Abstract
BACKGROUND:
Circulating tumor cells (CTCs) and circulating cell free DNA (ccfDNA) represent a liquid biopsy of a tumor allowing real time disease monitoring especially in advanced stages of cancer, but their analysis is technically challenging.
OBJECTIVE:
We aimed to demonstrate the feasibility of two different technical approaches to detect the BRAFV600E mutation in the liquid biopsy of 20 metastatic melanoma patients by using both the enriched CTC fraction and circulating ccfDNA from the same blood sample.
METHODS:
We detected CTCs by a filtration method in 20 metastatic melanoma patients and detected the BRAFV600E variant on CTCs and ccfDNA by an allele-specific qPCR assay; the mutated samples were confirmed by ICE-COLD PCR followed by Sanger sequencing.
RESULTS:
We found CTCs in 70% of the samples, and identified the BRAFV600E variant on CTCs. We correlated the results with those obtained on ccfDNA from the same blood draw. We found some discordant results between CTCs and ccfDNA.
CONCLUSIONS:
Our results underline the importance of investigating both CTCs and ccfDNA in a liquid biopsy approach to melanoma patients.
Keywords
Introduction
Circulating tumor cells (CTCs) and circulating cell free DNA (ccfDNA) represent an interesting investigation option for the study of cancer and its dissemination [1]. They recently reached popularity as the liquid biopsy of a tumor representing surrogate samples of tumor tissues. They appear in blood as early markers of cancer presence, presumably before appearance of tumor symptoms, and their persistence in the circulation has been indicated as an unfavourable prognostic index in many types of cancers including melanoma [2, 3].
Detection and counting of circulating tumor cells in metastatic cancer patients allowed the identification of a population characterized by a shorter survival rate [4]. Methodological challenges of CTC detection are mainly related to the requirement of high sensitivity combined with high specificity and to the identification of suitable markers for CTC enrichment. Since invasion can start very early during tumor development, identification and counting of these very rare cells (few CTCs per 10 ml of blood, which means few CTCs mixed with approximately 100 million leukocytes and 50 billion erythrocytes) could alert the oncologist about a developing tumor invasion process [5]. Specificity is also an absolute requirement in this field. In fact, a wrong identification of “non-tumor cells” (like epithelial non tumor cells, for instance) as “tumor cells”, could generate poor clinical and therapeutical choices having a negative impact on the quality and/or expectancy of life in patients with cancer.
Several recent reviews concerning the detection of CTCs have been published [1, 2, 3, 4, 6, 7, 8]. Many different methods have been developed and the production of commercially available CTC assays is increasing exponentially during the last few years. All of them possess advantages and disadvantages that have been recently reviewed in several papers [2, 3, 8, 9, 10].
Analogously, the detection of ccfDNA of tumor origin is rather challenging to be accomplished. ccfDNA has been proposed as a source of cancer biomarkers since the discovery of the presence of the same genetic and epigenetic changes in plasma DNA and primary tissue [11]. The studies conducted so far based on the measurement of total ccfDNA concentration showed limited sensitivity and specificity in distinguishing patients from healthy controls, preventing the achievement of any definitive conclusion on the diagnostic impact of this parameter. The identification of the fraction of ccfDNA deriving from the tumor may help reaching a higher specificity [12]. This task can be accomplished by detecting tumor specific alterations, such as genetic variants, among which BRAFV600E somatic mutation has been identified as a driver mutation particularly in melanoma and thyroid tumors.
The development of target-specific therapies directed against molecular alterations found in tumors with high frequency requires the identification of the mutational status of the neoplastic cells that can be easily studied within the tumor mass at the time of cancer surgical removal.
However, especially in patients with advanced or metastatic cancer, tumor tissue sampling can be difficult to obtain due to bad general conditions preventing surgery and to the risk related to biopsy the metastasis. Thus the molecular analysis of CTCs and/or ccfDNA would allow the development of non-invasive biomarkers of potential great interest for oncologists and benefit for patients.
Spreading of CTCs and of clusters of CTCs, Circulating Tumor Microemboli (CTM), in blood represents a key upstream step in the process driving the formation of metastases [5, 13, 14]. We previously demonstrated that CTCs are very heterogeneous even within a single patient [15, 16]; CTC characterization is thus fundamental to study their invasiveness and metastatic potential. As distant metastases are the major problems when treating solid tumors, the development of a specific, sensitive, non-invasive diagnostic method for early detection and monitoring of CTCs is expected to greatly improve patient prognosis. The early spreading of tumor cells in patients with no overt metastasis eludes detection, even by high-resolution imaging techniques. This has an impact on early intervention efforts. The limited or difficult availability of tumor tissues and the need to follow the mutational evolution of the invasive tumor cell population (in order to optimize the use of expensive targeted treatments) point to the analyses of circulating tumor cells as a future goal of utmost importance in clinical oncology. Because tumor cells evolve clonally, the detection of mutations in CTCs is clinically more relevant than in the primary tumor [17] allowing a real-time follow-up of disease progression [18].
We previously published the feasibility of CTC molecular analysis on CTCs isolated by blood filtration [19] evaluating HER2-neu amplification in CTCs isolated by ISET in breast cancer patients as a biomarker of tumor response to the targeted therapy trastuzumab (Herceptin
Here, we demonstrate the feasibility of two different technical approaches to assess BRAF mutational status in the liquid biopsy of metastatic melanoma patients by using both the enriched CTC fraction and circulating cell-free DNA from the same blood sample.
Our primary aim was to test the ScreenCell device [20] for molecular biology application followed by a qPCR assay method to detect BRAFV600E mutation. The reliability of the qPCR method was confirmed on selected samples by an ICE-COLD sequencing technique (Transgenomic, Omaha, NE, USA). In addition, we performed the quantitative measurement of the percentage of BRAFV600E mutated alleles in circulating cell-free DNA from the plasma obtained by the same blood draw. We finally correlated the results obtained by investigating the two different blood compartments for the presence of BRAFV600E mutation. The protocol of the study is schematically represented in Fig. 1.
Scheme of the study protocol.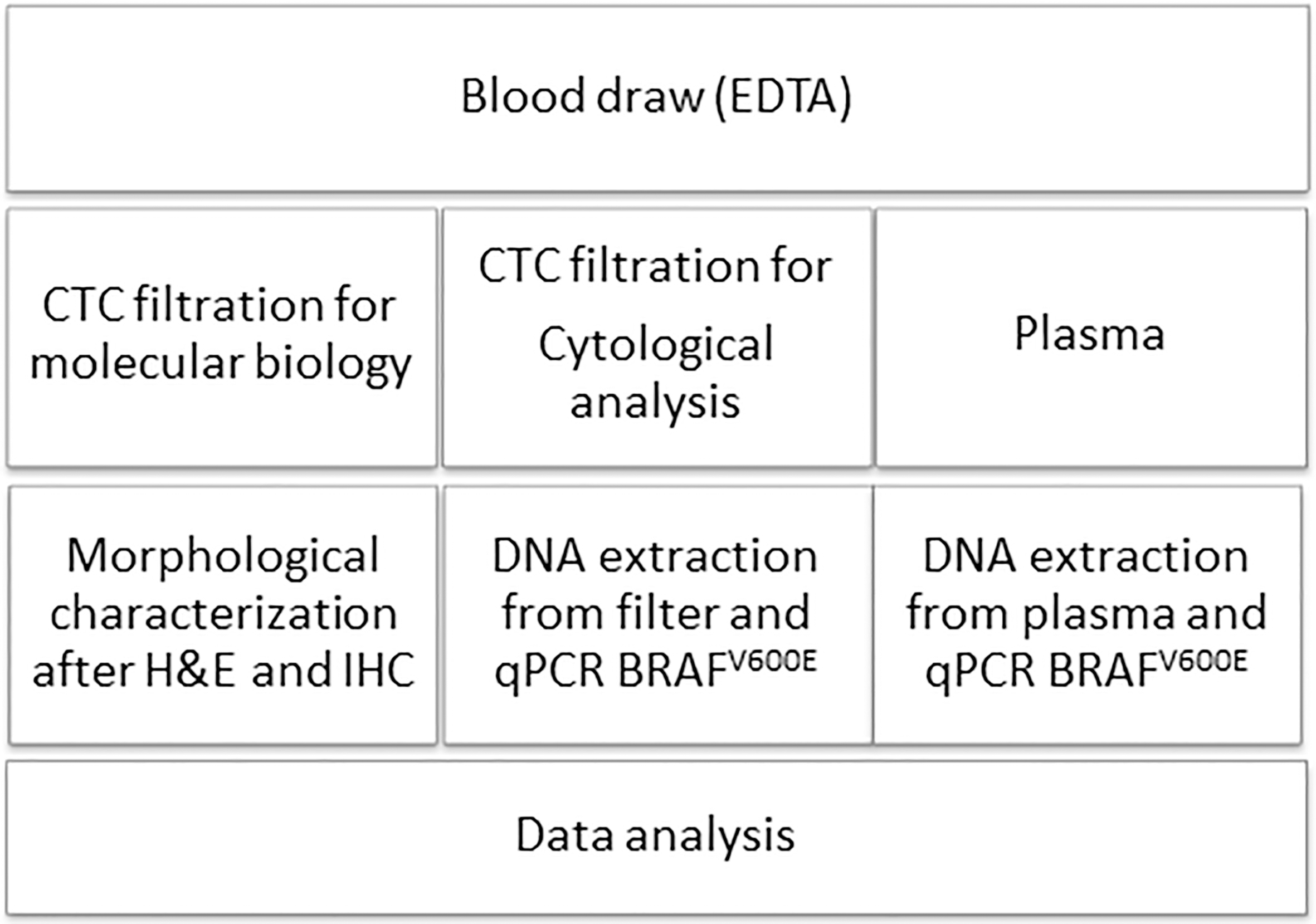
Patients and samples
Twenty metastatic melanoma consecutive patients were recruited at the Dermatology Unit, Department of Health Sciences, University of Florence, Italy. The group included patients in Stage IV disease who had developed distant metastases, the commonest sites being lung, liver, and bone.
Patients were submitted to venipuncture to collect three ethylenediaminetetraacetic (EDTA) blood samples. All blood samples were transported immediately to the laboratory and processed within 3 hours from drawing. Six ml blood were used for ScreenCell Cyto filtering in duplicate, while additional 6 ml were used in the molecular biology (MB) ScreenCell device. Cells on MB filters were devoted to DNA extraction. Moreover, 5 ml of blood were centrifuged twice at 4
All the patients gave their written informed consent to the study. The study was approved by the Local Ethical Committee. The study complies with the World Medical Association Declaration of Helsinki regarding ethical conduct of research involving human subjects.
In addition, for a subgroup of patients (5/20) formalin fixed paraffin embedded (FFPE) tissues have been recovered from the archives of the Anatomo-Pathology Institute of the University of Florence.
Cytological Evaluation of CTCs
Blood samples from 20 metastatic melanoma patients were filtered by the ScreenCell
Briefly, peripheral blood (2
CTCs were identified on the basis of the following morphologic criteria: cell size
CTC selection and DNA extraction from ScreenCell MB devices for molecular biology analysis
Blood was filtered by the ScreenCell
Briefly, before filtration the blood sample is diluted in the ScreenCell
DNA extraction methods for plasma and tissues
Cell-free circulating DNA was extracted from 500
DNA from formalin fixed paraffin embedded tissues was extracted by using the FFPE Tissue kit (Qiagen).
Cell-free DNA quantity and quality evaluation
The quantity and quality (in terms of DNA integrity) of ccfDNA were assessed by qPCR. In particular, a DNA integrity index was calculated by means of two qPCR assays targeting respectively a 67 bp and a 180 bp sequence on the single copy gene APP (Amyloid Precursor protein, chr. 21q21.2, accession NM_000484), as already published [22]. The primers and probe for the 67 bp amplicon were previously reported in ref. [23], while the reverse primer designed on the same target sequence for the 180 bp amplicon is reported in ref. [22]. The shorter amplicon (67 bp) was used to quantify total ccfDNA, while the ratio between the absolute concentration of the longer amplicon (180 bp) and the shorter one (67 bp) defined the integrity index 180/67, which was used to assess the fragmentation of ccfDNA.
The reactions were carried out in a 12.5
BRAFV600E mutational status
All patients were simultaneously investigated for BRAFV600E mutational status by analyzing DNA extracted by ScreenCell MB enriched samples. The investigation was performed by a modified version of the previously published allele specific real-time qPCR assay [24] on all the obtained DNA extracts (from CTCs and plasma). We designed a mutation-specific forward primer (5
The procedure was applied to both CTC enriched samples and plasma ccfDNA. Briefly, DNA extracted from the ScreenCell MB eluate was used to measure BRAFV600E mutated and wild type allele in distinct reactions containing the proprietary primers and probe for each variant.
qPCR for both variants was performed by denaturation at 95
The standard curve for BRAFV600E consisted of six dilutions (100%, 50%, 20%, 10%, 1% and 0% mutated alleles) obtained by mixing DNA from mutant SKMEL28 cell line and wild type MCF7 cell line. Linear regression analysis of the 2
The mutated samples were further confirmed by ICE-COLD PCR by using the REVEAL BRAF exon 15 kit (Transgenomic Inc, USA) according to the manufacturer’s instructions, followed by Sanger Sequencing according to a previously reported protocol [25].
The mutational status of DNA from FFPE tissues was assessed by conventional PCR followed by Sanger Sequencing using a previously reported protocol [25].
Results
Detection of circulating melanoma cells
CTCs were detected in 14 over 20 metastatic mel-anoma patients (70%). Tumor cells were observed mostly in single units (Fig. 2).
Picture of a single melanoma cell isolated on the ScreenCell Cyto filter.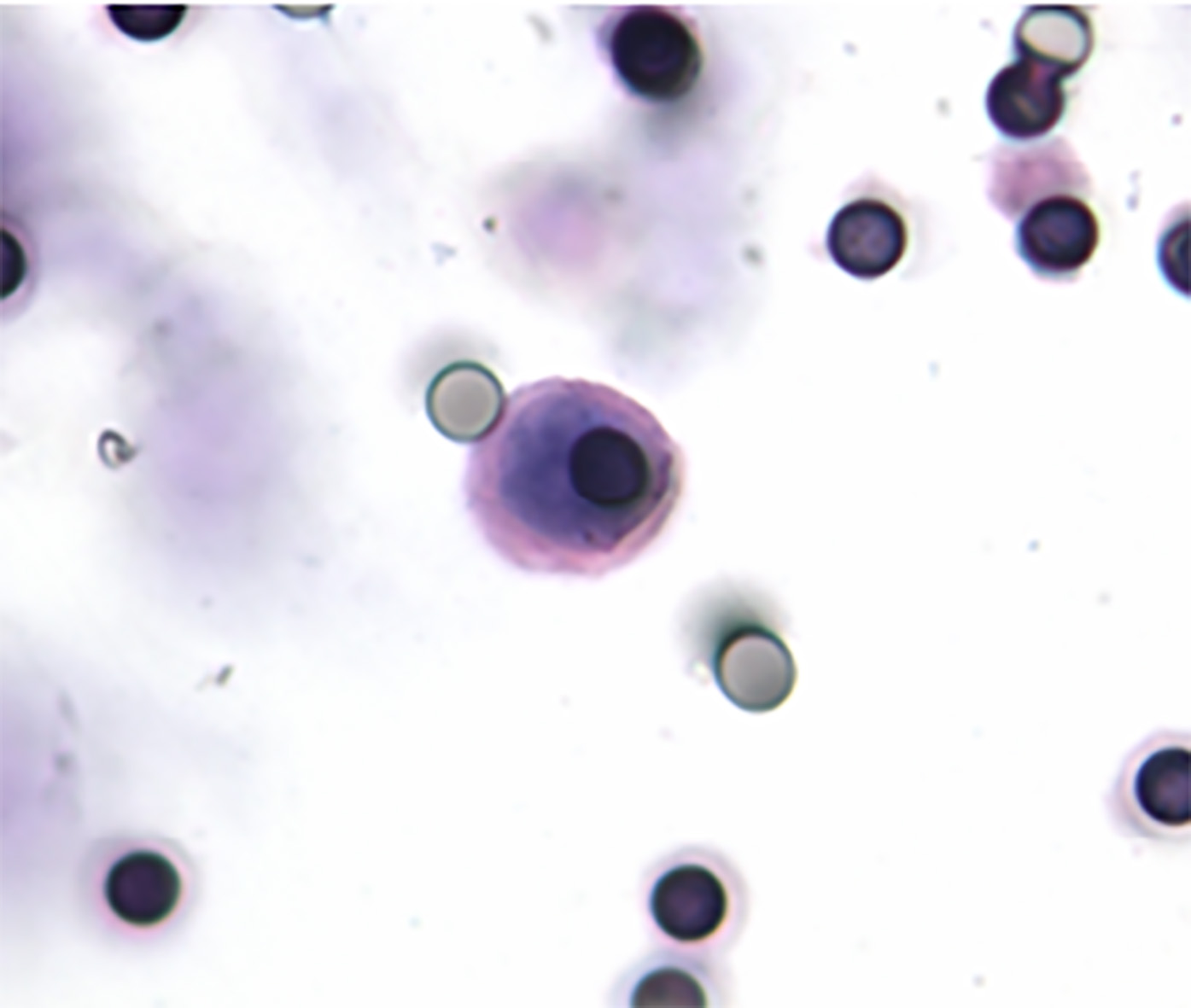
The allele specific qPCR method was modified in order to achieve data normalization on the basis of the BRAF wild type allele. In Fig. 3 (left panel) a typical standard curve for the evaluation of BRAFV600E percentage on DNA extracted from filters is reported. The standard curve derives from plotting the percentage of BRAFV600E mutated allele on the x axis and the 2
Detection of BRAFV600E mutation in CTC enriched samples and in corresponding tumor tissue
All patients were investigated for BRAFV600E mutational status by analyzing DNA extracted by ScreenCell MB enriched samples. All the obtained DNA extracts were successfully amplified by the allele specific qPCR assay. Among the 14 samples positive to the cytological analysis on the ScreenCell Cyto duplicate filters, 5 showed the presence of BRAFV600E mutation. The quantitative evaluation of the percentage of mutated allele resulted 1.37, 1.54, 5.26, 19.2 and 32.54 respectively (mean
Results related to the identification of CTCs by cytomorphological analysis of the ScreenCell Cyto filters and to the detection of BRAFV600E in CTCs and ccfDNA by qPCR
Results related to the identification of CTCs by cytomorphological analysis of the ScreenCell Cyto filters and to the detection of BRAFV600E in CTCs and ccfDNA by qPCR
Evaluation of the BRAFV600E assay. Left panel: typical standard curve for the evaluation of BRAFV600E percentage on DNA extracted from filters. Right panel: dilution test on control samples demonstrating the constancy of the 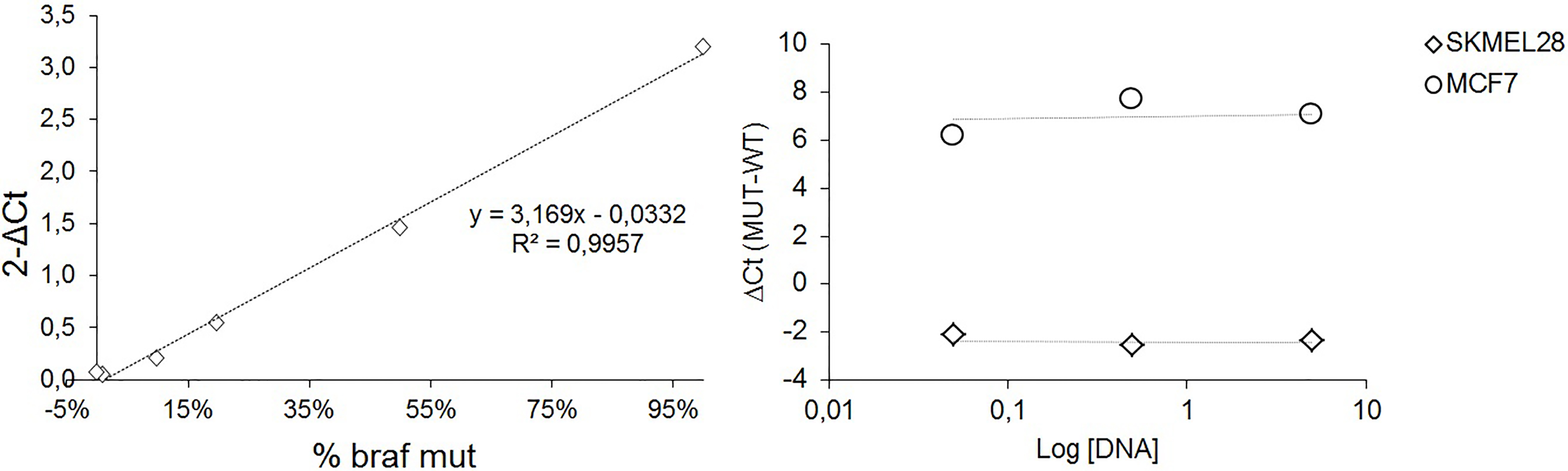
FFPE tissue specimens were available for 5/20 patients including 2 wild type and 3 mutated tissue specimens. All the analysed tissues derived from patients who showed CTCs in their blood. Both wild type tissues corresponded to BRAFV600E mutated CTC samples. Only one of the patients showing BRAFV600E mutation in the primary tumor was also BRAFV600E mutated in CTCs.
ICE-COLD PCR (Transgenomic) followed by Sanger Sequencing was used as a confirmatory test in qPCR mutated samples.
The mean quantity of ccfDNA extracted from plasma was 9.51 (
The evaluation of the presence of BRAFV600E mutation in DNA extracted from plasma obtained from the same samples submitted to CTC analysis detected four mutated samples. All the plasma-mutated samples were CTC positive and for three of them the mutation could be evidenced in paired DNA specimens extracted by ScreenCell MB enriched samples (Table 1).
Discussion
The term “liquid biopsy” refers to the possibility to obtain information from a minimally invasive blood draw analyzing circulating biomarkers, such as CTCs and ccfDNA, as surrogate materials of the tumor. This is particularly important in melanoma where tissue, whenever available, is scarce and mostly in metastatic patients for whom information obtained from the primary tumor does not reflect anymore the evolving disease. Furthermore, in the metastatic setting, the liquid biopsy reflects the heterogeneity of different metastatic sites and allows the longitudinal monitoring of the disease by serial blood sampling.
CTCs released into the bloodstream by the primary tumor and/or metastatic sites may play a role in the metastatic process but the mechanism is still not completely disclosed [26]. Despite the prognostic role of CTCs has been assessed in a variety of tumors, CTC analysis has not yet been translated into clinical practice mainly because CTC detection methods are technically challenging and lack a standardization. The CellSearch system, based on immunomagnetic capture of CTCs, is the only FDA-approved method for CTC detection and enumeration in metastatic breast, colon and prostate cancer. The CellSearch approach for melanoma relies on CD146 (MelCAM) and HMW-MAA (High Molecular Weight Melanoma-Associated Antigen) antibodies for melanoma CTC capture and detection, respectively. Nonetheless, some concerns have been raised about the opportunity of using methods based on the expression of cell surface markers for melanoma [27]. Other potential applications of CTC analysis are represented by screening and early detection of cancer and the real time monitoring of therapies by CTC counts [26]. On the other hand, shedding of CTCs may be discontinuous [28] and dependent on the characteristics of the neovascularization of the tumor [29].
This study was focused on the feasibility of CTC isolation and characterization based on the use of technically simple optimized filtration devices (ScreenCell Cyto and MB, ScreenCell, Paris, France). The filtration steps have been followed by cytological evaluation in the case of ScreenCell Cyto and by CTC genome characterization using a modified version of a previously developed allele specific qPCR-based assay for BRAFV600E quantitative detection after ScreenCell MB filtration and DNA extraction.
We found CTCs in 70% of the collected samples in line with our previously published paper with a different filtration method [21]. The rate of positive cases results higher than that found in two studies based on the immunomagnetic enrichment and CTC isolation by CellSearch [30, 31] but comparable to what reported by Roland et al. [32] using the same system. There are no papers from other groups performing the same CTC isolation method in melanoma patients with the exception of a proof-of principle demonstration for the application of three-dimensional telomeric analysis on cells isolated on filters [33]. On the other hand, by combining erythrocyte lysis or ficoll separation with immunomagnetic particles and qPCR [34], flow cytometry [35] or manual immunocytometry [36] higher percentages of CTC detection in metastatic melanoma patients have been reported (93, 88 and 100% respectively). The number of papers analyzing CTCs in melanoma patients is high and it is thus difficult to make a punctual comparison with each report (for a comprehensive review see references 3 and 4). Most of the works rely on the use of PCR-based indirect methods that possess high sensitivity, but that, on the other hand, are prone to false positive results due to illegitimate expression of tumor related markers from lymphocytes. The heterogeneity of the technological approaches and their combination with several different enrichment procedures and melanoma-related markers make it difficult to understand the features of an ideal method for CTC isolation.
Recently, with the emerging of biological therapies and precision medicine, the possibility to molecularly characterize the CTCs as regards some actionable mutations has become one of the features required in view of the clinical application of CTC analysis, thus opening new perspectives beyond the already established value of CTC count. In this context our approach might have a useful clinical application in treatment selection and in patients’ follow up during treatment or after interruption of combo-target therapy.
We were able to identify the BRAFV600E variant on CTCs of melanoma patients and the mutational status was confirmed by Sanger-Sequencing after ICE-COLD PCR amplification.
In our dataset, notwithstanding the low number of tissues available, it is interesting to notice that a wild type tissue corresponds to a CTC positive, BRAFV600E positive blood sample demonstrating that genetic alterations can change during tumor development as reported for other tumors [37].
Within the liquid biopsy approach, several studies in recent years have been focusing on the detection of mutations in ccfDNA, especially in relation with target therapies [1]. Sensitive assays for BRAFV600E detection in ccfDNA are still being developed and tested [38].
We are aware of the limited number of subjects investigated in the present study, not allowing to draw any general conclusion on the role of CTCs in this cluster of patients, nonetheless we found some promising results.
Interestingly in our study one subject showed mutant ccfDNA in contrast to wild type CTCs, while two patients with detectable BRAFV600E mutation in CTCs showed wild type ccfDNA. This discordance underlines the importance of analyzing both ccfDNA and CTCs, since they seem to represent two complementary aspects of the liquid biopsy providing different information. Nevertheless, we are convinced of the importance of analyzing DNA from CTCs, whenever possible, since CTCs derive directly from the primary tumor and/or metastatic sites and can be recognized for the tumor characteristic features reflecting tumor heterogeneity, while ccfDNA derives from both normal and tumor cells with not completely understood mechanisms. Both CTC and cfDNA analyses are technically challenging and need to be standardized also at the preanalytical level, in view of a clinical application of the liquid biopsy to melanoma patients.
The proposed approach benefits of a simple, fast filtration technique without the requirement of expensive equipment that could be easily managed as any other cytological specimen and used as a liquid biopsy of the tumor for the dynamic monitoring of the disease after surgery or whenever a tissue biopsy is not feasible.
Footnotes
Acknowledgments
We dedicate this paper to the memory of Prof. Claudio Orlando who started the research on liquid biopsy in our laboratory.
