Abstract
BACKGROUND:
Long non-coding RNAs (lncRNAs) are emerging as candidate biomarkers of cancer, having regulatory functions in both oncogenic and tumor-suppressive pathways. Concerning pancreatic cancer (PC), deregulation of lncRNAs involved in tumor initiation, invasion, and metastasis seem to play a key role. However, data is scarce about regulatory mechanism of lncRNA expression.
OBJECTIVE:
The aim of our study was to investigate the contribution of two lncRNAs polymorphisms (rs1561927 and rs4759313 of PVT1 and HOTAIR, respectively) in PC susceptibility.
METHODS:
A case-control study was conducted analysing rs1561927 and rs4759313 polymorphisms using DNA collected in a population-based case-control study of pancreatic cancer (111 pancreatic ductal adenocarcinoma cases (PDAC), 56 pancreatic neuroendocrine tumor (PNET), and 125 healthy controls).
RESULTS:
Regarding the PVT1 rs1561927 polymorphism the G allele was significantly overrepresented in both PDAC and PNET patients compared to the controls, while the presence of the HOTAIR rs4759314 G allele was found to be overrepresented in the PNET patients only compared to the controls. The PVT1 rs1561927 AG/GG genotypes were associated with poor overall survival in PDAC patients.
CONCLUSIONS:
Our results suggested that polymorphisms of these two lncRNA polymorphisms implicated in pancreatic carcinogenesis. Further large-scale and functional studies are needed to confirm our results.
Keywords
Introduction
Pancreatic cancer (PC) is one of the most fatal types of cancer, resulting in approximately 331,000 deaths per year, according to the estimates of GlOBOCAN [1]. The incidence of PC differs across countries; environmental and genetic factors, ageing and lifestyle seem to be responsible for these differences globally [2]. Among putative risk factors, tobacco smoking, obesity, diabetes mellitus, chronic pancreatitis and excessive use of alcohol are the most well established [3]. Although several pancreatic malignancies may appear, the highest prevalence is found on pancreatic ductal adenocarcinoma (PDAC) and pancreatic neuroendocrine tumour (PNET) which account for 90% and 5% of all pancreatic cancer types, respectively [4].
PDAC is a malignancy formed in the exocrine pancreas [5]. Most patients are diagnosed when they have already reached stage IV due to its asymptomatic course [6] and it has a very poor prognosis. Five-year survival rates fall dramatically as the disease proceeds to its later stages [7]. PNET can be characterised as either functioning or non-functioning. Differential diagnosis between these two types can be quite challenging [8]. The well-known serum marker for PC is CA19-9 [9] however, nearly 10% of Caucasians lack expression of this antigen [10]. In patients with PNET Chromogranin A (CHGA) has been recommended as a reliable serum tumour marker. However, studies on its effectiveness in distinguishing between PNET and non-PNET cases have yielded inadequate and controversial results [11, 12].
Genome Wide Associated Studies (GWAS) have revealed polymorphisms that increase PC susceptibility in 23 genes [13]. Several genetic and epigenetic changes are required for a pancreatic tumour to arise [14], including mechanisms that greatly affect tumour progression such as mutations and dysregulated methylation [15, 16]. Moreover, CDKN2A/B variants in germlines and variations in several chromosomal regions increase susceptibility to the disease [17, 18]. However, it is evident that a deeper understanding of the disease’s pathophysiological mechanisms is necessary to improve our diagnostic and/or therapeutic approaches. Similarly, the discovery of new biomarkers is of utmost importance; these biomarkers will be valuable tools in the disease’s signs, symptoms, and prognosis for each different cancer phenotype [19].
Evidence suggested that non-coding RNAs are important epigenetic factors since they seem to affect several cellular events in malignancies, including metastasis. Recent evidence shows that the expression of non-coding RNAs longer than 200 nucleotides (long non-coding RNAs, lncRNAs) is dysregulated in cancer [20, 21, 22, 23]. LncRNAs are key mediators of pathways involved in tumour suppression and oncogenesis [20, 23, 24, 25] affecting important cellular processes, such as chromatin reprogramming, cis- and trans-regulation of gene expression and mRNA processing [26, 27, 28] though DNA-RNA interactions and assemblage of specific protein combinations [29].
Unfortunately, little progress has been made towards finding a molecule that could serve as a biomarker; there is still no biomarker available for clinical use. Given the fact that the general population does not opt for screening and that the disease is only curable in its most early stages, the need for a biomarker is critical [30]. GWAS have shown a connection between cancer and lncRNA polymorphisms [31, 32]; PVT1 and HOTAIR are two lncRNAs which have been found to be dysregulated in several cancer types [33, 34], including PC [35]. Specifically, in PC cases, both PVT1 and HOTAIR have been found to be upregulated [36].
LncRNA HOTAIR is located on chromosomal region 12q.13.13 and is transcribed by the anti-sense strand of HoxC gene [34]. Its levels can be used to predict an ongoing metastasis while its overexpression is linked with a very poor prognosis [33]. Inhibition of HOTAIR was associated with decreased pancreatic cell invasion and proliferation [37]. Regarding HOTAIR polymorphisms, several studies concluded that are associated with an increased risk of cancer [38]. Indeed, polymorphism rs4759314, which is included in our study, was found to increase the risk of gastric cancer [39].
PVT1 lies at the 8q24.21 chromosomal region that also includes the oncogene MYC and is often found in extra copies in human cancers [40]. Upregulation of PVT1 seems to be responsible for the epithelial-to-mesenchymal transition (EMT), a crucial event for cancer metastasis [41]. Another study [42] also showed that PVT1 promotes EMT and that it also downregulates p21. PVT1 could serve as a new prognostic biomarker for gastric cancer [43], while genetic variation of its chromosomal region seems to affect the risk of renal cancer [44]. However, the function of PVT1 in pancreatic cancer is poorly understood. Recently, Zhao et al. [45] revealed that PVT1 promotes the proliferation and migration of pancreatic cancer cells. PVT1 polymorphisms have been associated with several cancer risks including pancreatic cancer [46].
In our study we focused on two polymorphisms of PVT1, and HOTAIR (rs1561927, and rs4759314 respectively), aiming to evaluate the contribution of these lncRNA polymorphisms with PC (PDAC and PNET) risk.
Materials and methods
Patients
The study included 111 PDAC cases, 56 PNET cases (23 non-functional and 33 functional), and 125 healthy controls. Controls were healthy blood donors with no evidence of inflammation or pancreatitis. For subsequent analysis, we excluded cases and controls with known genetic syndromes (i.e., MEN1, MEN2, VHL or TSC). The diagnosis was made following standard procedures and was confirmed histopathologically from operatively resected tumors. The PDAC cases were assessed according to the staging system established by the American Joint Committee on Cancer [47], whereas the classification of PNETs was performed according to the updated World Health Organization (WHO) classification [48]. Before commencement of the study, the Ethical committee at the participating centers approved the recruitment protocols. All participants were informed regarding the study, and their written consent was provided.
Genotyping
DNA was isolated from peripheral blood with the NucleoSpin Blood Kit (Macherey-Nagel, Germany) according to the manufacturer’s instructions. The rs1561927 (G/A) polymorphism was amplified by PCR-RFLP with a set of primers, 5’-AACAAAGAGG GGAAGTAAGCTAGT-3’ and 5’-TGACATGAACAG GTTAAAGACAG-3’. The 40 cycles of PCR were performed at 95
Statistical analysis
Genotype frequencies were compared with the chi-square with Yate’s correction using S-Plus (v. 6.2, Insightful, Seattle, WA, USA). Odds ratios (ORs) and 95% confidence intervals (CIs) were obtained with Graph Pad (v. 3.00, GraphPad Software, San Diego, CA). The
Clinicopathological data of the PDAC, PNET patients and controls
Clinicopathological data of the PDAC, PNET patients and controls
Table 1 shows the clinicopathological data of the studied population. Age and gender distributions were similar between patients and controls. Additionally, the distributions of smoking status, and alcohol consumption, were not different between the PDAC and PNET patients and controls (
Genotype and allele frequencies of the PVT1 rs1561927, and HOTAIR rs4759314 polymorphisms in PDAC, PNET, and controls
Genotype and allele frequencies of the PVT1 rs1561927, and HOTAIR rs4759314 polymorphisms in PDAC, PNET, and controls
PVT1 rs1561927 genotypes were associated with overall survival of PDAC patients
Cox proportional hazard estimation of overall survival
Association of overall survival with genotypes of PVT1 rs1561927 polymorphism. Kaplan-Mayer curve represents the percentage of survival in months of patient’s different genotypes of the rs1561927 polymorphism.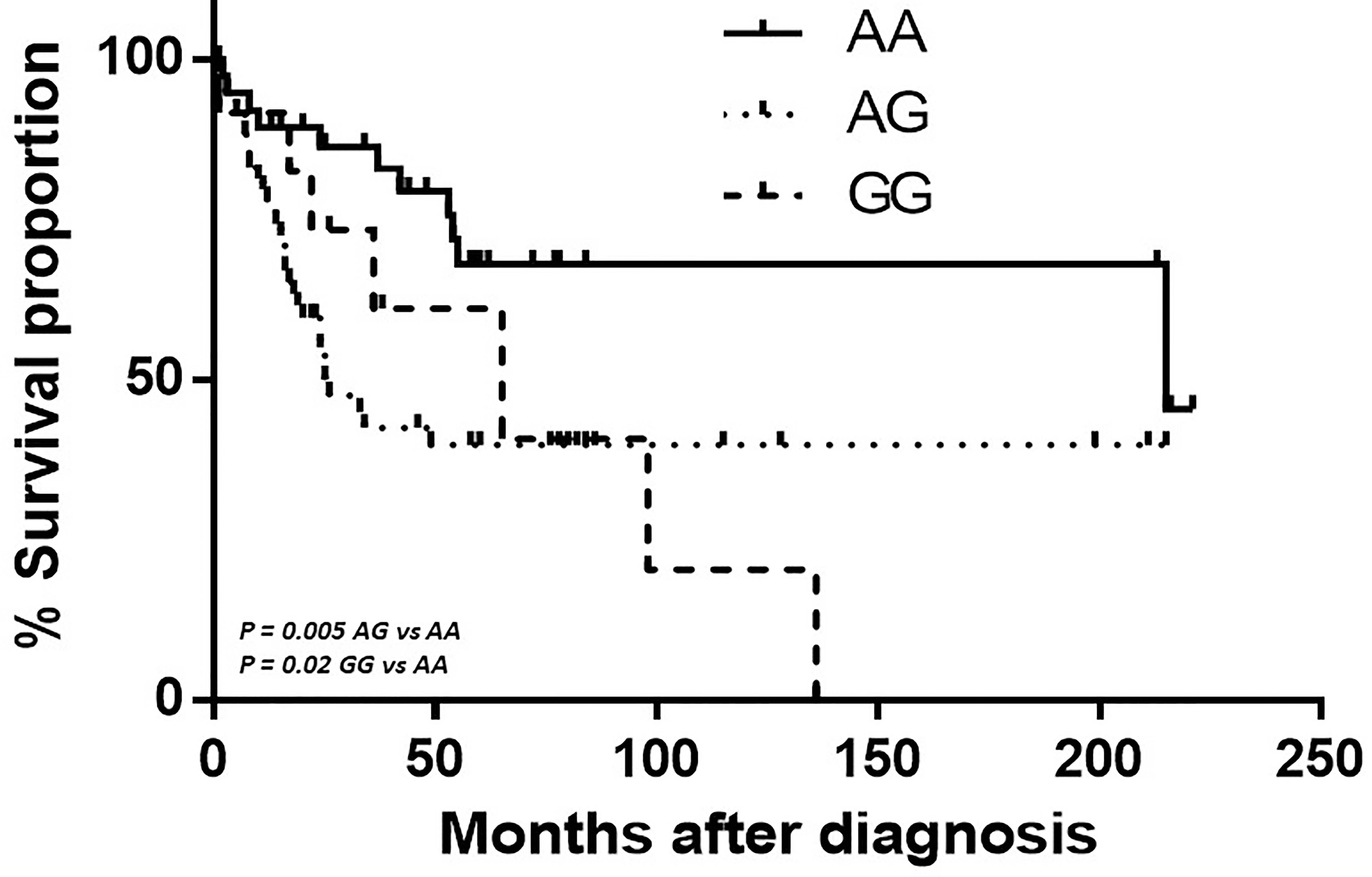
The genotype frequencies of PVT1 rs1561927, and HOTAIR rs4759314 polymorphisms between PDAC, PNET, patients, and controls are compared in Table 2. All genotype distributions were in Hardy-Weinberg equilibrium. The distribution of genotypes for the PVT1 rs1561927 polymorphism in the PDAC and PNET patient groups showed significant difference compared to the control group. The G allele of the PVT1 rs1561927 polymorphism was significantly overrepresented in both PDAC (10.81% vs 4.8%,
Follow-up information regarding survival was available only for PDAC patients. Since in our findings the PVT1 rs1561927 genotypes were found to be associated with PDAC risk, the hazard ratios of the rs1561927 genotypes of the PDAC patients for overall specific survival are presented in Table 3. The median duration of the follow-up was 25 months (range, 1–221 months). A total of 49 PDAC patients suffered cancer-related death during the follow-up period. The PVT1 rs1561927 AG and GG genotypes were significantly associated with poor survival (
Rising evidence has shown that lncRNAs play important roles in the pathogenesis of various cancers, including pancreatic cancer [49]. Several studies have shown higher levels of different lncRNA, such as H19, HOTAIR, HOTTIP, and MALAT-1 in pancreatic cancer tissue samples [50]. However, the PDAC, and PNET associated lncRNAs are still not well-documented.
In the present study, the association between SNPs in two key lncRNA genes (PVT1, and HOTAIR) and the risk of PDAC and PNET was analyzed. Our results showed a significant association of PVT1 rs1561927 G allele and the risk of PDAC and PNET in studied population. Recent findings reported that increased expression of lncRNA PVT1 in PDAC correlated with tumor progression and suggested that PVT1 might be a potential biomarker for predicting the prognosis of patients with PDAC [51]. Regarding the PDAC, our findings are in accordance with Wolpin et al. [52] and Zhang et al. [53] that identified a risk locus for pancreatic cancer at 8q24.21 in a non-genic region between PVT1 and LINC00977 that approached genome-wide significance located 455 kb telomeric of PVT1. However, in our study we found that PVT1 rs1561927 G allele is the risk allele in our population, as was also reported by Wolpin et al. [52] for Southern Europe population. Even if it is surprising that the PVT1 rs1561927 G allele was found in the present study to be associated with PNET risk also, our results confirmed previous study by Canzian et al. [54] and Obazee et al. [55], indicating that PDAC and PNET may share some common genetic background, and suggesting that common variants associated with PDAC carcinogenesis possibly influence carcinogenesis of pancreatic beta cells in a similar way as in exocrine pancreatic cells. Additionally, Haugvik et al. reported that although there is no obvious link between environmental factoes and risk of developing PNET, an overlap of risk factors in PDAC and PNET is possible [56]. You et al. [57] identified PVT1 as a regulator of Gemcitabine sensitivity in pancreatic cancer cells. The PVT1 is a non-protein encoding gene and seems to be implicated in the carcinogenesis of different types of human cancers, including pancreatic cancer, possibly through DNA rearrangement and co-amplifications with MYC. Additionally, it is noteworthy that our study provided indication for the association between PVT1 rs1561927 polymorphism and overall survival, on the grounds of lack of any association of the rs1561927 polymorphism with the tumor stage that is a typical prognostic parameter, that were taken into consideration. The possible association of rs1561927 AG/GG genotypes with poor survival merits further evaluation, since its significance might have significant prognostic and therapeutic implications. Further studies on the expression and regulation pathways of PVT1 may reveal the mechanisms of its action in pancreatic carcinogenesis.
Regarding the HOTAIR, recently polymorphisms in the HOX transcript antisense intergenic RNA (HOTAIR) have been associated with risk to different cancers [39]. HOTAIR reported to be involved in cancer proliferation, migration, invasion, progression, and poor prognosis [33]. Even if has been suggested that rs4759314 was a risk factor for gastric cancer mainly in Chinese population [58], we did not find any significant association between rs4759314 polymorphism and PDAC. Our results agree with the findings by Hu et al. [59] that they did not also find any significant association of rs4759314 polymorphism in pancreatic cancer, and with the results of meta-analysis by Zhang et al. [38] that they also did not find a significant association between rs4759314 (A
There is limited information available on lncRNAs, and pancreatic carcinogenesis in the literature, however, our results, confirm the recent observation that PVT1 rs1561927 polymorphism is associated with the risk of developing PDAC and PNET. Regarding the HOTAIR rs4759314 polymorphism, no significant association was noted with PDAC risk, but our results indicate that might be contributed to PNET risk. Wide functional researches and well-designed genetic studies in different ethnic groups are necessary to confirm and extend our observations.
Footnotes
Acknowledgments
This research is partly supported by the Hellenic Society of Medical Oncology (HeSMO) by a research grant to G. Aravantinos and M. Gazouli.
Conflict of interest
The authors declare that they do not have any conflict of interest that could influence this work.
