Abstract
Oral submucous fibrosis (OSF) is a chronic insidious disease which predisposes to oral cancer. Understanding the molecular markers for OSF is critical for diagnosis and treatment of oral cancer. In this study, the proteins expression profile of OSF tissues was compared to normal mucous tissues by 2 dimensional electrophoresis (2-DE). The 2-DE images were analyzed through cut, spot detection and match analysis using mass spectrometry (MS). Differentially expressed genes were identified as candidates. RT-PCR, Western Blot and immunohistochemistry were performed to validate the difference in expression of the candidates between OSF and normal mucous tissues. The shRNA targeted to the candidates were then transfected by Lipofectamine2000 to the 3T3 cells to study gene function. Cell proliferation and apoptosis were measured by MTT, clonogenic formation, PI and TUNEL staining. From the proteomic analysis, 94 of the 182 selected spots with differential expression were identified by MS analysis and Cyclophilin A (CYPA) was determined to be the OSF-associated protein candidate. The significant differences in expression between OSF and normal tissues were verified and confirmed by RT-PCR, Western blot and Immunohistochemical analysis. Inhibition of CYPA expression by RNA interference suggested its potential activities involved in cell proliferation and apoptosis process. In conclusion, these results indicated a novel molecular mechanism of OSF pathogenesis and demonstrated CYPA as a potential biomarker and gene intervention targets of OSF. These data may help the development for therapeutics of oral cancer.
Keywords
Introduction
Oral submucous fibrosis (OSF) is a chronic pathological condition which is predisposes to oral cancer [1]. Histologically, OSF is characterized by epithelial atrophy and progressive accumulation of collagen fibers in the lamina propria and submucous of the oral mucosa with a progressive loss of vascularity [2]. As the condition continues, the jaws eventually become so rigid that the patients may not be able to open their mouth [3]. The condition is linked to areca nut chewing, a habit similar to tobacco chewing, which is practiced predominantly in Southeast Asia and India [4]. Regarded as a precancerous condition by WHO, OSF has become one of the major oral premalignant lesions. Despite many years’ research, the pathogenesis for the malignant transformation of OSF, especially from a molecular perspective, is still not very clear. Meanwhile, molecular markers that specifically marks OSF condition and its transitioning process to oral cancer is still missing in clinical practice [5]. It’s believed that the main problem is traditional single gene or multiple genes studies were not able to uncover the overall changes of gene expression in the OSF conditions [6].
Since molecular marker plays such a crucial role in diagnosis, monitoring, and treatment of OSF, screening the gene expression profile by high through-put technology to identify specific markers for OSF tissue sounds like an attractive strategy. Proteomics, as a study of the complete protein complement of the cell, is a promising approach in the identification of protein biomarkers with changed levels which may be useful as invaluable new targets for therapeutic intervention and markers for early detection [7]. In the past decade, proteomic approach has been successfully used to study many types of cancer, such as brain, lung, liver, stomach and colon cancers [8, 9, 10]. It was considered to be one of the best technology to identify important marker proteins for the elucidation of the molecular mechanisms of cancerogenous pathogenesis [11]. Two-dimensional gel electrophoresis (2-DE), mass spectrometry (MS) and bioinformatics are three major techniques used in proteomic studies. With combination of these techniques, novel biomarkers have been reported on oral cancers in recent years [12, 13]. However, as far as we understand, very few proteomic studies on OSF have been reported.
In this study, total proteins in OSF and normal mucous tissues were analyzed by 2-DE and the differentially expressed proteins were identified with MS. Cyclophilin A was selected as a potential marker for OSF pathogenesis. The expression of Cyclophinli A was validated by RT-PCR, Western Blot. Its activity in cell proliferation and apoptosis was valued with RNA interference. Our data could be used for diagnosis and treatment OSF related oral cancer.
Materials and methods
Patient samples
Twenty five samples of OSF tissue were obtained from Department of stomotolgy of Xiangya Hospital, Changsha, China, between August 2006 and December 2007. Normal mucous tissues were collected at the West China School of Stomatology, Sichuan University in the same time period. Fresh tissue were separated in two halves. One half of the sample was snap frozen and stored in
Two-dimensional electrophoresis and image analysis
Tissue were minced in liquid nitrogen and lysed in lysis buffer (7 M urea, 2 M thiourea, 4% CHAPS, 2 mM TBP, 2 mM PMSF, 1 mg/ml DNase I and 0.2 mg/ml RNase A) [14]. After centrifugation followed by vortex and incubation, the supernatant was precipitated with cold actone/trichloroacetic acid. The pellet was resolved in rehydration buffer (7 M urea, 2 M thiourea, 4% CHAPS, 2 mM TBP, 0.2% carrier ampholytes). Total protein concentration was determined with Pierce
In-gel digestion
Spots of interest were excised and in-gel digested according to the manufacturer’s instruction (Promega, Madison, WI) with some modifications. Briefly, the gel spots were destained twice with 0.2 ml of 100 mM NH
Mass spectrometry analysis
1.5
Semi-quantitative RT-PCR
The primer sequences and the expected sizes of PCR products were as follows: Cyclophilin A: (Forward) 5’-CATGGTCAACCCCA CGTGTTCTT-3’ and (Reverse) 5’-TAG ATG GAC TTG CCA CCA GTG CCA T-3’ (236 bp); GAPDH: (Forward) 5’-AGCTTCGGCA CATATTTCATCTG-3’ and (Revserse) 5’-CGTTCAC TCCCATGACAAACA-3’ (311 bp). Total RNA was extracted using Trizol reagent (Invitrogen, Carlsbad, CA) and RT-PCR was performed with conditions as follows: reverse transcription at 48
Western blotting
Tissue samples were ground in liquid nitrogen and lysed in RIPA lysis buffer (50 mM Tris-HCl (pH 7.4), 0.25 % sodium deoxycholate, 150 mM NaCl, 1% NP-40, 1 mM EDTA, 1 mM NaF, 1 mM Na3V4, 1 mM PMSF). Lysates were subjected to 12% SDS-PAGE, transferred to polyvinylidene difluoride (PVDF) membranes (Millipore, Bedford, MA), and immunoblotted with rabbit anti-human Cyclophilin A antibody (Upstate Biotechnology, Charlottesville, VA). The blots were labeled with horseradish peroxidaseconjugated secondary antibody, visualized by chemiluminescent detection.
Immunohistochemistry
The sections were stained by the Envision System-HRP method (DakoCytomation Inc, Carpinteria, CA), according. Specific antibodies performed included rabbit anti-human Cyclophilin A, and goat anti-human PCNA (Santa Cruz Biotechnology, Santa Cruz, CA). Immunostaining intensity (A) was classified as lack of staining (0), mild staining (1), moderate staining (2), and strong staining (3). The proportion of staining-positive cells (B) was semi-quantitatively divided into 4 grades, as
siRNA synthesis and shRNA plasmid vector construction
A double strand siRNA oligonucleotide targeting Cyclophilin A (sense: 5’-AAGAUGAGAACUUCAUC CUdTdT-3’; antisense: 5’-AGGAUGAAGUUCUCAU CUUdTdT-3’) with dTdT-overhangs at 3’ end was designed according to the published sequence of human Cyclophilin A (NM_021130). This sequence, corresponding to the nucleotides 265–283 bp, had been used in a previous study [14], and proved to be specific and effective. The HK sequence, with no homology to any human gene, was used as negative control. The plasmid Pgenesil-2 containing kanamycin resistance gene was linearized with BamHI and HindIII, and the annealed oligonucleotide templates were ligated into a plasmid vector using T4 DNA ligase. Chemically competent DH5
Cell culture and transfection
The murine fetal 3T3 fibroblast cell line (ATCC, VA) was maintained in Roswell Park Memorial Institute (RPMI)
Cell proliferation and apoptosis assays
Cells seeded on 96-well plates in triplicate were transfected with shRNA at various concentrations. After incubation at 37
For clonogenic formation assay, 100 counted cells transfected with siRNA were seeded in triplicate in a 6-well plate, and cultured continuously for 14 days. Clones were stained with Gimsa, and counted under a microscope. A cluster with more than 50 cells was considered as a clone.
Propidium Iodide (PI, Invitrogen, Carlsbad, CA) staining was carried out 72 hrs after transfection. TUNEL staining was performed according to the manufacturer direction (Promega, Madison, WI). The cells were then observed under a fluorescence microscope (Olympus, Tokyo, Japan).
Data analysis and Statistics
All quantitative data were recorded as mean
Proteins up-regulated in OSF
Proteins up-regulated in OSF
Proteins down-regulated in OSF
Protein profile differences between oral submucous fibrotic tissue and normal mucous tissue. (A) H&E staining of OSF tissue and normal mucous tissue. Bar 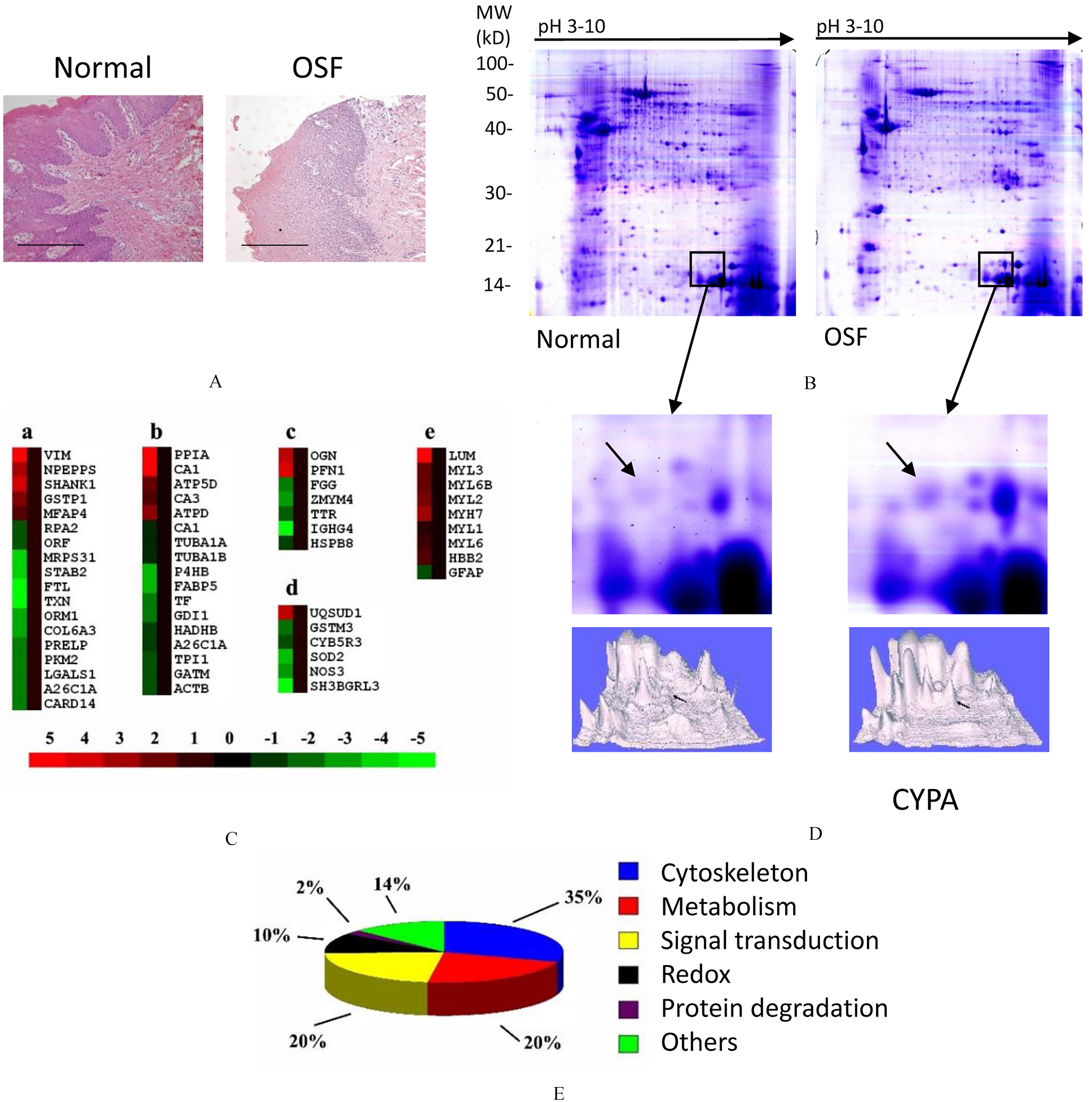
Identification of Cyclophinlin A in croppted 2-DE gel. MS/MS analysis revealed 1 matched-peptides for Cyclophilin A, with 8% sequence coverage and a MOWSE score of 114.
Confirmation of Cyclophilin A up-regulation in SOF tissue. (A) RT-PCR indicated 2.1-fold (t test, 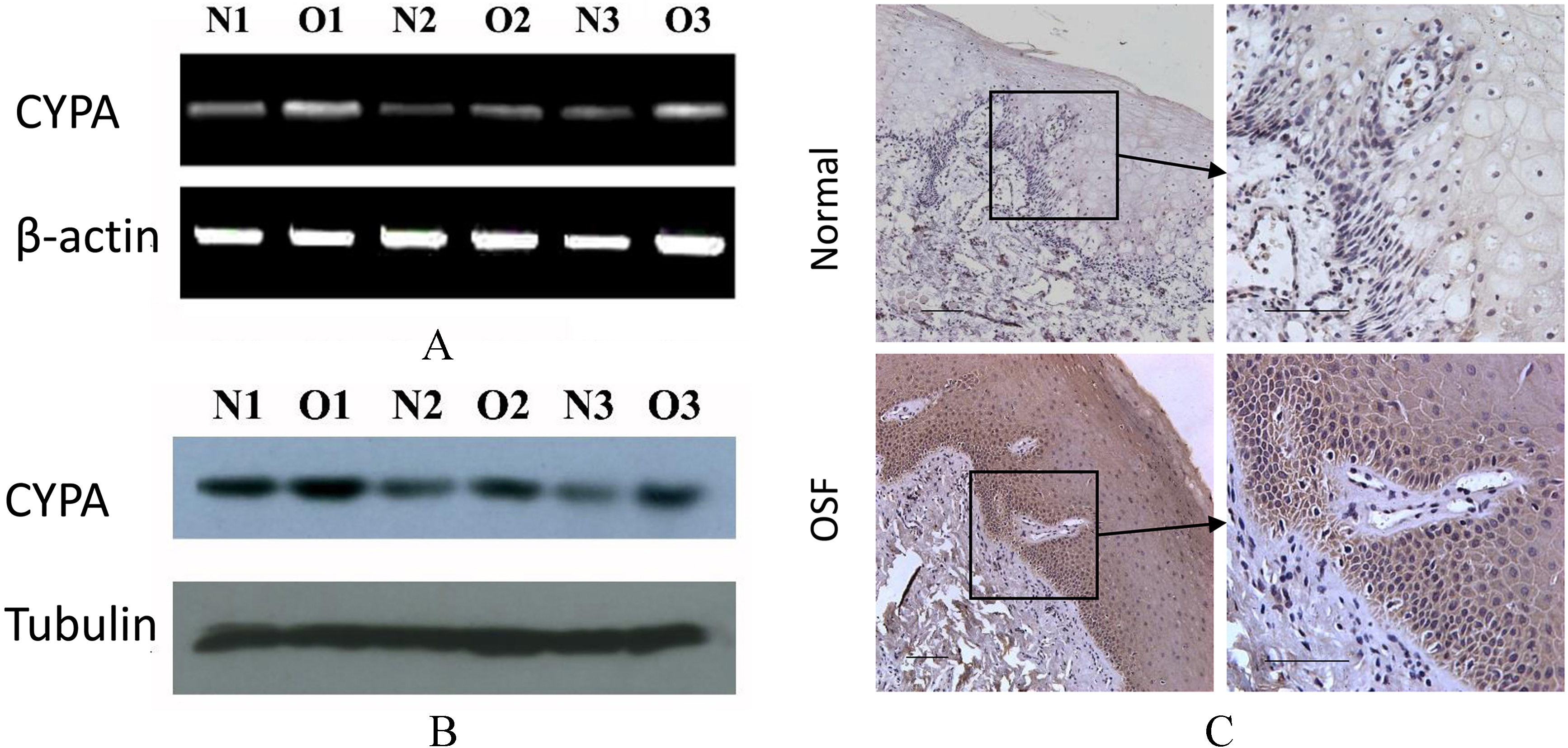
Suppression of Cyclophilin A by shRNA inhibits the proliferation of fibroblasts. (A) Seventy-two hours after transfection, total proteins were collected for western blot. (B) Quantification of western blots indicated that CYPA-shRNA transfected 3T3 fibroblasts had significantly less expression of CYPA (Dunnett-t test, 
Suppression of Cyclophilin A by shRNA increases the apoptosis of fibroblasts. (A) Propidium iodide (PI) staining was performed 72 hr after transfection. Bar 
Comparison of global expression profile of proteins between SOF and normal mucous tissue
Before proteomic analysis, H&E staining was carried out to confirm the pathological conditions of tissue samples. Normal submucous tissue contained regular epithelial spikes and clear basement membrane, while OSF tissue had edema between collagen fibers, invasion of lymphocyte, and irregular epithelial spikes (Fig. 1A). Two-dimensional electrophoresis was then performed on 25 normal mucous tissue samples and 25 OSF tissue samples. Image analysis was performed using PDQuest 7.0 software. 2D gels displayed well-resolved and reproducible protein profiles for both tissues. Comassie staining visualized on averages of 1000 spots (Fig. 1B) in samples both tissues. Spots at different expression levels (182 spots in total) were analyzed using MALDI-Q-TOF (matrix-assisted laser desorption/ionization quadrupole time-of-flight) tandem mass spectrometry. A total of 94 proteins were identified, among which 38 were up-regulated in OSF tissue while 56 were down-regulated in OSF tissue (Fig. 1C). Among these proteins, most of them were involved in cellular functions, including cytoskeleton, metabolism, signaling transduction, Redox, and protein degradation (Fig. 1E). Full lists of proteins that differentially expressed in two types of tissues were in Tables 1 and 2.
Among the 94 differentially expressed proteins, we chose Cyclophinlin A for detailed analysis due to several reasons. Firstly, the expression of Cyclophilin A was found to be reliably different between normal and OSF tissues in all detected samples (Fig. 1D). Then, the mass spectrum results revealed that the peptide had8% sequence coverage and a MOWSE score of 114 (Fig. 2), which is an indication of accuracy for protein matching. At last, Cyclophinlin A had been reported in literature as an important marker in many carcinomas [15].
Up-regulation of Cyclophilin A in OSF tissue was confirmed at both mRNA and protein levels
To confirm the difference in expressions of Cyclophilin A in OSF and normal mucous tissue, RT-PCR was carried out to measure the gene expression at mRNA level. As shown in Fig. 3A, the mRNA level of Cyclophilin A was observed between the two tissues (OSF tissues: 70.35
Suppression of Cyclophilin A by shRNA inhibit proliferation of fibroblasts
To investigate the function of Cyclophilin A in fibroblasts, shRNA sequences were used to transfect fibroblasts 3T3 cells. As shown by western blot, the expression of Cyclophilin A was significantly suppressed 72 hr after transfection (Fig. 4A). Quantification of blots showed its expression was reduced by more than 70%. Scrambled shRNA did not inhibit the expression of Cyclophilin A in a side by side experiment (Fig. 4B). The proliferation of transfected cells was studied by MTT assay and clonogenic formation. Data from MTT assay showed that the proliferation was significantly suppressed by transfection of Cyclophilin A shRNA in a duration-dependent manner (Fig. 4C). After 14 days culture in the clonogenic formation experiment, the clone numbers in untreated control, lipo and HK-shRNA groups, were all significantly larger than the clone number in Cyclophilin A-shRNA group (Dunnett-t test,
Suppression of Cyclophilin A by shRNA induced apoptosis of fibroblasts
At last, we examined the effects of Cyclophilin A on the apoptosis of fibroblasts by PI and TUNEL staining. Upon PI staining, apoptotic cells represented smaller cell size and lower forward scattered light than normal cells (Fig. 5A). TUNEL staining visualized a large amount of apoptotic cells in CYPA-shRNA transfected group comparing to control groups (Fig. 5B). There were only minimal numbers of cells in untreated and negative control groups showed apoptotic features upon PI staining, however, the CYPA-shRNA transfected group had almost 40% of apoptotic cells (Dunnett-t test,
Discussion
In this study, high-resolution and reproducible protein expression patterns were obtained from 25 samples of OSF tissue and 25 samples of normal mucous tissue by 2D electrophoresis. Approximately 182 protein spots were successfully identified including 94 different proteins. 38 proteins were up-regulated while 56 proteins were down-regulated in OSF tissue comparing to normal tissue. By analysis and classification of those proteins, Cyclophilin A was identified as OSF-associated candidate protein. RT-PCR, western blot and Immunohistochemical analysis of tissues exhibited an increased pattern of Cyclophilin A expression in OSF than in normal mucous tissues. Transfection of CYPA-shRNA into 3T3 cells resulted significant inhibition of Cyclophilin A expression. The MTT assay and clonogenic formation test showed that proliferation and cloning efficiency were inhibited by knocking down Cyclophilinin in 3T3 cells. Apoptosis was also induced by the shRNA transfection.
The peptidylprolyl cis-trans-isomerase (PPIse) family includes Cyclophilin (CyP), FK506 binding protein (FKBP) and parvulin [16]. The common feature of this family of protein is the enzymatic active group consist of 120 amino acid residuals which regulate the activity and stability of the target protein. PPIse was believed to regulate the functions of proteins involved in cell cycles [17]. This family of proteins was also shown to play important roles in autoimmune diseases with its regulatory effects on immunological response [15]. It’s been reported to be up-regulated in the serum of patients with systemic lupus erythematosus, rheumatoid arthritis and psoriasis [18, 19, 20]. As a member of the peptidylprolyl cis-trans-isomerase (PPIse) family, Cyclophilin A is an 18 kDa protein with a broad range of functions in cell proliferation and apoptosis. Initial studies mainly focused on its role in protein folding, immune response and HIV-1 infection [21, 22, 23]. It was generally regarded as the intracellular receptor for cyclosporin A [24]. But it’s later been found in the secretomes of tumor cells as a response to inflammatory stimulation [25]. The Cyclophilin-Acyclosporin A complex was reported to suppress the activity of calcineurin and nuclear transportation of nuclear factor of activated T-cells (NFAT) in T lymphocytes and other immune cells [26, 27].
Recent studies had identified Cyclophilin A as a candidate marker gene in many carcinomas. Up-regulation of Cyclophilin A had been observed in lung cancer [28], cholangiocarcinoma [29], pancreatic cancer [30], myeloma [31] and gastric carcinoma [32]. Our study was the first one to report the up-regulation of Cyclophilin A in oral submuscous fibrotic tissue. This broadened our understanding of the roles of Cyclophilin A in carcinogenesis. These results demonstrated that overexpression of Cyclophilin A was important for the pathogenic process of fibroblasts predispose to cancer cells. Regarding the mechanism how Cyclophilin A involved in carcinogenesis, it’s generally believed to promote cells proliferation and inhibit apoptosis through caspase deactivation [33]. In support of this, our data showed that knockdown of Cyclophilin A in fibroblasts by shRNA inhibit cell proliferation and increase apoptosis.
In conclusion, the present study demonstrated that CYPA is a potential biomarker and gene intervention targets of OSF. These results may uncover novel mechanisms for OSF pathogenic process. These data improve our understanding of the molecules in the initiation and development of OSF, and help the development for therapeutics of oral cancer.
