Abstract
BACKGROUND:
Liposarcoma constitute about 13% of all soft tissue sarcoma and are associated with a high risk of metastases. As the preoperative differentiation between benign and malign lipomatous tumors is restricted to magnetic resonance imaging, computed tomography and biopsy, we performed a miRNA array to distinguish dedifferentiated liposarcoma patients from healthy controls and lipoma patients.
METHODS:
Blood samples of patients with dedifferentiated liposarcoma, healthy controls and lipoma patients were collected. Whole blood RNA was extracted and samples of patients with dedifferentiated liposarcoma (
RESULTS:
As shown by the microarray, two miRNAs (miR-3613-3p, miR-4668-5p) were shown to be significantly upregulated (fold change:
CONCLUSION:
We identified a specific whole blood miRNA (miR-3613-3p) that may serve to distinguish between dedifferentiated liposarcoma patients and healthy controls, thus potentially serving as a specific biomarker for dedifferentiated liposarcoma.
Background
Liposarcoma are a heterogeneous group of adipocy-tic tumors which can be classified into four main subtypes: atypic lipomatous tumors, dedifferentiated liposarcoma, myxoid liposarcoma and pleomorphic liposarcoma [1]. Since these subtypes differ significantly in terms of clinical behaviour, metastasis rate and sensitivity to radio- and chemotherapy [2, 3, 4], the correct diagnosis is of utmost importance.
MicroRNAs (miRNAs) are small, non-coding mole-cules which regulate the expression of target genes through mRNA degradation and translation inhibition [5] and have been described as potential biomarkers in various malignancies [6, 7, 8]. Recently, it has been shown that several miRNAs are deregulated in liposarcoma tumor tissue [9, 10] and even in the plasma of patients with dedifferentiated liposarcoma [11]. However, we felt that it was crucial to establish a miRNA biomarker for liposarcoma which was easier to transfer into clinical practice: Thus, in this study, blood sample collection was carried out by means of PAXgene Blood RNA Tubes, which immediately stabilize intracellular RNA, even if stored at room temperature for up to 72 hours.
Demographic patient data (age, BMI) and blood count of patients with G2 and G3 dedifferentiated liposarcoma and healthy donors
Demographic patient data (age, BMI) and blood count of patients with G2 and G3 dedifferentiated liposarcoma and healthy donors
Data are presented as mean value
Demographic data (Age and Sex), grading and therapy status of dedifferentiated liposarcoma patients
M0
MiRNAs with a fold change of
Since the availability of a non-invasive “liquid biopsy” for dedifferentiated liposarcoma might possibly improve survival rates due to earlier detection of sarcoma recurrence as well as allow more accurate differential diagnosis between the different subtypes, we performed a microarray-based miRNA screen of whole blood RNA of dedifferentiated liposarcoma patients compared to healthy controls, which was further validated by qRT-PCR.
Study population
The liposarcoma patients included in the study were patients receiving treatment from specialists in the interdisciplinary tumor board of the Comprehensive Cancer Center Freiburg (CCCF). Patients with a history of cancer other than liposarcoma, any type of systemic inflammatory disease or autoimmune disorder were excluded from the study. The control groups included healthy adults as well as lipoma patients matched to the liposarcoma groups in terms of body mass index (BMI).
Ethics, consent and permissions
Signed informed consent was obtained from all participants, allowing analysis of blood samples and all clinical data. The Ethics Committee of the Albert-Ludwigs University of Freiburg, Germany, approved the study. The design and performance of the study is in accordance with the Declaration of Helsinki.
Hierarchical clustering of all covered human mature miRNAs and human pre-miRNAs separated dedifferentiated liposarcoma samples from control samples and G2 liposarcoma samples from G3 liposarcoma samples.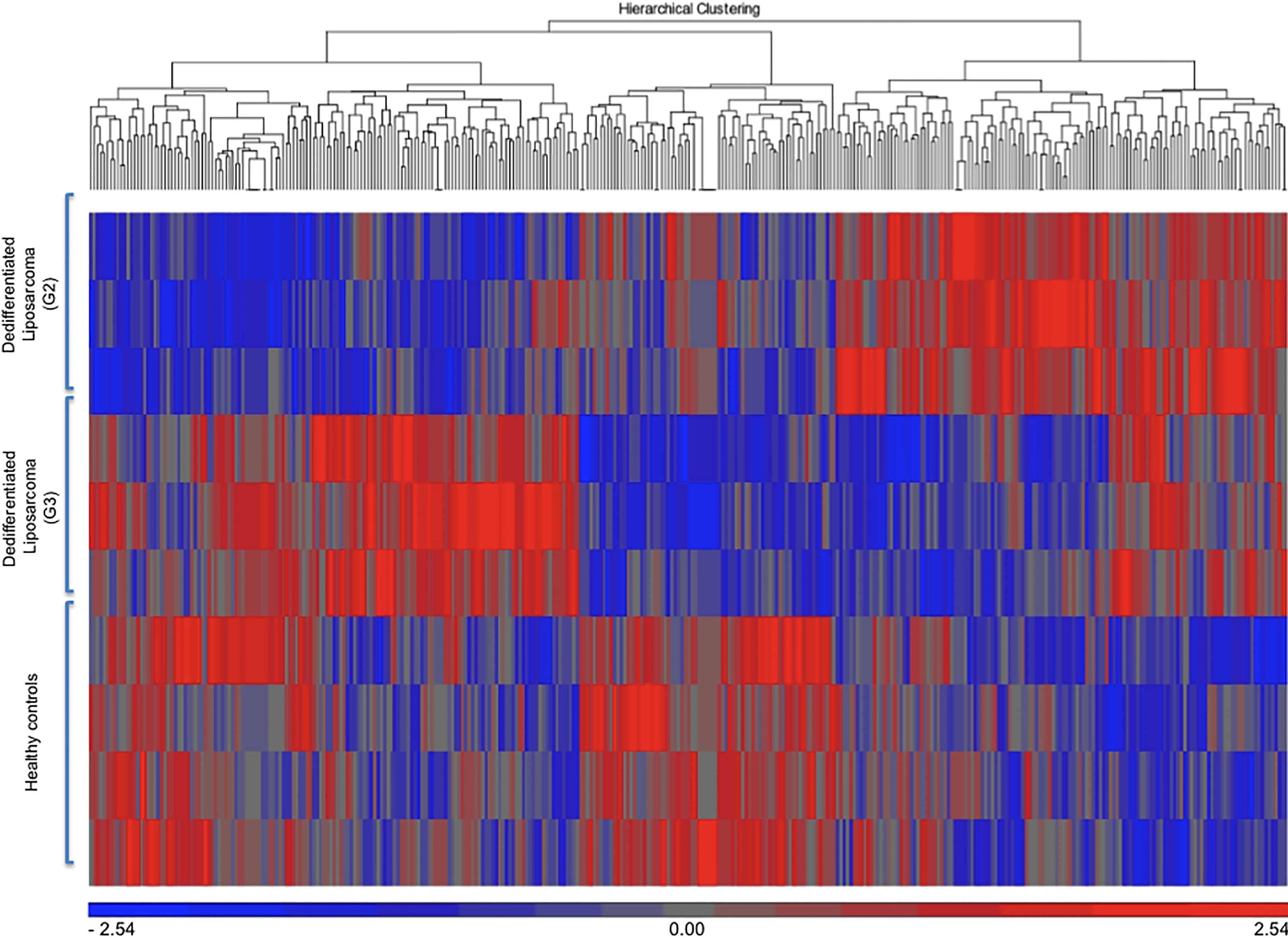
The blood samples were collected by puncture of the antecubital vein without tourniquet through a 20-gauge needle; each 2.5 ml of whole blood were collected and stabilized in PAXgene Blood RNA Tubes (PreAnalytiX, Hombrechtikon, Switzerland). The first 3 ml of blood were discarded. The RNA Tubes were left at least 2 hours at room temperature after blood collection; then being stored at
miRNA array
RNA quality and quantity was evaluated by capillary electrophoresis by the Fragment Analyzer and Standard sensitivity RNA Analysis kits (Advanced Analytical Technologies, Ames, IA, U.S.). Total whole blood RNA of patients with liposarcoma (
Preparation of cDNA
For detection of miRNA expression levels, cDNA synthesis of whole blood RNA (275 ng) was carried out using the miScript II RT Kit (Qiagen) according to the manufacturer’s protocol. The reverse transcription reaction was incubated for 60 min at 37
qRT-PCR
qRT-PCR was carried out using the miScript SYBR Green PCR Kit (Qiagen, Hilden, Germany) according to the manufacturer’s protocol. Cycling conditions consisted of an initial activation step of the HotStarTaq DNA Polymerase for 15 min at 95
Statistics
CEL-files of raw data were produced with Affym-etrix GeneChip Command Console Software Version 4.0. (Affymetrix, Santa Clara, CA, U.S.), then further being analyzed by Partek Genomics Suite software (Version 6.14.0923; Partek, Inc., St. Louis, MO, USA). CEL-files were imported including control and interrogating probes. Arrays were normalized using quantile normalization. Probeset summarization was performed using Median Polish. Probe values were log2 transformed. In order to detect differential miRNA expression between the 3 groups, 1-way ANOVA was performed [14] and Fisher’s Least Significant Difference (LSD) was used as contrast method.
Results
Dedifferentiated G2 and G3 liposarcoma patients (
Hierarchical clustering of all covered human mature miRNAs and human pre-miRNAs separated dedifferentiated liposarcoma samples from healthy control samples (Fig. 1). Following hierarchical clustering, the array results were narrowed down to deregulated miRNAs with a fold change >
MiR-3613-3p-expression in whole blood of patients with G2 dedifferentiated liposarcoma and G3 dedifferentiated liposarcoma compared to healthy donors.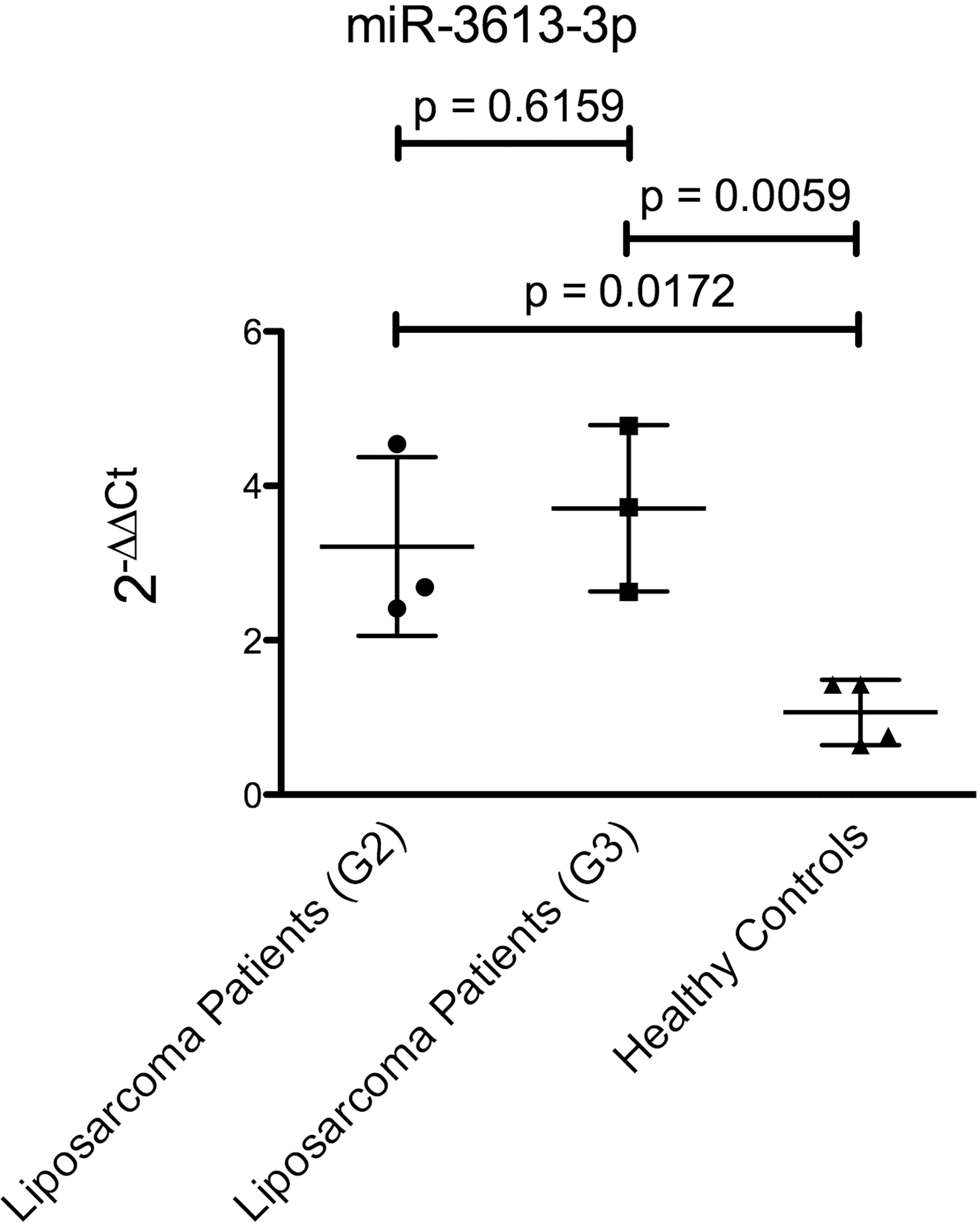
For further validation of the array, qRT-PCR was performed to confirm an upregulation of miR-3613-3p in the dedifferentiated liposarcoma samples (
MiR-3613-3p-expression in whole blood of patients with G2 dedifferentiated liposarcoma and G3 dedifferentiated liposarcoma compared to an independent cohort of healthy donors and lipoma patients.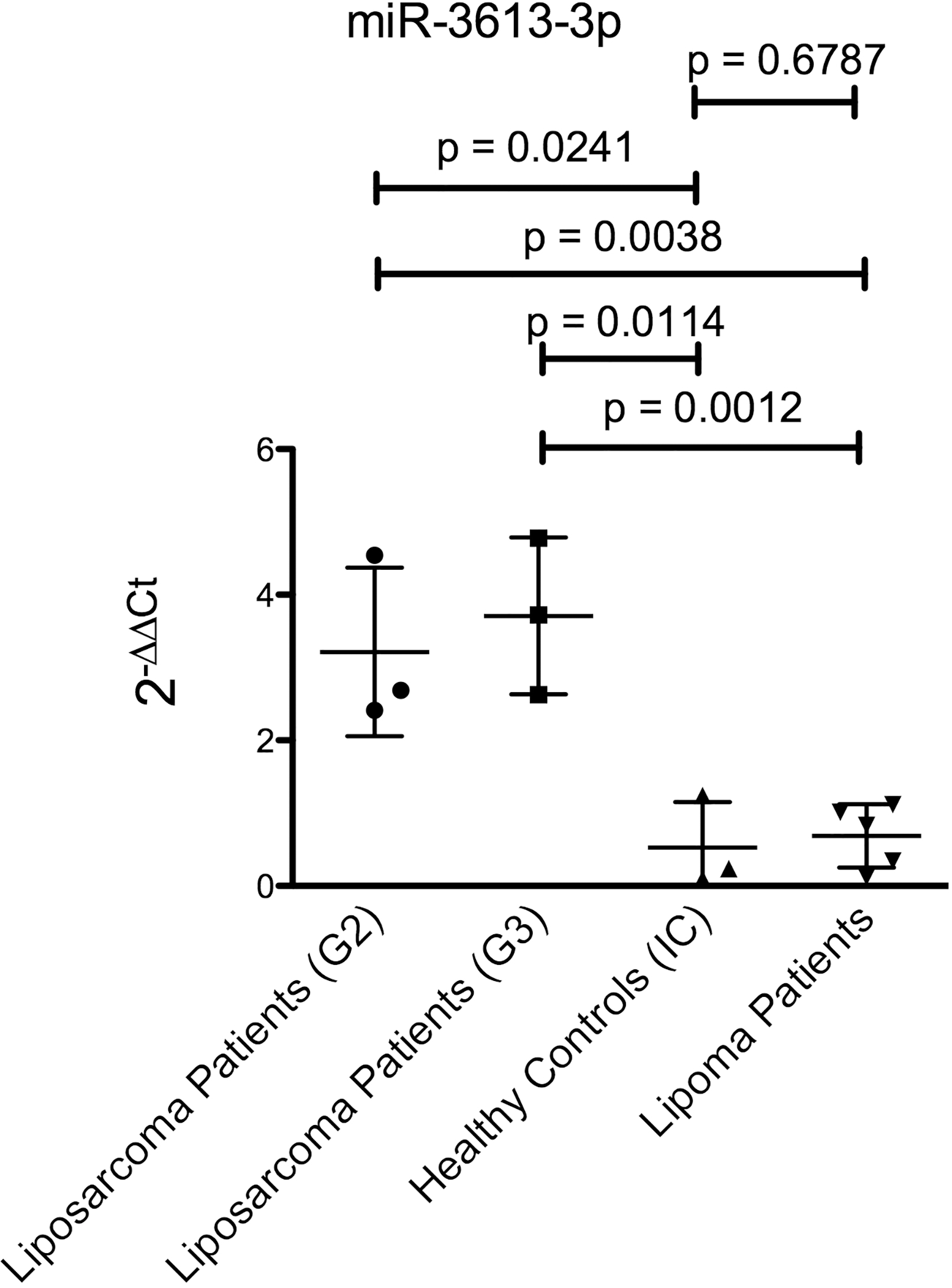
Biomarkers for liposarcoma are urgently needed to distinguish benign tumors such as lipoma (Fig. 4A and B) from liposarcoma (Fig. 4C and D), as well as identify patients with local recurrence and metastatic disease. Although several radiological predictors differentiating malignant liposarcoma from benign lipoma have been established [15], imaging studies are often inconclusive, since both lipoma and liposarcoma can present as well-circumscribed tumors, demonstrating large lobulated components of fat-signal (Fig. 4). In this context, it has been shown that lipoma cannot always be accurately differentiated from liposarcoma using imaging studies [16, 17]. Coran et al. examined MRI findings of 54 patients with lipomatous tumors, judging accuracy in distinguishing between lipoma and malignant lipomatous tumors according to size, localization, septa, nodules and signal homogeneity. Hereby, the authors found that although there are quite a few generally known signs that can indicate malignancy of the lesion such as tumor size
Magnetic resonance imaging of a 62-year male old patient with a lipoma of the right medial compartment of the thigh (A, coronal; B, axial) and an 87-year old female patient with a dedifferentiated liposarcoma (G3) of the ventromedial thigh (C, coronal; D, axial; T1-weighted images). Both tumors are well-circumscribed, demonstrating large lobulated components of fat-signal.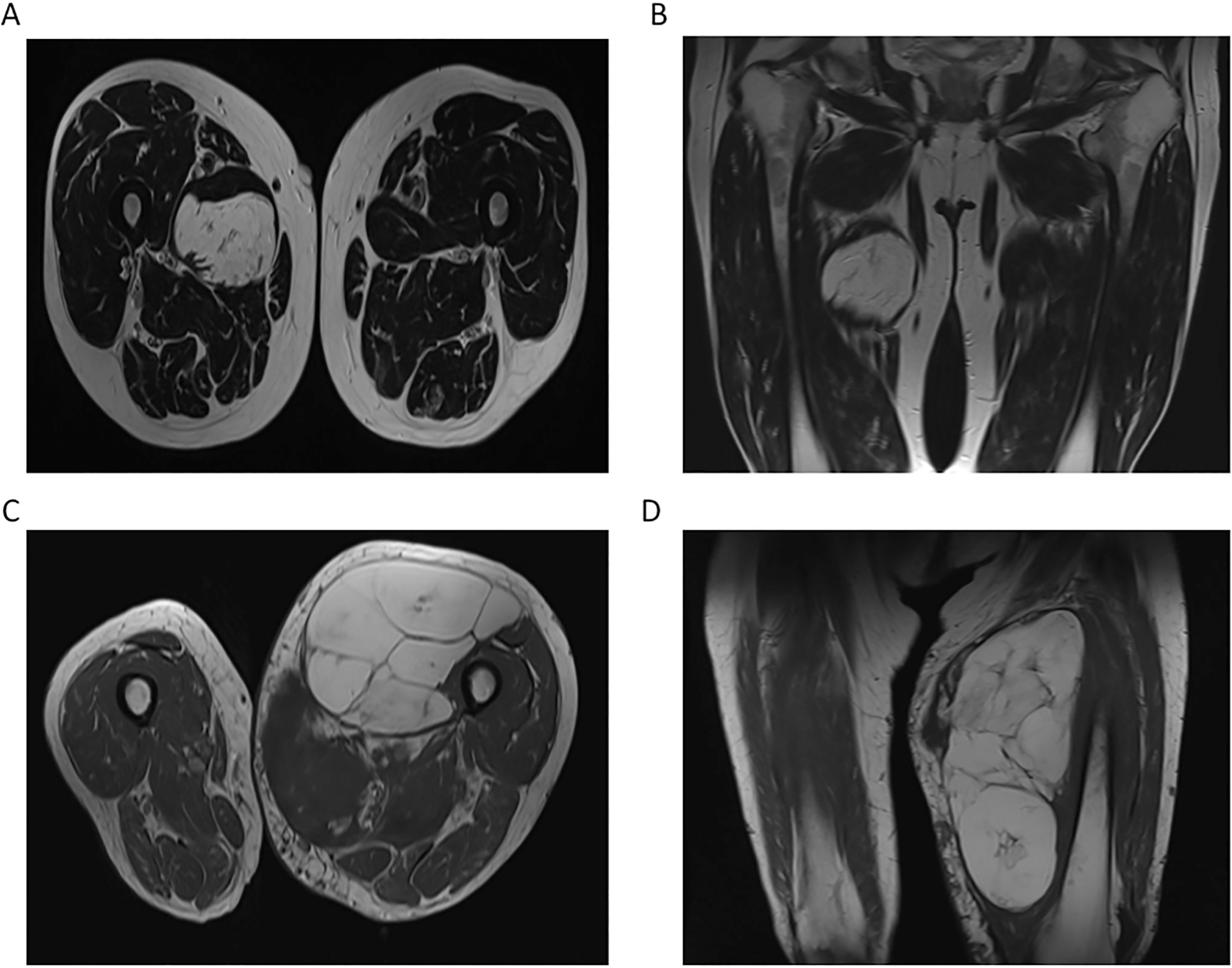
The aim of our study was to establish a miRNA biomarker for dedifferentiated liposarcoma which could easily be transferred into clinical practice: Using PAXgene Blood RNA Tubes for blood sample collection, samples can be stored at room temperature for up to 72 hours before being further processed, which might facilitate multi-centre studies. On the contrary, for serum or plasma miRNA profiling, immediate processing of the samples is necessary, which might lead to potential changes in miRNA levels by different processing conditions [21, 22].
Our results showed that the detection of miR-3613-3p in whole blood of dedifferentiated liposarcoma patients may serve as a ,,liquid biopsy“, a non-invasive way to differentiate between lipoma and dedifferentiated liposarcoma as well as detect distant metastasis or local recurrence of dedifferentiated liposarcoma. Interestingly, miR-3613-3p-values were higher in patients with localized disease compared to patients with metastatic liposarcoma (Supplement Figs 1 and 2). Nevertheless, the fact that localized liposarcomas yield higher miR-3613-3p-values than metastatic disease warrants further studies.
MiR-3613-3p has been described to play an important role in the histological classification and TNM staging of lung adenocarcinoma [23] and to be potentially involved in several important biological processes and functional pathways such as Wnt and Notch signaling [24]. Furthermore, bioinformatic analysis using the miRDB, Target Scan and TargetMiner databases found DFFB, APAF1, DICER, RORA, KIF3A, MAP3K1, NF1 and VHL to be possible mRNA targets binding miR-3613-3p prevalently in their 3’UTRs; however, performing qRT-PCR and western blot analyses, only target genes of the apoptotic machinery (APAF1, DFFB) were proven to be direct targets of miR-3613-3p [25].
Demographic patient data (age, BMI) and blood count of patients with dedifferentiated liposarcoma, healthy donors (Individual Cohort (IC)) and lipoma patients
Data are presented as mean value
Interestingly, the recently discovered MCPIP1 (monocyte chemoattractant protein-induced protein 1), a multidomain protein encoded by the MCPIP1 (ZC3H12A) gene which has been described to be downregulated in primary neuroblastoma, has been postulated to repress the expression of miR-3613-3p by cleavage of its precursor form [25]. Furthermore, MCPIP1 has been shown to play an important role in the immune system, being induced by inflammation-related factors, such as MCP-1, TNF
MiR-4668-5p was shown to be upregulated in dedifferentiated liposarcoma samples as compared to heal-thy donors throughout the array; however, this miRNA was not validated to be upregulated by qRT-PCR. Nevertheless, miR-4668-5p was only shown to be of importance in IgA nephropathy and tuberculosis [30, 31], not being linked to tumorigenesis, apoptosis or tumor-immune system interactions.
Contradictory findings of miRNA profiles in the peripheral blood and tumor tissue have been shown throughout previous studies [32, 33, 34]. In this context, Fehlmann et al., who identified several miRNAs which are rather specific for certain tissues or body fluids, also argued that miRNAs which are highly expressed in solid tissues are not necessarily found in the peripheral blood [35]. Interestingly, Mookherjee and El-Gabalawy found that whole blood miRNA profiles reflect miRNA expression patterns of peripheral blood mononuclear cells (PBMC) [36]. As previously hypothesized by the authors [21], these findings point out that the miRNA signature detected in whole blood RNA constitutes a response of the innate immune system to the tumor rather than the miRNA expression profile of the tumor itself. Therefore, it is not surprising that when analyzing whole blood RNA, we did not find an upregulation of miRNAs which have been previously been described to be overexpressed in liposarcoma tissue [9, 37, 38]. Correspondingly, Keller et al. found that only a few of the miRNAs deregulated in whole blood RNA were also previously described to be deregulated in solid tissues derived from patients with the same diseases [21, 39].
However, whole blood miRNA expression might be influenced by differing quantities of circulating blood cells. Analysis of age, BMI, Hb level, platelet count and leukocyte count was not statistically different when comparing dedifferentiated liposarcoma patients to the healthy donors evaluated in the microarray (Table 1), but G2 and G3 liposarcoma patients were significantly older than the independent cohort of healthy controls (
In summary, although the biological significance of miR-3613-3p in the liposarcoma-immune system interaction remains to be further clarified, the detected upregulation of miR-3613-3p might constitute a sensitive biomarker to distinguish dedifferentiated liposarcoma patients from lipoma patients.
Nevertheless, a further limitation of this study constitutes its low number of patients. However, one has to consider that liposarcoma is a rare disease; furthermore, dedifferentiated liposarcoma constitutes one of the less frequent subtypes of liposarcoma, therefore making the collection of a large number of samples challenging. Thus, further studies are warranted to validate the detected miRNA deregulation in a larger cohort of independent dedifferentiated liposarcoma patients.
MiRNA-3613-3p may serve as an independent bio-marker for the diagnosis of dedifferentiated liposarcoma, distinguishing dedifferentiated liposarcoma from healthy controls as well as from lipoma patients.
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-170496.
Footnotes
Acknowledgments
This study was supported by a project grant appointed to Dr. Alba Fricke by the Mattern foundation. Dr. Steffen U. Eisenhardt is supported by a Heisenberg Fellowship of the German Research Foundation (DFG) (EI866/3-1) and project grants EI866/1-2 and EI866/2-1 that are not related to the study.
Conflict of interest
The authors declare that they have no competing interests.
