233_CBM-203077.pdf
sj-docx-1-cbm-10.3233_CBM-203077.docx - Supplemental material
Supplemental material, sj-docx-1-cbm-10.3233_CBM-203077.docx
Footnotes
Acknowledgments
We thank Karnataka Biotechnology and Information Technology Services (KBITS), the Government of Karnataka, for the support to the Center for Systems Biology and Molecular Medicine at Yenepoya (Deemed to be University) under the Biotechnology Skill Enhancement Programme in Multiomics Technology (BiSEP GO ITD 02 MDA 2017). We thank Yenepoya (Deemed to be University) for access to instrumentation. The first author is a recipient of the Women Scientist-A award from the Department of Science and Technology (DST), Government of India. This work was supported by the Department of Science and Technology (DST), Government of India (SR/WOS-A/LS-32/2018). We thank Dr. Sneha M. Pinto, Centre of Molecular Inflammation Research (CEMIR), Norwegian University of Science and Technology, Norway for critical reading of the manuscript, inputs to data analysis and editing.
Conflict of interest
The authors declare that they have no conflict of interest.
