Abstract
BACKGROUND
: Nucleoporin NUP153 (NUP153) is well known to be involved in the regulating of nuclear transport. Although NUP153 is associated with several cancers, its role in colorectal cancer (CRC) and the underlying mechanism are still unknown.
OBJECTIVE:
The aim of this study was to access the effect of NUP153 on the prognosis of patients with CRC, and cancer cell proliferation.
METHODS:
The expression levels of NUP153 in CRC tissues and matched normal colon tissues were examined by real-time quantitative PCR and immunohistochemistry. Then the association between NUP153 levels with clinical variables as well as survival time was investigated. Moreover, overexpression of NUP153 in HCT116 cells was established to study its influence on cell proliferation in vitro, and a xenograft model was performed to explore this effect in vivo.
RESULTS:
We found that NUP153 was highly expressed in adjacent normal tissues than in cancer tissues, and elevated NUP153 expression was negatively associated with pathological grade (
CONCLUSIONS:
NUP153 might be a promising prognostic factor, a potential tumor suppressor and therapeutic target in human CRC through an interaction with the Wnt/
Introduction
Colorectal cancer (CRC) is one of the most common gastrointestinal cancer worldwide, with 37,000 newly diagnosed cases and 19,000 estimated deaths in China in 2015 [1, 2]. Although new methods for CRC diagnosis and treatment have continuously emerged, the survival rate of patients in advanced CRC remains low [3, 4]. Many tumor related gene mutations are identified to associate with prognosis of patients with CRC. Thus, understanding the role of regulatory genes and its related pathway in CRC is essential to optimizing current therapeutic strategies.
Successive acquisition of genetic alterations in Wnt pathway are the driving force for several diseases, especially for CRC [5, 6]. Several key molecular gene mutations in this pathway causing CRC formation have been identified [7]. Namely that, adenomatous polyposis coli (APC) mutations occurs in 90% of spontaneous colorectal cancer [8]. Activating Axin2 mutations occurs in some cases of CRC patients. Therefore, Wnt/
NUP153 is an essential part of the structure of mammalian nuclear pore complexes (NPCs) and is a key part of NPCs anchoring on the surface of the nuclear membrane and plays an important role in nuclear transport [12, 13]. More studies about NUP153 are focusing on embryo development, HIV and chromosomal translocation [14, 15, 16]. For example, Toda et al. reported that NUP153 interacted with sox2 to enable bimodal gene regulation and maintenance of neural progenitor cells, which could control cell fate [17]. Nanni et al. indicated that NUP153 was an epigenetic regulator, associating with chromatin and regulating cardiac gene expression in dystrophic mdx hearts [18]. Furthermore, NUP153 is reported to be associated with several cancers. In human breast carcinoma, silencing NUP153 in MDA231 cell line significantly reduced the speed of directional cell migration [19]. Chromosomal rearrangements of NUP153 were related with its increased expression in urothelial and retinoblastoma cancer [20]. Shain et al. revealed a possible oncogenic functions of NUP153 by regulating TCF
Here we explored the role of NUP153 in colorectal cancer. First we found that NUP153 highly expressed in adjacent normal tissues compared with the cancer tissues, and patients with higher NUP153 expression had a good prognosis. Also, our work uncovered that NUP153 suppressed the proliferation of CRC and inactivated the Wnt/
Materials and methods
Human samples
With the approval of the Ethnic Committee of Fifth Hospital of Shanghai, Fudan University. A total of 18 primary CRC (I–II: 11, III–IV: 7; T1–2: 9, T3–4: 9; Tumor diameter: 5.00
Cell culture and materials
Plasmids of NUP153, Topflash vector, p-cDNA3.1, renilla luciferase pRL-TK reporter vector were got from East China Normal U niversity. Two hundred ninety three T and HCT116 cells were maintained in DMEM with 10% fetal bovine serum (Gibco) in 37
RNA extraction and qRT-PCR
Total cellular or tissue RNA was extracted from the different groups using trizol reagent (Invitrogen, MA, USA), the cDNAs were amplified by PCR using Thunderbird SYBR qPCR Mix (Toyobo, Osaka, Japan) in the Applied Bio-systems 7500 Real-Time PCR System (Applied Biosystems, Foster City, CA, USA). The thermal cycler protocols included 3 min at 95
Human primers set for qRT-PCR analysis
Human primers set for qRT-PCR analysis
Immunohistochemistry for NUP153 expression in CRC and adjacent non-tumor samples was performed using standard methods. Colorectal cancer tissue microarray (TMA) was deparaffinized by xylene, followed by rehydration through a graded series of ethanol, and then incubated with blocking solution for 30 min at room temperature. Next, tissue sections were successively incubated with rabbit anti-NUP153 diluted 1:100 (ab96462; abcam; US) overnight at 4
Western blots
Cell lysis buffer was used to extracted proteins. Subsequently, those proteins were boiled with SDS sample buffer and separated by sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE). Then, the proteins were transferred to PVDF membranes (Millipore, USA). After transmembrane, membranes contained proteins were respectively incubated with anti-NUP153 antibody (1:500; ab96462; abcam; US) overnight at 4
MTT assay
HCT116-NUP153 cells were seeded into 96-well plates at a density of 4,000 cells per well and cultured for 24 h at 37
Clone formation assay
HCT116-NUP153 cells were seeded into 6-well plates at a density of 1,000 cells per well and cultured for 8 days at 37
Xenograft experiment
Nude mice aged 5 weeks were purchased from the Experimental Animal Center of East China Normal University. To demonstrate the effect of NUP153 on inhibit tumor growth in vivo, HCT116-NUP153 cells were implanted into nude mice at 2
Luciferase reporter assay
HCT116-NUP153 cells were plated at a density of 4
The expression levels of NUP153 in colorectal cancer (CRC). (A) qRT-PCR analysis showed that the expression levels of NUP153 mRNA highly expressed in adjacent normal tissues compared with cancer tissues. (B) IHC analysis for the expression pattern of NUP153 protein in CRC. Strong expression in normal colorectal mucosa. The staining of NUP153 protein was mainly located in the membrane of cells. Scale bars 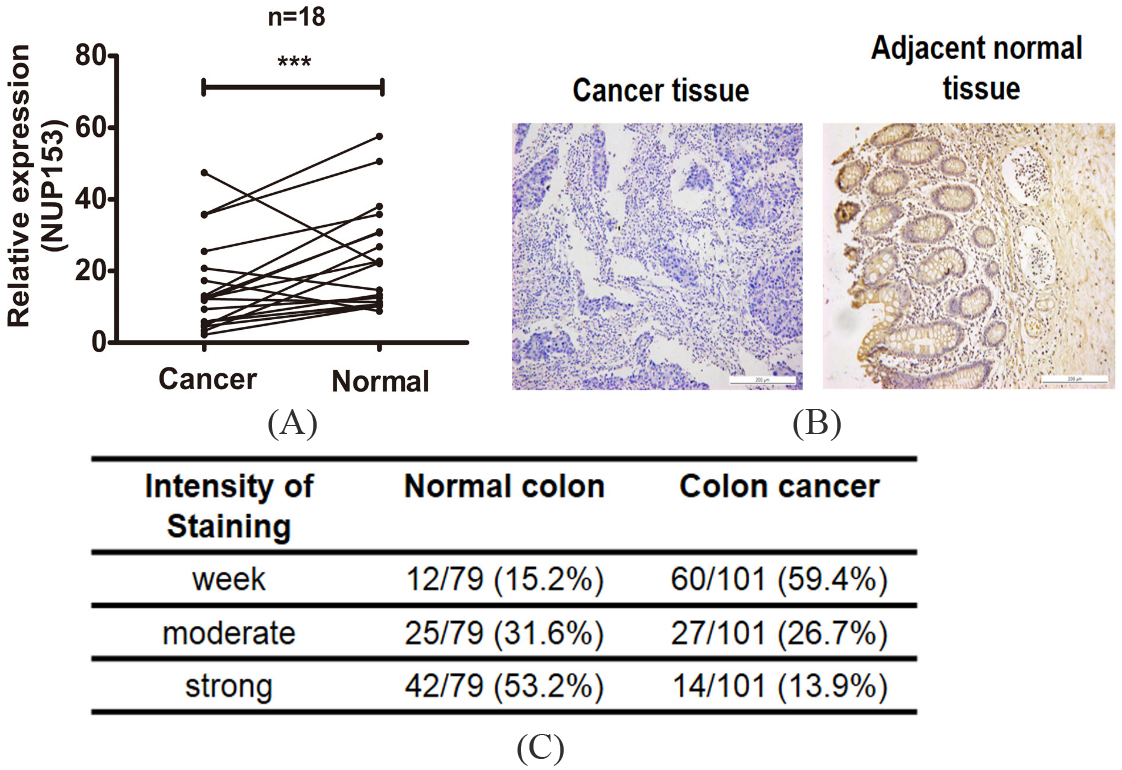
All statistical analyses were performed using SPSS17.0 (Chicago, IL, USA). Graphpad Prism 5.0 (La Jolla, CA, USA) was used for graphs. Student t test was used to compare the expression levels of NUP153 in cancer tissues and matched normal non-cancer tissues. Chi-square test was used to analyze the correlation between NUP153 expression and clinicopatholigical features. Cumulative survival time was calculated by the Kaplan-Meier method and analyzed by the log-rank test. Logistic regression analysis was performed to assess the effects of NUP153 on patients’ OS and tumor recurrence.
Results
Expression of NUP153 in colorectal cancer samples
To explore the expression pattern of NUP153 in cancer tissues. We first detected the expression levels of NUP153 mRNA in 18 pairs of CRC tumor tissue with matched normal tissue. The result showed that NUP153 was highly expressed in normal tissues than cancer tissues (Fig. 1A). Furthermore, we purchased a colorectal cancer tissue microarray (TMA) with complete pathological information. Beside 101 cases of cancer tissues, 79 matched normal tissues spots were found on this TMA. As showed in colorectal adenocarcinoma cells, NUP153 was found to be stained mainly in the membrane of cells (Fig. 1B). Meanwhile, two pathologists with no knowledge of the clinical characteristics of the patients were employed to score staining intensity (0–4) and extent (0–100) of NUP153. As showed in Fig. 1C, NUP153 expression was markedly reduced in the cancer tissues compared with the normal tissues. 69.3% colorectal cancer tissues showed negative staining for NUP153, while 84.8% normal tissues showed obvious positive staining.
Association between NUP153 and clinicopatholigical characteristics in TMA cohort
Association between NUP153 and clinicopatholigical characteristics in TMA cohort
*P-values
According to the information of IHC, we further investigated the association between NUP153 expression and clinicopatholigical features. As show in Table 2, higher NUP153 expression was positively associated with better tumor outcomes: pathological grade (
Correlation between NUP153 expression with OS and RFS
In order to explore whether patients with higher NUP153 expression could prolong their survival time, we then analyzed the association between their survival information and NUP153 expression. As showed in Fig. 2A, patients with higher NUP153 expression was found to have a favorable 9-years OS (log-rank
Kaplan-Meier curves for OS rates between CRC patients with high and low NUP153 expression. (A and B) Survival analysis showed that patients with higher NUP153 expression was significantly had a longer OS and RFS (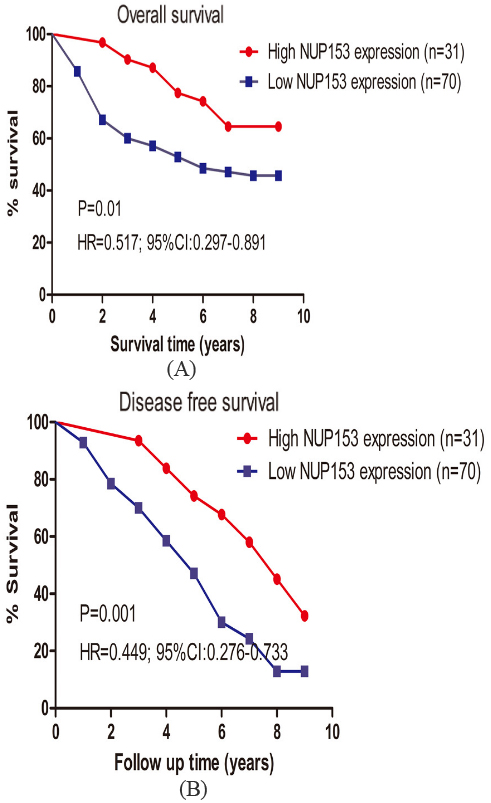
The influence of NUP153 expression on OS and recurrence were then analyzed based on TMA data. Univariate logistic regression analysis showed that higher NUP153 expression (
Overexpression of NUP153 inhibited the proliferation of CRC
In order to determine the function of NUP153 in CRC cells, we overexpressed NUP153 in HCT116 human colon adenocarcinoma cell line. Protein levels were markedly increased after transfection of NUP153 plasmid (Fig. 3A). The MTT experiment showed that cell growth was markedly reduced in HCT116 cells in responsive to NUP153 overexpression (Fig. 3B). We further used soft agar colony formation assay to compare cell grow of HCT116 cells with or without NUP153 plasmid (Fig. 3C and D). Consistent with MTT results, overexpression of NUP153 significantly reduced cell proliferation compared with the control, indicating a tumor suppressor potential for NUP153. In xenograft experiment, compared with control group, the results showed that the tumor weights and volumes of node mice injected with HCT-NUP153 cells were significantly smaller (Fig. 3E–G).
Univariate and multivariate analyses for over survival (OS) in TMA cohort
Univariate and multivariate analyses for over survival (OS) in TMA cohort
Univariate and multivariate analyses for recurrence in TMA cohort
NUP153 overexpression suppressed the proliferation of CRC. (A) Western blot blanket analysis of NUP153 protein blanket expression in HCT116 cells treated with or without NUP153 plasmid. (B) After 72 hours post transfection with or without NUP153, HCT116 cell viability was monitored by MTT assay at the indicated times. (C and D) Clone formation assay and statistical analysis for HCT116 cells transfected with or without NUP153, respectively. The experiments were repeated independently for three times. (E–G) Cells were subcutaneously injected into the flank of nude mice. Tumor volumes and weights were measured on the indicated days to evaluate the effects of NUP153 on tumor growth. Data points are showed as the mean 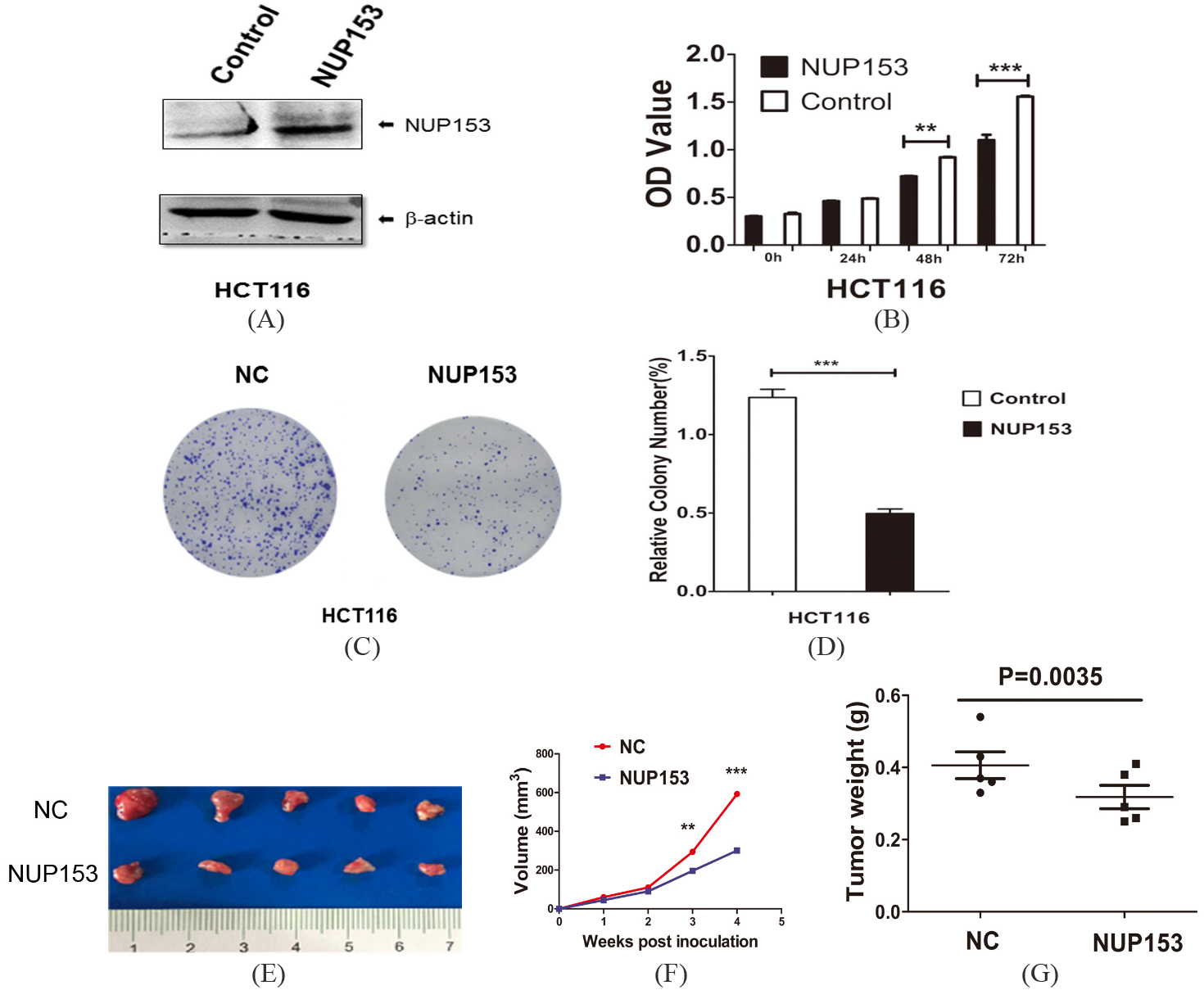
NUP153 overexpression in HCT116 cells inhibited the Wnt/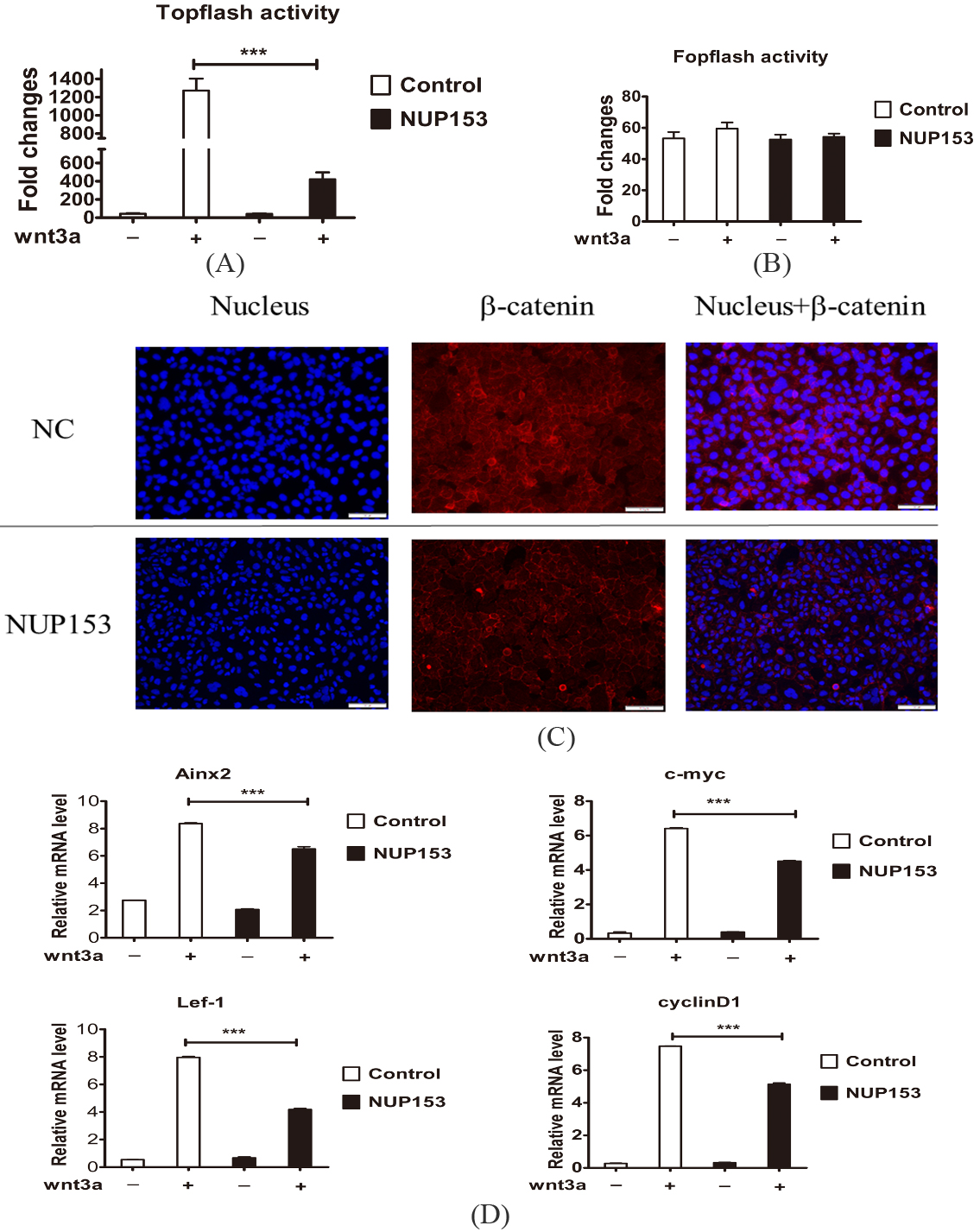
According to the previous studies, we had found that NUP153 could inactivate the Wnt/
Discussion
Colorectal cancer (CRC) develops through a series of genetic and epigenetic changes that cause the transformation of normal mucosa to a premalignant polyp, and finally to a cancer [22, 23, 24]. These regulatory genes are found to be closely associated with prognosis of patients, and also provide strong evidences for disease diagnosis and treatment [25, 26]. Some studies have reported that the role of the human NUP153 gene accounts for different type of cancers. However, the role for NUP153 in CRC tumorigenesis remains unknown.
In this study, our work first discovered that NUP153 was significantly down-regulated in colorectal cancer tissues, which suggested that NUP153 might be a potential suppressor gene in human CRC. Currently, cancer recurrence and metastasis remain to be the leading death cause for CRC patients. As reported, the 5-years survival time of patients with early tumor stage was up to 90%, while only 14% in those with distant metastasis [27]. According to our study, the results showed that higher NUP153 expression was positively related with some better tumor outcomes: pathological grade, T stage and distant metastasis. These findings supported that NUP153 might work as favorable biomarker in screening out those with risk factors. In order to demonstrate to effects of NUP153 on survival time, we followed up all of the patients for 9 years, and found out that patients with lower NUP153 expression had a risk in OS and RFS. Next, logistic regression analysis demonstrated that higher NUP153 expression increased the favorable prognosis of patients with CRC in OS and recurrence, which identified NUP153 as an independent safe prognostic indicator. As a result, we have revealed the clinical significance of NUP153 in CRC formation, which might function as a useful therapeutic target for CRC in the future. Just as KRAS mutation status, which is a strong predictive marker of resistance to EGFR-targeted therapy in patients with metastatic colorectal cancer [26, 28, 29].
NUP153 was reported to associate with cancer cells invasion, proliferation, and metastasis [30]. Both estrogens and NO cooperate and contribute to aggressiveness in prostate cancer through Nup153 upregulation [31]. Our work showed that NUP153 overexpression markedly suppressed colorectal cancer cell proliferation, which uncovered that NUP153 could inhibit the progression of CRC formation. Consistent with the result, our xenograft model experiment also confirmed that NUP153 could inhibit tumor growth in vivo. Thus, these findings support that NUP153 might be a potential biomarker and a therapeutic target of CRC through its interaction with various signaling pathway involved in this disease.
Wnt/
NUP153 expression was generally down-regulated in CRC, and patients with higher NUP153 expression was correlated with a longer OS and RFS. Meanwhile, NUP153 was identified as an independent safe prognostic factor in OS and tumor recurrence. Furthermore, overexpression of NUP153 was found to inhibit the proliferation of CRC via inactivating Wnt/
Footnotes
Acknowledgments
This study was funded by Science and Technology Commission of Shanghai Municipally (No. 17411967 600), Science and Technology Commission of Minhang Municipally (No. 2015MW07).
